

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.6
(378 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
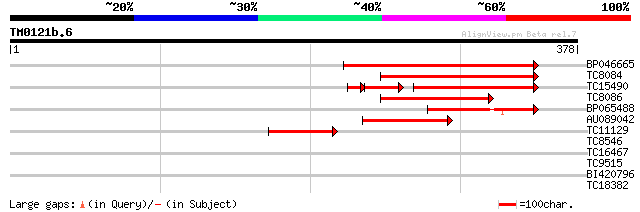
Score E
Sequences producing significant alignments: (bits) Value
BP046665 159 7e-40
TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombin... 132 7e-32
TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like prote... 101 3e-28
TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination p... 86 1e-17
BP065488 69 1e-12
AU089042 57 7e-09
TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protei... 55 3e-08
TC8546 28 2.5
TC16467 UP|NUCC_LOTJA (Q9BBN8) NAD(P)H-quinone oxidoreductase ch... 27 4.3
TC9515 similar to UP|O49081 (O49081) Clp-like energy-dependent p... 27 5.6
BI420796 27 5.6
TC18382 similar to PIR|S44026|S44026 thioredoxin-disulfide reduc... 27 7.3
>BP046665
Length = 524
Score = 159 bits (402), Expect = 7e-40
Identities = 80/131 (61%), Positives = 96/131 (73%), Gaps = 1/131 (0%)
Frame = -1
Query: 223 DYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGLCNGTRMIVNALTKYIIVAIVLNRNTMGE 282
D C GIPNH+I LK G PIML+RNI Q G CNGTR+IV L +I A V+ +G+
Sbjct: 524 DITCFGIPNHKITLKEGAPIMLLRNIYQAVGFCNGTRLIVADLGTNVIKATVIT*TNIGD 345
Query: 283 TTFIPRMSLTPSNADIPFKFQRRQFPVILCFAMTINKSQGQSSTHVGLYLPRPVFTHVQL 342
FIPRM + PS++ PFKF+RRQFP+ LC AMTINKSQGQS +HVGLYL R VFTH QL
Sbjct: 344 DIFIPRMDMVPSDSGYPFKFERRQFPISLCSAMTINKSQGQSLSHVGLYLSRHVFTHGQL 165
Query: 343 YVAL-SRVKSR 352
YVAL +++K R
Sbjct: 164 YVALHAKIKKR 132
>TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (4%)
Length = 560
Score = 132 bits (333), Expect = 7e-32
Identities = 65/105 (61%), Positives = 79/105 (74%)
Frame = +1
Query: 248 IDQTAGLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQF 307
I + LCNGTR+IV L Y+I A V+ +G+ FIPR+ + PS++ PFKF+RR F
Sbjct: 43 IQKFENLCNGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*F 222
Query: 308 PVILCFAMTINKSQGQSSTHVGLYLPRPVFTHVQLYVALSRVKSR 352
P+ LCFAMTINKSQGQS +HV LYL RPVFTH QLYVALSRV+SR
Sbjct: 223 PISLCFAMTINKSQGQSLSHVSLYLSRPVFTHGQLYVALSRVRSR 357
>TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like protein, partial
(8%)
Length = 634
Score = 101 bits (251), Expect(3) = 3e-28
Identities = 50/83 (60%), Positives = 61/83 (73%)
Frame = +2
Query: 270 IVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFAMTINKSQGQSSTHVG 329
I V+ +G+ IPRM + PS++ PFKF+RRQ P+ LCFAMTINKSQG+S +HVG
Sbjct: 137 IQVTVITGTHIGDDISIPRMDMVPSDSSYPFKFERRQSPISLCFAMTINKSQGRSLSHVG 316
Query: 330 LYLPRPVFTHVQLYVALSRVKSR 352
LYL RPV TH LYVAL RV+SR
Sbjct: 317 LYLSRPVSTHG*LYVALPRVRSR 385
Score = 39.3 bits (90), Expect(3) = 3e-28
Identities = 18/26 (69%), Positives = 21/26 (80%)
Frame = +1
Query: 237 KVGVPIMLIRNIDQTAGLCNGTRMIV 262
K G IML+RNI Q +GLCNGTR+IV
Sbjct: 43 KEGALIMLLRNIVQASGLCNGTRLIV 120
Score = 21.2 bits (43), Expect(3) = 3e-28
Identities = 8/12 (66%), Positives = 9/12 (74%)
Frame = +3
Query: 226 CSGIPNHQIKLK 237
CS IPNH+I K
Sbjct: 9 CSDIPNHKITEK 44
>TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination protein
DMC1/LIM15 homolog, partial (12%)
Length = 628
Score = 85.5 bits (210), Expect = 1e-17
Identities = 40/75 (53%), Positives = 53/75 (70%)
Frame = +1
Query: 248 IDQTAGLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSNADIPFKFQRRQF 307
I + LC+GTR+IV L Y+I A V+ +G+ FIPR+ + PS++ PFKF+RR F
Sbjct: 151 IQKFENLCHGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*F 330
Query: 308 PVILCFAMTINKSQG 322
P+ LCFAMTINKSQG
Sbjct: 331 PISLCFAMTINKSQG 375
>BP065488
Length = 439
Score = 68.9 bits (167), Expect = 1e-12
Identities = 41/78 (52%), Positives = 52/78 (66%), Gaps = 4/78 (5%)
Frame = -1
Query: 279 TMGETTFIPRMSLTPSNADIPFKFQRRQFPVILCFAMTINKSQGQSSTH---VGLYLPRP 335
T G+ FIPRM++ PS + P KF+R QFP+ LCFAMTINKS Q S H V L
Sbjct: 385 TXGDDIFIPRMNMVPSVSGDPLKFERCQFPISLCFAMTINKS--QXSVHYPTVALISFLA 212
Query: 336 VFTHV-QLYVALSRVKSR 352
++H+ +LYVALS V+SR
Sbjct: 211 CYSHMDKLYVALSGVRSR 158
>AU089042
Length = 191
Score = 56.6 bits (135), Expect = 7e-09
Identities = 27/60 (45%), Positives = 40/60 (66%)
Frame = +3
Query: 236 LKVGVPIMLIRNIDQTAGLCNGTRMIVNALTKYIIVAIVLNRNTMGETTFIPRMSLTPSN 295
LK GVP+ML+ N+ + GLCNGTR+IV+ L +I A +L+ +G +I M+L PS+
Sbjct: 9 LKXGVPVMLMXNLXISTGLCNGTRLIVDYLGPNVIGATILSGTHIGXVVYISMMNLXPSD 188
>TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protein disulfide
isomerase A6 precursor (P5) , partial (7%)
Length = 600
Score = 54.7 bits (130), Expect = 3e-08
Identities = 24/46 (52%), Positives = 33/46 (71%)
Frame = +2
Query: 173 PTLESVEEINNFMLAMILGEEIEYLNCDTPCKSDEDSGVNAEWVTS 218
PTLESVE++N FML ++ G EYL+ DT CK DED+ + + W T+
Sbjct: 458 PTLESVEKVNEFMLDLLPGNTTEYLSSDTTCKYDEDTELQS*WFTT 595
>TC8546
Length = 541
Score = 28.1 bits (61), Expect = 2.5
Identities = 15/41 (36%), Positives = 21/41 (50%)
Frame = -2
Query: 217 TSEFLNDYKCSGIPNHQIKLKVGVPIMLIRNIDQTAGLCNG 257
TS F YKC G Q+K K+ P + +NI++ C G
Sbjct: 123 TSPFFIIYKCLG---QQVKKKLQCPTLKRKNIEEK*NTCTG 10
>TC16467 UP|NUCC_LOTJA (Q9BBN8) NAD(P)H-quinone oxidoreductase chain H,
chloroplast (NAD(P)H dehydrogenase, chain H)
(NADH-plastoquinone oxidoreductase 49 kDa subunit) ,
complete
Length = 2407
Score = 27.3 bits (59), Expect = 4.3
Identities = 14/37 (37%), Positives = 17/37 (45%)
Frame = -1
Query: 117 TEQLQPLMKLNHSFRFLLIFIEPCKDPLLEIVNFSYP 153
T L P K + LIF PC+ + EI F YP
Sbjct: 694 THPLNPFQKNRIARNKFLIFNNPCEKIIAEIKTFIYP 584
>TC9515 similar to UP|O49081 (O49081) Clp-like energy-dependent protease
(ATP-dependent Clp protease proteolytic subunit)
(Endopeptidase Clp) , partial (74%)
Length = 1158
Score = 26.9 bits (58), Expect = 5.6
Identities = 14/41 (34%), Positives = 21/41 (51%)
Frame = -1
Query: 300 FKFQRRQFPVILCFAMTINKSQGQSSTHVGLYLPRPVFTHV 340
+K+ ILCF TINK++ QS + +P V H+
Sbjct: 867 YKYHANNLGSILCFQYTINKTKLQSLRNRKKTVPLCVLLHL 745
>BI420796
Length = 552
Score = 26.9 bits (58), Expect = 5.6
Identities = 19/51 (37%), Positives = 22/51 (42%)
Frame = +2
Query: 287 PRMSLTPSNADIPFKFQRRQFPVILCFAMTINKSQGQSSTHVGLYLPRPVF 337
P L PS + +P + Q P IL QGQS YLPRP F
Sbjct: 296 PPPLLPPSPSPLPQPRRHGQPPKILLNHRPFPCRQGQSGLVARPYLPRPRF 448
>TC18382 similar to PIR|S44026|S44026 thioredoxin-disulfide reductase 2 -
Arabidopsis thaliana (fragment)
{Arabidopsis thaliana;} , partial (53%)
Length = 503
Score = 26.6 bits (57), Expect = 7.3
Identities = 21/84 (25%), Positives = 43/84 (51%), Gaps = 5/84 (5%)
Frame = -1
Query: 124 MKLNHSFRFLLIFIEPCKDPLLE--IVNFSYPKLLFNLEKNSFFQERAI---LAPTLESV 178
+++ + F ++ ++P K+PL + + ++ ++L+ + Q RA+ P E V
Sbjct: 497 LQIRNLFSHQILHLQPTKNPLAISLAIRLDHSRVPYHLDL-AIAQRRALHNLRGP--ERV 327
Query: 179 EEINNFMLAMILGEEIEYLNCDTP 202
++ L ILGEEI +L+ P
Sbjct: 326 PPVDYIHLRAILGEEIRFLHGRVP 255
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.343 0.151 0.491
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,671,455
Number of Sequences: 28460
Number of extensions: 93356
Number of successful extensions: 744
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 742
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 743
length of query: 378
length of database: 4,897,600
effective HSP length: 92
effective length of query: 286
effective length of database: 2,279,280
effective search space: 651874080
effective search space used: 651874080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0121b.6


