

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0131.12
(78 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
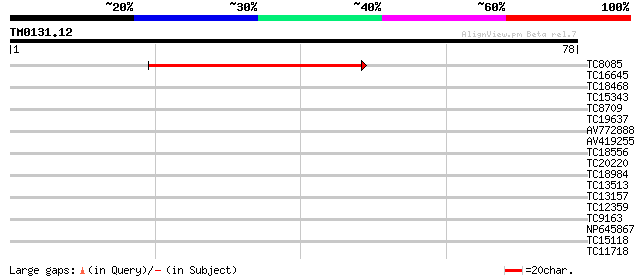
Score E
Sequences producing significant alignments: (bits) Value
TC8085 similar to GB|BAB03033.1|11994717|AP001300 AtDMC1 (meioti... 69 1e-13
TC16645 36 0.001
TC18468 26 1.5
TC15343 similar to UP|Q9SJM8 (Q9SJM8) Expressed protein, partial... 25 2.0
TC8709 similar to UP|O49929 (O49929) Pore protein of 24 kDa (OEP... 25 2.6
TC19637 similar to UP|Q9T0D0 (Q9T0D0) T5C23.60 protein, partial ... 25 2.6
AV772888 25 3.4
AV419255 25 3.4
TC18556 similar to UP|Q9XIN0 (Q9XIN0) At2g27230 protein (BHLH tr... 24 5.9
TC20220 23 7.6
TC18984 homologue to UP|AGO1_ARATH (O04379) Argonaute protein, p... 23 7.6
TC13513 similar to UP|O65516 (O65516) Aldehyde dehydrogenase lik... 23 10.0
TC13157 23 10.0
TC12359 homologue to UP|Q08150 (Q08150) GTP-binding protein 6, p... 23 10.0
TC9163 similar to UP|Q9AT45 (Q9AT45) ADP-glucose pyrophosphoryla... 23 10.0
NP645867 proliferating floral organs protein [Lotus japonicus] 23 10.0
TC15118 23 10.0
TC11718 23 10.0
>TC8085 similar to GB|BAB03033.1|11994717|AP001300 AtDMC1 (meiotic
recombination protein)-like protein {Arabidopsis
thaliana;} , partial (8%)
Length = 500
Score = 69.3 bits (168), Expect = 1e-13
Identities = 29/30 (96%), Positives = 29/30 (96%)
Frame = +1
Query: 20 MNNPDHVWNITWKLLANDIQHEYRGTLTQP 49
M NPDHVWNITWKLLANDIQHEYRGTLTQP
Sbjct: 115 MTNPDHVWNITWKLLANDIQHEYRGTLTQP 204
>TC16645
Length = 596
Score = 36.2 bits (82), Expect = 0.001
Identities = 14/36 (38%), Positives = 23/36 (63%)
Frame = +2
Query: 11 FASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTL 46
F +L+TN+++ P VW +W LL + I ++ R TL
Sbjct: 362 FTRMLMTNLISRPQEVWRKSWSLLCDGILYDRRKTL 469
>TC18468
Length = 630
Score = 25.8 bits (55), Expect = 1.5
Identities = 18/77 (23%), Positives = 33/77 (42%), Gaps = 1/77 (1%)
Frame = +2
Query: 2 HSGNELRRLF-ASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSK 60
H+ ++L+ + +S LIT + NPD +W++++
Sbjct: 191 HASSKLKAIVRSSPLITQKLLNPDVLWSLSY----------------------------- 283
Query: 61 PILLKIDNHLQSNGKSL 77
+DNHL++NGK L
Sbjct: 284 -----VDNHLRTNGKEL 319
>TC15343 similar to UP|Q9SJM8 (Q9SJM8) Expressed protein, partial (49%)
Length = 689
Score = 25.4 bits (54), Expect = 2.0
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = -1
Query: 48 QPGMIPTFSLSSKPILLKIDNHLQSNGKS 76
Q IPTFS+S ++I+NH+ N ++
Sbjct: 560 QSRAIPTFSIS*YGYNIRIENHMVQNARA 474
>TC8709 similar to UP|O49929 (O49929) Pore protein of 24 kDa (OEP24),
complete
Length = 997
Score = 25.0 bits (53), Expect = 2.6
Identities = 11/30 (36%), Positives = 18/30 (59%)
Frame = -1
Query: 16 ITNIMNNPDHVWNITWKLLANDIQHEYRGT 45
I +++NP+ V + K+L DI +Y GT
Sbjct: 328 IQRLLSNPNSVHELKPKVLFGDIVVDYEGT 239
>TC19637 similar to UP|Q9T0D0 (Q9T0D0) T5C23.60 protein, partial (48%)
Length = 603
Score = 25.0 bits (53), Expect = 2.6
Identities = 11/25 (44%), Positives = 15/25 (60%)
Frame = +1
Query: 1 HHSGNELRRLFASLLITNIMNNPDH 25
HHS RRL +SLL + + +P H
Sbjct: 37 HHSDMASRRLLSSLLRSPLTRSPPH 111
>AV772888
Length = 465
Score = 24.6 bits (52), Expect = 3.4
Identities = 21/78 (26%), Positives = 35/78 (43%), Gaps = 5/78 (6%)
Frame = +1
Query: 1 HHSGNELRRLFASLLITNIMNNPDHV-WNITWKLLANDIQHEYRGTLTQP----GMIPTF 55
HH N L + +I N + W I W++ ++ + +G + P GM+ T
Sbjct: 31 HHKNN----LTGAKIIKK*WNGIQYAGWFINWEI*SHRL*---KGRASVPTGCLGMVNTS 189
Query: 56 SLSSKPILLKIDNHLQSN 73
+ K LKI+ HL+ N
Sbjct: 190 TFPMKTFNLKIERHLEIN 243
>AV419255
Length = 421
Score = 24.6 bits (52), Expect = 3.4
Identities = 8/38 (21%), Positives = 21/38 (55%)
Frame = -3
Query: 2 HSGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQ 39
H+ L +++ + ++ PD+ W +W+ ++D+Q
Sbjct: 269 HTRYHLSSYKSTVDVNCLVQKPDYTWQRSWQNYSHDLQ 156
>TC18556 similar to UP|Q9XIN0 (Q9XIN0) At2g27230 protein (BHLH
transcription factor), partial (24%)
Length = 710
Score = 23.9 bits (50), Expect = 5.9
Identities = 9/23 (39%), Positives = 12/23 (52%)
Frame = -2
Query: 31 WKLLANDIQHEYRGTLTQPGMIP 53
WK N QH TLT+ ++P
Sbjct: 97 WKYFENSFQHHPCYTLTEENLLP 29
>TC20220
Length = 514
Score = 23.5 bits (49), Expect = 7.6
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = +1
Query: 15 LITNIMNNPDHVWNITWKLL 34
++T+ NP +WN TW LL
Sbjct: 190 VVTHAQPNP*PLWNGTWCLL 249
>TC18984 homologue to UP|AGO1_ARATH (O04379) Argonaute protein, partial
(22%)
Length = 698
Score = 23.5 bits (49), Expect = 7.6
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +2
Query: 23 PDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLS 58
P H+ ++K+ + Q RGTLT GMI +S
Sbjct: 365 PKHIGRSSFKIFSXQWQDPVRGTLT-GGMIKELLIS 469
>TC13513 similar to UP|O65516 (O65516) Aldehyde dehydrogenase like protein,
partial (12%)
Length = 540
Score = 23.1 bits (48), Expect = 10.0
Identities = 8/14 (57%), Positives = 10/14 (71%)
Frame = +2
Query: 26 VWNITWKLLANDIQ 39
VWN+ WK+L IQ
Sbjct: 110 VWNVPWKVLV*HIQ 151
>TC13157
Length = 548
Score = 23.1 bits (48), Expect = 10.0
Identities = 7/21 (33%), Positives = 12/21 (56%)
Frame = +3
Query: 24 DHVWNITWKLLANDIQHEYRG 44
D +W +T + L D H++ G
Sbjct: 402 DKLWGLTCRFLGYDFSHQFIG 464
>TC12359 homologue to UP|Q08150 (Q08150) GTP-binding protein 6, partial
(30%)
Length = 360
Score = 23.1 bits (48), Expect = 10.0
Identities = 13/26 (50%), Positives = 16/26 (61%)
Frame = -1
Query: 41 EYRGTLTQPGMIPTFSLSSKPILLKI 66
EY+ LT G T SL S+ I+LKI
Sbjct: 225 EYQLCLTASG*AATNSLQSRHIILKI 148
>TC9163 similar to UP|Q9AT45 (Q9AT45) ADP-glucose pyrophosphorylase large
subunit (Glucose-1-phosphate adenylyltransferase)
(ADP-glucose synthase) , partial (5%)
Length = 663
Score = 23.1 bits (48), Expect = 10.0
Identities = 8/13 (61%), Positives = 10/13 (76%)
Frame = +2
Query: 40 HEYRGTLTQPGMI 52
H Y GT T+PGM+
Sbjct: 377 HSYSGTHTKPGML 415
>NP645867 proliferating floral organs protein [Lotus japonicus]
Length = 1350
Score = 23.1 bits (48), Expect = 10.0
Identities = 16/43 (37%), Positives = 20/43 (46%), Gaps = 3/43 (6%)
Frame = -1
Query: 22 NPDHVWNITWKLLANDIQHEYRGTLTQ---PGMIPTFSLSSKP 61
+P WNI LL QH+ TL Q P P SL+ +P
Sbjct: 1026 HPQTPWNIQLALLNRRNQHQLLPTLNQVRRP*KPPHRSLNLEP 898
>TC15118
Length = 657
Score = 23.1 bits (48), Expect = 10.0
Identities = 6/10 (60%), Positives = 7/10 (70%)
Frame = +2
Query: 22 NPDHVWNITW 31
NPDH W + W
Sbjct: 128 NPDHCWEMMW 157
>TC11718
Length = 759
Score = 23.1 bits (48), Expect = 10.0
Identities = 10/29 (34%), Positives = 18/29 (61%)
Frame = -2
Query: 49 PGMIPTFSLSSKPILLKIDNHLQSNGKSL 77
P IPT + +LLK+D +++G+S+
Sbjct: 644 PESIPTSGSYPQKLLLKMDLDFRNDGRSI 558
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,635,387
Number of Sequences: 28460
Number of extensions: 22134
Number of successful extensions: 86
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 85
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 86
length of query: 78
length of database: 4,897,600
effective HSP length: 54
effective length of query: 24
effective length of database: 3,360,760
effective search space: 80658240
effective search space used: 80658240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0131.12


