

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.2
(253 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
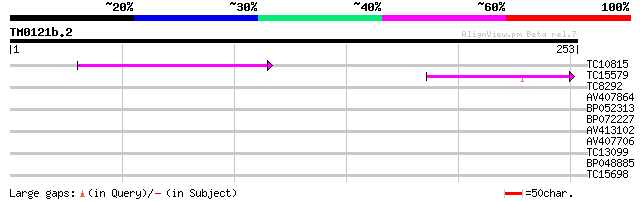
Score E
Sequences producing significant alignments: (bits) Value
TC10815 similar to UP|Q940Y3 (Q940Y3) At2g17400 (At2g17400/At2g1... 45 2e-05
TC15579 weakly similar to UP|Q8L7T9 (Q8L7T9) AT5g54930/MBG8_20, ... 41 2e-04
TC8292 29 0.89
AV407864 28 1.5
BP052313 27 3.4
BP072227 27 4.4
AV413102 26 7.5
AV407706 25 9.8
TC13099 weakly similar to UP|CBL9_ARATH (Q9FJ13) COBRA-like prot... 25 9.8
BP048885 25 9.8
TC15698 25 9.8
>TC10815 similar to UP|Q940Y3 (Q940Y3) At2g17400 (At2g17400/At2g17400),
partial (18%)
Length = 1027
Score = 44.7 bits (104), Expect = 2e-05
Identities = 24/87 (27%), Positives = 40/87 (45%)
Frame = +2
Query: 31 DAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSV 90
+ + F+ L F+ ++ P G L+ L+ V R GY+ V K WR+VG
Sbjct: 533 EREAFMKELENFYRERSLEFKPPKFYGEPLNCLKLWRAVIRLGGYDVVTGSKLWRQVGES 712
Query: 91 FNFSTTTTSASYVLKKHYWNLLYNFEQ 117
F+ T T+ S+ + Y L +E+
Sbjct: 713 FHPPKTCTTVSWTFRIFYEKALLEYEK 793
>TC15579 weakly similar to UP|Q8L7T9 (Q8L7T9) AT5g54930/MBG8_20, partial
(27%)
Length = 592
Score = 41.2 bits (95), Expect = 2e-04
Identities = 25/68 (36%), Positives = 33/68 (47%), Gaps = 2/68 (2%)
Frame = +3
Query: 187 KHANNGPESNVEAKGYPRYGEGRIEEKFDCGYLVSVKLGSE--VLRGVIFHPEETVPPPS 244
+H G E++ + G IE FD GYL+SV++G LRGV+F P VP
Sbjct: 384 QHLQRGQENDSRDALVSQVVSGVIEAVFDAGYLLSVRVGDSDTTLRGVVFKPGRYVPISP 563
Query: 245 SRVVGTGV 252
V GV
Sbjct: 564 ENDVAPGV 587
>TC8292
Length = 851
Score = 28.9 bits (63), Expect = 0.89
Identities = 12/17 (70%), Positives = 14/17 (81%)
Frame = +1
Query: 78 VVAEKKWREVGSVFNFS 94
V+A KKWREVG+VF S
Sbjct: 664 VIA*KKWREVGNVFRIS 714
>AV407864
Length = 414
Score = 28.1 bits (61), Expect = 1.5
Identities = 13/26 (50%), Positives = 21/26 (80%), Gaps = 1/26 (3%)
Frame = +3
Query: 89 SVFNFST-TTTSASYVLKKHYWNLLY 113
S+F+ S+ TT+S+S+ L +H WNLL+
Sbjct: 48 SIFSLSSNTTSSSSFSL*QHPWNLLH 125
>BP052313
Length = 403
Score = 26.9 bits (58), Expect = 3.4
Identities = 15/48 (31%), Positives = 22/48 (45%)
Frame = +3
Query: 87 VGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTV 134
+ + N T+SA L Y L NF++ FF V PP+G +
Sbjct: 120 ISGINNKFKATSSARITLHMLYPGTLRNFQKAIFFSVNFFPQPPSGII 263
>BP072227
Length = 472
Score = 26.6 bits (57), Expect = 4.4
Identities = 8/20 (40%), Positives = 14/20 (70%)
Frame = -2
Query: 109 WNLLYNFEQVHFFKVQGPIT 128
W+L +NF + F++ GP+T
Sbjct: 402 WSLWWNFTSLSAFRIHGPVT 343
>AV413102
Length = 374
Score = 25.8 bits (55), Expect = 7.5
Identities = 9/20 (45%), Positives = 14/20 (70%)
Frame = -3
Query: 138 FTLCFMEMDSLQGMYPNIHA 157
FTLC M S++ +PN+H+
Sbjct: 321 FTLCIFFMLSIRSKFPNLHS 262
>AV407706
Length = 428
Score = 25.4 bits (54), Expect = 9.8
Identities = 9/24 (37%), Positives = 15/24 (62%)
Frame = +1
Query: 21 PLASHEVIVNDAKLFLDTLRQFHF 44
P SHE ++N ++D++ Q HF
Sbjct: 349 PKESHEFLINFIAKYVDSVAQIHF 420
>TC13099 weakly similar to UP|CBL9_ARATH (Q9FJ13) COBRA-like protein 9
precursor, partial (14%)
Length = 426
Score = 25.4 bits (54), Expect = 9.8
Identities = 11/30 (36%), Positives = 20/30 (66%)
Frame = -1
Query: 216 CGYLVSVKLGSEVLRGVIFHPEETVPPPSS 245
C ++ S + + VLR V+ +P +TVP P++
Sbjct: 402 CIWVKSPAVATAVLRSVMGYPAKTVPLPTA 313
>BP048885
Length = 523
Score = 25.4 bits (54), Expect = 9.8
Identities = 12/32 (37%), Positives = 17/32 (52%), Gaps = 5/32 (15%)
Frame = +1
Query: 114 NFEQVHF-----FKVQGPITPPAGTVLNSFTL 140
N Q+HF + QGP+TPP + S T+
Sbjct: 34 NIVQMHFKVKYLIQTQGPVTPPIS*NITSITI 129
>TC15698
Length = 486
Score = 25.4 bits (54), Expect = 9.8
Identities = 10/47 (21%), Positives = 22/47 (46%)
Frame = -3
Query: 108 YWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTLCFMEMDSLQGMYPN 154
+W+L + + +H + + P+ P T+CF + + YP+
Sbjct: 355 FWHLDHKDDGIHVYSLSLPLFTPLSVYYQYSTVCFYS-NQIYASYPS 218
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.137 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,911,934
Number of Sequences: 28460
Number of extensions: 67642
Number of successful extensions: 351
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 351
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 351
length of query: 253
length of database: 4,897,600
effective HSP length: 88
effective length of query: 165
effective length of database: 2,393,120
effective search space: 394864800
effective search space used: 394864800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0121b.2


