

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
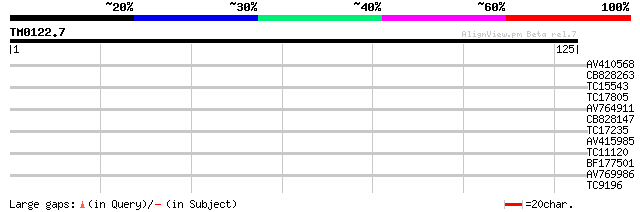
Query= TM0122.7
(125 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV410568 27 0.77
CB828263 26 1.7
TC15543 similar to UP|Q84K86 (Q84K86) 4-coumarate:CoA ligase-lik... 25 3.8
TC17805 weakly similar to UP|O82681 (O82681) Lanatoside 15'-O-ac... 25 5.0
AV764911 24 6.6
CB828147 24 6.6
TC17235 UP|Q93XE5 (Q93XE5) Glutathione synthetase, complete 24 6.6
AV415985 24 6.6
TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete 24 6.6
BF177501 24 6.6
AV769986 24 8.6
TC9196 weakly similar to UP|Q9LUC0 (Q9LUC0) Gb|AAD10662.1, parti... 24 8.6
>AV410568
Length = 394
Score = 27.3 bits (59), Expect = 0.77
Identities = 13/45 (28%), Positives = 22/45 (48%)
Frame = -1
Query: 62 LQIDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAE 106
+ + +G G E + + H PSR+F+PL + +TK E
Sbjct: 142 ISFSTRTIGAGCSSESNNFTQPLLHLAWPSRAFNPLYKESITKVE 8
>CB828263
Length = 570
Score = 26.2 bits (56), Expect = 1.7
Identities = 12/21 (57%), Positives = 13/21 (61%)
Frame = +3
Query: 79 DGSNSFPHFYGPSRSFSPLSL 99
+G PHF GPSR S LSL
Sbjct: 15 EGMRFHPHFSGPSRRQSSLSL 77
>TC15543 similar to UP|Q84K86 (Q84K86) 4-coumarate:CoA ligase-like, partial
(21%)
Length = 579
Score = 25.0 bits (53), Expect = 3.8
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = +2
Query: 103 TKAEKLSLSDGRFTCCLLDQV 123
TKA KL L R CCL+ Q+
Sbjct: 56 TKASKLLLQS*RLFCCLIPQL 118
>TC17805 weakly similar to UP|O82681 (O82681) Lanatoside
15'-O-acetylesterase precursor, partial (12%)
Length = 484
Score = 24.6 bits (52), Expect = 5.0
Identities = 10/30 (33%), Positives = 16/30 (53%)
Frame = -2
Query: 9 ISTAKEWEELQANGSTFGGNLDKSSAFIHL 38
+S K W+ELQ +G G ++ +I L
Sbjct: 402 VSNLKHWKELQVSGFHLGSGINMMHDWIFL 313
>AV764911
Length = 321
Score = 24.3 bits (51), Expect = 6.6
Identities = 10/18 (55%), Positives = 15/18 (82%)
Frame = +1
Query: 95 SPLSLDVVTKAEKLSLSD 112
SPL+ ++ ++AEKLSL D
Sbjct: 196 SPLAWEIKSEAEKLSLED 249
>CB828147
Length = 570
Score = 24.3 bits (51), Expect = 6.6
Identities = 11/28 (39%), Positives = 16/28 (56%)
Frame = -3
Query: 90 PSRSFSPLSLDVVTKAEKLSLSDGRFTC 117
P SFS L+ V++ +L+ S RF C
Sbjct: 97 PPASFSIRDLNAVSRRRRLTCSQVRFQC 14
>TC17235 UP|Q93XE5 (Q93XE5) Glutathione synthetase, complete
Length = 2152
Score = 24.3 bits (51), Expect = 6.6
Identities = 11/30 (36%), Positives = 19/30 (62%)
Frame = -1
Query: 41 LDQVRSTLERFYMNCKEELYLLQIDAKKLG 70
LDQ+ L RFY+ + + YLL++ + +G
Sbjct: 976 LDQL---LPRFYVQSQSDAYLLELQKENVG 896
>AV415985
Length = 277
Score = 24.3 bits (51), Expect = 6.6
Identities = 14/35 (40%), Positives = 20/35 (57%)
Frame = -3
Query: 91 SRSFSPLSLDVVTKAEKLSLSDGRFTCCLLDQVVS 125
SR S L+ + T K S + RFT C++ +VVS
Sbjct: 275 SREISSLT-SMATG*SKFSAAGSRFTTCMVRRVVS 174
>TC11120 UP|Q84PP4 (Q84PP4) Transcription factor MYB102, complete
Length = 1281
Score = 24.3 bits (51), Expect = 6.6
Identities = 18/70 (25%), Positives = 25/70 (35%)
Frame = +2
Query: 29 LDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAKKLGDGLVYEMVDGSNSFPHFY 88
L AF L+ + V+ L + + L D + + SN PHF
Sbjct: 638 LTNMEAFSLLNSISNVKENLGVDFSQMDTQTSLFSHDGNNMASS--QPLHHPSNMLPHFL 811
Query: 89 GPSRSFSPLS 98
P SFS S
Sbjct: 812 DPQVSFSSQS 841
>BF177501
Length = 247
Score = 24.3 bits (51), Expect = 6.6
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = -1
Query: 100 DVVTKAEKLSLSDGRFTCCLLD 121
DV K E L+ G+ TCCLLD
Sbjct: 154 DVHLKEEFLNQFLGKGTCCLLD 89
>AV769986
Length = 178
Score = 23.9 bits (50), Expect = 8.6
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = -1
Query: 86 HFYGPSRSFSPLSLDVVTK 104
HF+ PSRS P+SL ++ K
Sbjct: 76 HFFPPSRS*RPVSLCILYK 20
>TC9196 weakly similar to UP|Q9LUC0 (Q9LUC0) Gb|AAD10662.1, partial (25%)
Length = 799
Score = 23.9 bits (50), Expect = 8.6
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +3
Query: 41 LDQVRSTLERFYMNCKEELYLLQIDAKKLG 70
L + RS+ +RF M +E LLQ+ + LG
Sbjct: 195 LGRWRSSAQRFRMQRREHALLLQLGIQVLG 284
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,122,549
Number of Sequences: 28460
Number of extensions: 24380
Number of successful extensions: 108
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 108
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 108
length of query: 125
length of database: 4,897,600
effective HSP length: 80
effective length of query: 45
effective length of database: 2,620,800
effective search space: 117936000
effective search space used: 117936000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 50 (23.9 bits)
Lotus: description of TM0122.7


