

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0132.6
(162 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
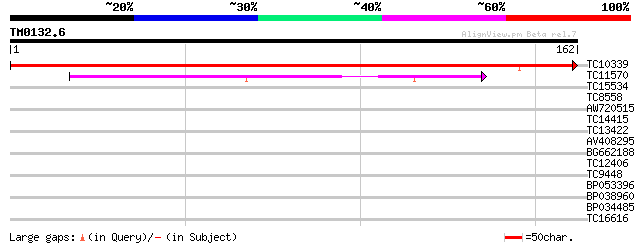
Score E
Sequences producing significant alignments: (bits) Value
TC10339 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protei... 294 4e-81
TC11570 similar to UP|O22165 (O22165) 60S ribosomal protein L30 ... 62 5e-11
TC15534 weakly similar to GB|AAO63420.1|28950993|BT005356 At5g28... 30 0.16
TC8558 weakly similar to UP|Q9NDI0 (Q9NDI0) 200 kDa antigen p200... 30 0.26
AW720515 29 0.35
TC14415 similar to UP|O04237 (O04237) Transcription factor, part... 29 0.35
TC13422 27 1.3
AV408295 27 1.7
BG662188 27 1.7
TC12406 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, p... 26 2.9
TC9448 similar to GB|CAB81029.1|7269936|ATCHRIV72 cyclic nucleot... 25 5.0
BP053396 25 6.5
BP038960 25 8.5
BP034485 25 8.5
TC16616 similar to GB|AAM47365.1|21360499|AY113057 AT4g32140/F10... 25 8.5
>TC10339 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 677
Score = 294 bits (753), Expect = 4e-81
Identities = 147/164 (89%), Positives = 155/164 (93%), Gaps = 2/164 (1%)
Frame = +2
Query: 1 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQ 60
MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCK YFHNRLKPSKLTWTAM+RKQ
Sbjct: 74 MVLKTELCRFSGAKIYPGRGIRFIRSDSQVFLFVNSKCKHYFHNRLKPSKLTWTAMYRKQ 253
Query: 61 HKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALREIKERVK 120
HKKDIAQEA +K+RR+TKKPYSRSIVGATLEVIQK+RTEKPEVRDAAREAALREIKER+K
Sbjct: 254 HKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDAAREAALREIKERIK 433
Query: 121 KTKDEKRAKKAEVQTKAQKSQGKG--GKVAAPKGPKLGGGGGKR 162
KTKDEK+AKKAE K+QKSQGKG K AAPKGPKLGGGGGKR
Sbjct: 434 KTKDEKKAKKAETTAKSQKSQGKGNVSKGAAPKGPKLGGGGGKR 565
>TC11570 similar to UP|O22165 (O22165) 60S ribosomal protein L30
(At2g44860/T13E15.13), partial (64%)
Length = 1282
Score = 62.0 bits (149), Expect = 5e-11
Identities = 37/123 (30%), Positives = 67/123 (54%), Gaps = 4/123 (3%)
Frame = +1
Query: 18 GRGIRFIRSDSQVFLFVNSKCKRYFHNRLKPSKLTWTAMFRKQHKKDIA--QEAARKKRR 75
G GI+F+R+D+++F F SKC + F + P K+ WT +R+ H K++ Q +++R
Sbjct: 424 GHGIQFVRNDAKIFRFYRSKCHKNFKMKRNPRKVKWTKAYRRVHGKEMTRIQTFEFERKR 603
Query: 76 STKKPYSRSIVGATLEVIQKRRTEKPEVRDAAREAALRE--IKERVKKTKDEKRAKKAEV 133
+ + Y R++ L+ I K A+ A+RE +++K+ K EK ++AE
Sbjct: 604 NRPERYDRNVKEDVLKAIPK----------IAKIRAIREETHHKKMKQGKHEKLRREAEK 753
Query: 134 QTK 136
+ K
Sbjct: 754 ELK 762
>TC15534 weakly similar to GB|AAO63420.1|28950993|BT005356 At5g28040
{Arabidopsis thaliana;}, partial (8%)
Length = 937
Score = 30.4 bits (67), Expect = 0.16
Identities = 26/99 (26%), Positives = 44/99 (44%), Gaps = 10/99 (10%)
Frame = -3
Query: 67 QEAARKKRRSTKKPYSR--------SIVGATLEVIQKRRTEKPEVRDAAREAALREIKER 118
+E +K +KK ++R +I+ + I K + + DA + +R ++
Sbjct: 365 EEDGKKSGNDSKKQFTRLWSDEDEIAILKGLADFISKTGNDPLKYPDAFYDFVIRSLQAD 186
Query: 119 VKKT--KDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKL 155
+T KD+ R K + QT A GKG PK PK+
Sbjct: 185 ATRTQVKDKVRRLKKKFQTLA----GKGKNGETPKFPKV 81
>TC8558 weakly similar to UP|Q9NDI0 (Q9NDI0) 200 kDa antigen p200
(Fragment), partial (4%)
Length = 585
Score = 29.6 bits (65), Expect = 0.26
Identities = 27/94 (28%), Positives = 42/94 (43%), Gaps = 19/94 (20%)
Frame = +1
Query: 82 SRSIVGATLEVIQKRRTEK-PEVRDAAREAAL------------------REIKERVKKT 122
S ++V A L+ ++R + E +DAA AAL RE E +K
Sbjct: 10 SPTLVAARLDAEERRNNIRLREEQDAAYRAALEADQARERQRREEQERLAREAAEAERKR 189
Query: 123 KDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLG 156
K+E+ A++ E + A+K Q K+ K LG
Sbjct: 190 KEEEEAREREAREAAEK-QAALAKIRQQKAESLG 288
>AW720515
Length = 472
Score = 29.3 bits (64), Expect = 0.35
Identities = 29/106 (27%), Positives = 44/106 (41%), Gaps = 2/106 (1%)
Frame = -2
Query: 44 NRLKPSKLTWTAMFRKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQK--RRTEKP 101
N LK + W F H KDI +E +RRS G + K R+ +
Sbjct: 279 NFLK*FLVIWV*FF---HSKDIVKENLNHRRRS----------GLLTKACMKMNRKLREG 139
Query: 102 EVRDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGKGGKV 147
VR ++ +A + E K+ KR K + + + +S G GG V
Sbjct: 138 VVRSSSSKAFPLIVYEDPFGDKERKRVSKVQKKEEQWRSGGIGGTV 1
>TC14415 similar to UP|O04237 (O04237) Transcription factor, partial (83%)
Length = 1666
Score = 29.3 bits (64), Expect = 0.35
Identities = 14/27 (51%), Positives = 18/27 (65%)
Frame = +2
Query: 136 KAQKSQGKGGKVAAPKGPKLGGGGGKR 162
KA + +G G AP+GP GGGGG+R
Sbjct: 587 KASERRGYG----APRGPYRGGGGGRR 655
>TC13422
Length = 490
Score = 27.3 bits (59), Expect = 1.3
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = -1
Query: 32 LFVNSKCKRYFHNRLKPSKLTWTAMF 57
LF+ S +YFH+ L PSK+ A F
Sbjct: 226 LFLTSSQDKYFHSYLLPSKVNTCAFF 149
>AV408295
Length = 426
Score = 26.9 bits (58), Expect = 1.7
Identities = 17/62 (27%), Positives = 28/62 (44%)
Frame = +1
Query: 43 HNRLKPSKLTWTAMFRKQHKKDIAQEAARKKRRSTKKPYSRSIVGATLEVIQKRRTEKPE 102
H+ L+P + +K+ K + RKKR KK + + +++KRR K
Sbjct: 133 HHPLRPRRTKLQLQLQKKK*KKKKRRKQRKKRNLLKKRRRKR*LKKKRNLLKKRRRPKLR 312
Query: 103 VR 104
VR
Sbjct: 313 VR 318
>BG662188
Length = 358
Score = 26.9 bits (58), Expect = 1.7
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = +1
Query: 123 KDEKRAKKAEVQTKAQKSQGKGGKVAAPK 151
++EKR KAE + KA K K K A P+
Sbjct: 217 EEEKRCGKAEAERKAAKKGSKSKKGAPPE 303
>TC12406 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, partial (6%)
Length = 450
Score = 26.2 bits (56), Expect = 2.9
Identities = 20/48 (41%), Positives = 25/48 (51%), Gaps = 3/48 (6%)
Frame = +2
Query: 116 KERVKKTKDEKRAKKAEVQTKAQK---SQGKGGKVAAPKGPKLGGGGG 160
K R K+ K K KKAE + K +K +Q + A GP GGGGG
Sbjct: 194 KRRGKRRKKHK--KKAEKKRKIRKILMAQVQVQPQNAMPGPNGGGGGG 331
>TC9448 similar to GB|CAB81029.1|7269936|ATCHRIV72 cyclic nucleotide and
calmodulin-regulated ion channel-like protein
{Arabidopsis thaliana;} , partial (5%)
Length = 553
Score = 25.4 bits (54), Expect = 5.0
Identities = 12/20 (60%), Positives = 14/20 (70%)
Frame = +3
Query: 141 QGKGGKVAAPKGPKLGGGGG 160
+G+GGK GPK GGGGG
Sbjct: 411 KGEGGKKL---GPKKGGGGG 461
>BP053396
Length = 467
Score = 25.0 bits (53), Expect = 6.5
Identities = 13/39 (33%), Positives = 16/39 (40%)
Frame = +2
Query: 45 RLKPSKLTWTAMFRKQHKKDIAQEAARKKRRSTKKPYSR 83
+L+ L W QH I R KRR T P S+
Sbjct: 197 KLRKRLLFW*PSMPSQHMDRIGTRIPRTKRRRTMSPLSK 313
>BP038960
Length = 524
Score = 24.6 bits (52), Expect = 8.5
Identities = 16/52 (30%), Positives = 25/52 (47%)
Frame = +3
Query: 93 IQKRRTEKPEVRDAAREAALREIKERVKKTKDEKRAKKAEVQTKAQKSQGKG 144
+QK E+P V D+ EAA + E ++ ++E+ V A GKG
Sbjct: 270 LQKGEAEQPRVEDSEEEAADYDDDEEEEEEEEEEDEHADRV---AVSGPGKG 416
>BP034485
Length = 485
Score = 24.6 bits (52), Expect = 8.5
Identities = 11/28 (39%), Positives = 14/28 (49%)
Frame = +3
Query: 134 QTKAQKSQGKGGKVAAPKGPKLGGGGGK 161
+ K + GG +G K GGGGGK
Sbjct: 180 KNKNKNKNNGGGNNKGKEGKKGGGGGGK 263
>TC16616 similar to GB|AAM47365.1|21360499|AY113057 AT4g32140/F10N7_50
{Arabidopsis thaliana;}, partial (27%)
Length = 883
Score = 24.6 bits (52), Expect = 8.5
Identities = 14/43 (32%), Positives = 22/43 (50%)
Frame = -3
Query: 119 VKKTKDEKRAKKAEVQTKAQKSQGKGGKVAAPKGPKLGGGGGK 161
V + + RA+ Q + ++ + GG A G + GGGGGK
Sbjct: 194 VNENRATNRARSDPKQERQRRGRDGGG--GAGGGGRDGGGGGK 72
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,167,329
Number of Sequences: 28460
Number of extensions: 24803
Number of successful extensions: 206
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 204
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 204
length of query: 162
length of database: 4,897,600
effective HSP length: 83
effective length of query: 79
effective length of database: 2,535,420
effective search space: 200298180
effective search space used: 200298180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0132.6


