

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0118c.6
(278 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
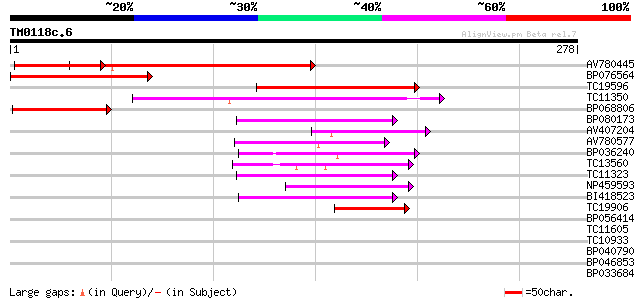
AV780445 182 7e-47
BP076564 75 1e-14
TC19596 70 3e-13
TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3,... 61 2e-10
BP068806 60 3e-10
BP080173 45 2e-05
AV407204 44 4e-05
AV780577 43 5e-05
BP036240 43 7e-05
TC13560 43 7e-05
TC11323 41 2e-04
NP459593 Pge1 protein [Lotus japonicus] 40 4e-04
BI418523 40 6e-04
TC19906 similar to PIR|AC1649|AC1649 phage related protein [impo... 39 7e-04
BP056414 39 0.001
TC11605 38 0.002
TC10933 similar to UP|Q9LNS0 (Q9LNS0) F1L3.4, partial (6%) 37 0.003
BP040790 33 0.003
BP046853 36 0.008
BP033684 35 0.011
>AV780445
Length = 504
Score = 182 bits (461), Expect = 7e-47
Identities = 91/125 (72%), Positives = 97/125 (76%), Gaps = 4/125 (3%)
Frame = +1
Query: 30 ALAWTNFGVGDHGSWIRSQA----RSRNSVRFLAGVWGVWKWRNNMVFEDSPWLVDEACR 85
A AW GV G W SQ R RNSV+ LAGVW VWKWRNNMVFEDSPW VDEA R
Sbjct: 133 AKAW-GMGVD*FGCWRSSQLDQQPRIRNSVKCLAGVW*VWKWRNNMVFEDSPWSVDEAWR 309
Query: 86 RICHEHDEFITFSNGVAGDHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDD 145
++ HEHDEFITFS+G AGD GSWLSSRWQPP+QG VKLN+D S EQDRCMGAGGLIRDD
Sbjct: 310 KLGHEHDEFITFSDGDAGDPGSWLSSRWQPPRQGTVKLNVDGSGREQDRCMGAGGLIRDD 489
Query: 146 QGIWL 150
QG WL
Sbjct: 490 QGKWL 504
Score = 90.1 bits (222), Expect = 4e-19
Identities = 39/45 (86%), Positives = 39/45 (86%)
Frame = +2
Query: 3 RCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDHGSWIRS 47
RC APEEDI HCLRDCPHSQEIWLRLGA AWTN GVGDH SWI S
Sbjct: 65 RCPAPEEDI*HCLRDCPHSQEIWLRLGAWAWTNLGVGDHRSWISS 199
>BP076564
Length = 371
Score = 75.1 bits (183), Expect = 1e-14
Identities = 32/70 (45%), Positives = 45/70 (63%)
Frame = +2
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDHGSWIRSQARSRNSVRFLAG 60
C RCSA +DILHCL+DCPHS+E+W RLG ++ F D SW+ S + N + +
Sbjct: 161 CGRCSAAVKDILHCLQDCPHSREVWNRLGCSSFGGFLSSDGWSWLLSIMKQPNVEKLIYI 340
Query: 61 VWGVWKWRNN 70
+W +W+ RNN
Sbjct: 341 LWWIWR*RNN 370
>TC19596
Length = 499
Score = 70.5 bits (171), Expect = 3e-13
Identities = 32/80 (40%), Positives = 50/80 (62%)
Frame = +1
Query: 122 KLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALLLGLELVWDLG 181
+++ D +F D MG GG++RD G W+ GFYA GG+AL AE AL GL L+W+
Sbjct: 4 RMDSDGAFKHDDDRMGMGGIVRDAHGAWISGFYAGSLGGDALRAEIAALKHGLTLLWNAH 183
Query: 182 YRQVMVEVDCGELLQVMADE 201
R+ EVDC ++++ + ++
Sbjct: 184 VRRATCEVDCLDIVEALEND 243
>TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3, P1 clone:
MIG10, partial (18%)
Length = 1118
Score = 61.2 bits (147), Expect = 2e-10
Identities = 45/162 (27%), Positives = 71/162 (43%), Gaps = 9/162 (5%)
Frame = -3
Query: 61 VWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFITFSNGVAGDHG---------SWLSS 111
+W +WK RN ++F+ +I H + S + S
Sbjct: 978 LWAIWKNRNRVIFQLEEPNPVNTIFQIRHMQQDANLMSTSQSNKQTTRERNINVTSARDR 799
Query: 112 RWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALL 171
W+PP GV KLN D S+ G +IRD+ G+ + G + + +++EA AL
Sbjct: 798 SWRPPPLGVYKLNSDASWKAPSPSCTVGVVIRDNLGLLISGTASRPLAPSPIVSEALALR 619
Query: 172 LGLELVWDLGYRQVMVEVDCGELLQVMADEESCRFLCLS*MS 213
GL L LG +++ E DC L+ E+CR C +*+S
Sbjct: 618 EGLILARSLGLDRILSESDCQPLI------EACRSSCPA*IS 511
>BP068806
Length = 418
Score = 60.5 bits (145), Expect = 3e-10
Identities = 24/49 (48%), Positives = 34/49 (68%)
Frame = +3
Query: 2 PRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDHGSWIRSQAR 50
PRC EDI HCLRDC ++++W+RL AL W F V + +W++SQA+
Sbjct: 252 PRCQHQFEDIWHCLRDCRIARDVWVRLNALGWPGFFVSNAQAWVKSQAQ 398
>BP080173
Length = 382
Score = 44.7 bits (104), Expect = 2e-05
Identities = 22/79 (27%), Positives = 39/79 (48%)
Frame = -2
Query: 112 RWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALL 171
RW P +G +K+N+D SF C GG++ D +G ++ G AE +
Sbjct: 348 RWPVPMEGWIKINIDGSFAPSSNCAACGGVMWDHEGTFITGASVRLGCYFINHAELWGIF 169
Query: 172 LGLELVWDLGYRQVMVEVD 190
G ++ G+++++VE D
Sbjct: 168 HGAKIAQARGFKKILVESD 112
>AV407204
Length = 432
Score = 43.5 bits (101), Expect = 4e-05
Identities = 22/60 (36%), Positives = 32/60 (52%), Gaps = 2/60 (3%)
Frame = -2
Query: 149 WLGGFYAH--RSGGNALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVMADEESCRF 206
W+ GF + + + +AEA AL GL L WD G R+++ DC EL+ +AD F
Sbjct: 428 WVNGFISQYLEAASCSFLAEALALRDGLRLAWDNGTRKLVCNSDCKELIDAIADPSRASF 249
>AV780577
Length = 486
Score = 43.1 bits (100), Expect = 5e-05
Identities = 26/77 (33%), Positives = 41/77 (52%), Gaps = 1/77 (1%)
Frame = +2
Query: 111 SRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWL-GGFYAHRSGGNALIAEATA 169
S W PP G +K+N+ + Q MG G ++RD G+ L G HR + + EA A
Sbjct: 230 STWWPPPHGKLKINIVDVVVPQVDMMGFGFVMRDANGLVLVSGASKHRRSLSVAMGEALA 409
Query: 170 LLLGLELVWDLGYRQVM 186
+++ +DLG+R +M
Sbjct: 410 *RWAMQVSFDLGHRVLM 460
>BP036240
Length = 567
Score = 42.7 bits (99), Expect = 7e-05
Identities = 23/92 (25%), Positives = 45/92 (48%), Gaps = 3/92 (3%)
Frame = -3
Query: 113 WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSG---GNALIAEATA 169
W P +G K ++D + +D GG++RD + W+ GF + N + E +A
Sbjct: 562 WTAPDEGWTKFDVDGAL-SRDLLASCGGVLRDSRRHWIKGFCRNMGDLTYHNVFLVELSA 386
Query: 170 LLLGLELVWDLGYRQVMVEVDCGELLQVMADE 201
+ +E+ + + V+VE D E + +++ E
Sbjct: 385 IQSAVEIALSMDLQHVIVESDALEAIHLLSSE 290
>TC13560
Length = 571
Score = 42.7 bits (99), Expect = 7e-05
Identities = 29/94 (30%), Positives = 45/94 (47%), Gaps = 5/94 (5%)
Frame = +2
Query: 110 SSRWQPPKQGVVKLNMDESFWEQDRCMGAG--GLIRDDQGIWLGGF---YAHRSGGNALI 164
SS W P G VK+N+D + C+ A G++R+ G WL GF + S N +
Sbjct: 137 SSDWIKPDAGWVKINVDGAV---SNCLQASCRGVLRNADGSWLKGFRWNFGTFSSANVFL 307
Query: 165 AEATALLLGLELVWDLGYRQVMVEVDCGELLQVM 198
E A+ +E L +V+VE D E++ +
Sbjct: 308 TELMAVKTAVEAAMSLDLARVIVESDSLEVVDFL 409
>TC11323
Length = 657
Score = 41.2 bits (95), Expect = 2e-04
Identities = 26/79 (32%), Positives = 35/79 (43%)
Frame = +2
Query: 112 RWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALL 171
RW P G KLN D S G GGL+RD ++ GF + + A+L
Sbjct: 2 RWLKPVHGRFKLNFDGSSLGNRGNAGGGGLLRDGSSNFIFGFSIFLAVAQIMKLSLCAIL 181
Query: 172 LGLELVWDLGYRQVMVEVD 190
GL + LGY + +E D
Sbjct: 182 EGLLVCKPLGYDGIGIECD 238
>NP459593 Pge1 protein [Lotus japonicus]
Length = 633
Score = 40.0 bits (92), Expect = 4e-04
Identities = 19/63 (30%), Positives = 31/63 (49%)
Frame = +1
Query: 136 MGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALLLGLELVWDLGYRQVMVEVDCGELL 195
M ++RD +G WL + G+A + E A+ LG+ D+G+ V DC + L
Sbjct: 1 MRCAAVVRDSRGAWLSATLVCYNYGSAFLTEILAVKLGIRHALDIGHIYVFCLSDCSQAL 180
Query: 196 QVM 198
Q +
Sbjct: 181 QAL 189
>BI418523
Length = 496
Score = 39.7 bits (91), Expect = 6e-04
Identities = 22/78 (28%), Positives = 35/78 (44%)
Frame = +1
Query: 113 WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALLL 172
W PP G +K N+D + G GG+ RD LG F + G A A+ ++L
Sbjct: 187 WSPPSAGFIKCNVDGASQGNPGPSGVGGVFRDANRKILGYFSLNSGNGWAYEAKVRSILN 366
Query: 173 GLELVWDLGYRQVMVEVD 190
L + + +++E D
Sbjct: 367 ALVFIQKFLLKNILIESD 420
>TC19906 similar to PIR|AC1649|AC1649 phage related protein [imported] -
Listeria innocua (strain Clip11262)
{Listeria innocua;}, partial (4%)
Length = 509
Score = 39.3 bits (90), Expect = 7e-04
Identities = 19/37 (51%), Positives = 24/37 (64%)
Frame = -3
Query: 160 GNALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQ 196
G+ AE AL LGL L+WD+ R + EVDC EL+Q
Sbjct: 114 GDPFKAEVEALKLGLILMWDVNMRSAVCEVDCIELVQ 4
>BP056414
Length = 606
Score = 38.5 bits (88), Expect = 0.001
Identities = 30/105 (28%), Positives = 48/105 (45%), Gaps = 10/105 (9%)
Frame = -3
Query: 104 DHGSWLSSR-----WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRS 158
D SW SR W P +G K+N+D + D GG++RD G W GF RS
Sbjct: 604 DMFSWGRSRASSYHWTRPPEGWFKVNVDGAL-SGDLQASCGGVVRDAAGSWSKGF--ARS 434
Query: 159 GG-----NALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVM 198
G A E A+ ++ + QV++E D ++++++
Sbjct: 433 LGVLRWHXAFFVELMAVQTAVDFIMSWDIPQVIIESDSQQVVELL 299
>TC11605
Length = 546
Score = 37.7 bits (86), Expect = 0.002
Identities = 22/63 (34%), Positives = 32/63 (49%)
Frame = +3
Query: 113 WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALLL 172
W PP+ ++K N+D S GAGG++R+ + LG F A AE A+LL
Sbjct: 348 WCPPEANILKFNVDGSARGSPGISGAGGILRNAESAVLGRFSKPLGVLCAYQAEVKAILL 527
Query: 173 GLE 175
L+
Sbjct: 528 ALK 536
>TC10933 similar to UP|Q9LNS0 (Q9LNS0) F1L3.4, partial (6%)
Length = 596
Score = 37.4 bits (85), Expect = 0.003
Identities = 24/63 (38%), Positives = 28/63 (44%), Gaps = 4/63 (6%)
Frame = +2
Query: 83 ACRRICH--EHDEFITFS--NGVAGDHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGA 138
+ R CH ++ I F + D S RW PP G VKLN D SF MG
Sbjct: 5 SARAACHYASREKTIEFQPKSFTFADLASL*KDRWTPPSVGWVKLNTDGSFTRGSSLMGC 184
Query: 139 GGL 141
GGL
Sbjct: 185 GGL 193
>BP040790
Length = 344
Score = 33.5 bits (75), Expect(2) = 0.003
Identities = 17/38 (44%), Positives = 23/38 (59%)
Frame = +2
Query: 169 ALLLGLELVWDLGYRQVMVEVDCGELLQVMADEESCRF 206
AL GL VW+ G V +VDC E+LQ + D ++ RF
Sbjct: 176 ALRNGLMSVWNWGLIYVACKVDCVEVLQALYDPDNWRF 289
Score = 22.7 bits (47), Expect(2) = 0.003
Identities = 8/18 (44%), Positives = 10/18 (55%)
Frame = +3
Query: 147 GIWLGGFYAHRSGGNALI 164
G+WL GF GGN +
Sbjct: 108 GLWLLGFALREEGGNTFM 161
>BP046853
Length = 612
Score = 35.8 bits (81), Expect = 0.008
Identities = 22/58 (37%), Positives = 32/58 (54%)
Frame = -3
Query: 139 GGLIRDDQGIWLGGFYAHRSGGNALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQ 196
G LIRD L G +A + L AEA A+ GL L ++LG +V++E D L++
Sbjct: 391 GVLIRDKYSTLLSGTHAKIYCSSPLTAEACAIREGLNLAYNLGCTEVVLESDNKSLVE 218
Score = 26.6 bits (57), Expect = 5.0
Identities = 17/62 (27%), Positives = 27/62 (43%), Gaps = 13/62 (20%)
Frame = -1
Query: 37 GVGDHGSWIR---------SQARSRNSVRFLAGVWGVWKWRNNMVFE----DSPWLVDEA 83
G+G SW+ SQ R + +W +W+ RNN VF+ D + + +A
Sbjct: 609 GMGPCDSWLLQRITILKRLSQDFDREFSLLASLLWSIWRGRNNSVFKAICPDPEFTIQQA 430
Query: 84 CR 85
R
Sbjct: 429 TR 424
>BP033684
Length = 557
Score = 35.4 bits (80), Expect = 0.011
Identities = 26/104 (25%), Positives = 44/104 (42%)
Frame = -1
Query: 67 WRNNMVFEDSPWLVDEACRRICHEHDEFITFSNGVAGDHGSWLSSRWQPPKQGVVKLNMD 126
W V+++ P+ D+ R C + + + G W PP +G++K N+D
Sbjct: 398 WWIKAVWKECPYNFDQFHRDFCMLSFKKLIPKQRLKG---------WTPPSRGILKFNVD 246
Query: 127 ESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATAL 170
+ G GG++ D + + LG F + G A AE L
Sbjct: 245 GASQGNPGPSGVGGVLCDYRSMVLGFFSINMGHGWAYEAEVLYL 114
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.333 0.143 0.509
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,977,971
Number of Sequences: 28460
Number of extensions: 118743
Number of successful extensions: 708
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 702
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 708
length of query: 278
length of database: 4,897,600
effective HSP length: 89
effective length of query: 189
effective length of database: 2,364,660
effective search space: 446920740
effective search space used: 446920740
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0118c.6


