

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0114.10
(1082 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
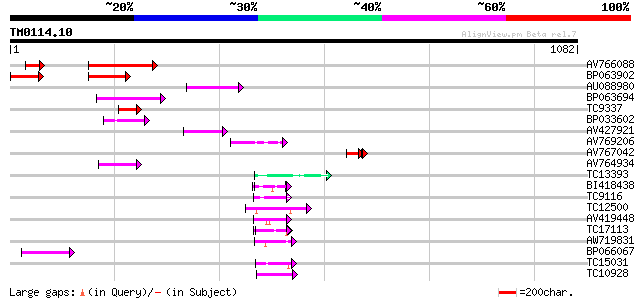
Score E
Sequences producing significant alignments: (bits) Value
AV766088 248 4e-66
BP063902 120 1e-27
AU088980 82 5e-16
BP063694 78 9e-15
TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B pre... 75 7e-14
BP033602 67 2e-11
AV427921 66 3e-11
AV769206 60 1e-09
AV767042 51 6e-08
AV764934 55 8e-08
TC13393 similar to UP|Q9FMU6 (Q9FMU6) Mitochondrial phosphate tr... 53 2e-07
BI418438 53 3e-07
TC9116 similar to UP|Q9ZW85 (Q9ZW85) 3-isopropylmalate dehydrata... 52 7e-07
TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein... 50 2e-06
AV419448 49 6e-06
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 48 7e-06
AW719831 48 7e-06
BP066067 48 9e-06
TC15031 weakly similar to UP|Q69107 (Q69107) LAT protein, partia... 47 1e-05
TC10928 weakly similar to PIR|G96668|G96668 protein F1N19.7 [imp... 47 1e-05
>AV766088
Length = 501
Score = 248 bits (633), Expect = 4e-66
Identities = 119/132 (90%), Positives = 123/132 (93%)
Frame = -3
Query: 150 QTGIPLMRCNSLGDLYPVTPSVPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLNS 209
QTGIPLMRCNS GDLYPVT S PFAG+AQSLWHSRLGHP SSAL+ LRSNKFI+YE LN
Sbjct: 397 QTGIPLMRCNSPGDLYPVTLSFPFAGIAQSLWHSRLGHPSSSALRYLRSNKFISYELLNY 218
Query: 210 PTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFL 269
VCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHR+YVLFLDDFTDFL
Sbjct: 217 SPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRFYVLFLDDFTDFL 38
Query: 270 WTFPLSNKSQVF 281
WTFPLSNKSQVF
Sbjct: 37 WTFPLSNKSQVF 2
Score = 68.9 bits (167), Expect = 4e-12
Identities = 32/35 (91%), Positives = 34/35 (96%)
Frame = -2
Query: 31 VPTDIATAMHTMTLAPPDSTWYMDTGASSHVASSP 65
VPTD+ATAMHTMTLAPPDST YMDTGASSHVA+SP
Sbjct: 500 VPTDLATAMHTMTLAPPDST*YMDTGASSHVAASP 396
>BP063902
Length = 494
Score = 120 bits (301), Expect = 1e-27
Identities = 57/80 (71%), Positives = 64/80 (79%)
Frame = -3
Query: 150 QTGIPLMRCNSLGDLYPVTPSVPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLNS 209
+TG+PL+RCNSLGDLYPVT S PFAGLA S+WH RLGHP SSAL LR+NK I E S
Sbjct: 273 KTGMPLLRCNSLGDLYPVTRSSPFAGLASSVWHHRLGHPASSALNHLRNNKLIFCEPSRS 94
Query: 210 PTVCESCVFGKHVRLPFVSS 229
+VC+SCV GKHVRLPF SS
Sbjct: 93 SSVCDSCVLGKHVRLPFSSS 34
Score = 102 bits (254), Expect = 3e-22
Identities = 45/64 (70%), Positives = 53/64 (82%)
Frame = -2
Query: 1 TSQWTRPSAPPRQPGILGPRPQAHAASTSPVPTDIATAMHTMTLAPPDSTWYMDTGASSH 60
TSQ +RP P+QPG+LGPRPQA+ A+ SP PTDIA AMHTM+L PPD+ WYMDTGASSH
Sbjct: 466 TSQXSRPMGVPQQPGVLGPRPQAYTATASPAPTDIAAAMHTMSLTPPDTPWYMDTGASSH 287
Query: 61 VASS 64
A+S
Sbjct: 286 TAAS 275
>AU088980
Length = 360
Score = 82.0 bits (201), Expect = 5e-16
Identities = 43/110 (39%), Positives = 57/110 (51%)
Frame = +2
Query: 337 NGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRR 396
N ERK R I N+ R L+ H+ +P + W A++ A YL+N +P L P Q L
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFLNND 181
Query: 397 DPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDL 446
P+ L+VFG LCY + KL PR+ CVFLG+ GY DL
Sbjct: 182 SPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGYIVMDL 331
>BP063694
Length = 511
Score = 77.8 bits (190), Expect = 9e-15
Identities = 45/132 (34%), Positives = 62/132 (46%), Gaps = 2/132 (1%)
Frame = -2
Query: 167 VTPSVPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEH-LNSPTVCESCVFGKHVRLP 225
+T S P + LWHSRLGH ++ L + + L P C SC K RL
Sbjct: 399 LTTSSPLPRASFELWHSRLGHVNFDIIKQLHKHGCLDVSSILPKPICCTSCQMAKSKRLV 220
Query: 226 FVSSNNVTVMPFDILHSDL-WTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMF 284
F +N D++H DL SPV S G Y+V+F+DDF+ F W +PL KS ++
Sbjct: 219 FHDNNKRASAVLDLIHCDLRGPSPVASIDGFSYFVIFVDDFSRFTWFYPLKRKSDFSDVL 40
Query: 285 TSLTHQIRTHFS 296
+ FS
Sbjct: 39 LRFKVFMENRFS 4
>TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B precursor
(Pectin methylesterase) (PE) , partial (54%)
Length = 1145
Score = 74.7 bits (182), Expect = 7e-14
Identities = 33/43 (76%), Positives = 37/43 (85%)
Frame = +3
Query: 209 SPTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLS 251
S +VC+SCV GKHVRLPF SS +T+ PFDILHSDLWTSPVLS
Sbjct: 1017 SNSVCDSCVLGKHVRLPFSSSETITLRPFDILHSDLWTSPVLS 1145
>BP033602
Length = 533
Score = 66.6 bits (161), Expect = 2e-11
Identities = 35/89 (39%), Positives = 53/89 (59%), Gaps = 2/89 (2%)
Frame = +3
Query: 180 LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-CESCVFGKHVRLPFVSSNNVTVMPFD 238
LWH+RLGHP L+ L + F +++N ++ CESCV K+ R P+ S PF
Sbjct: 276 LWHNRLGHPNFQYLRHLFPDLF---KNVNCSSLECESCVLAKNQRAPYYSQPYHASRPFY 446
Query: 239 ILHSDLW-TSPVLSSAGHRYYVLFLDDFT 266
++HSD+W S + + G R++V F+DD T
Sbjct: 447 LIHSDVWGPSKITTQFGKRWFVTFIDDHT 533
>AV427921
Length = 284
Score = 65.9 bits (159), Expect = 3e-11
Identities = 33/84 (39%), Positives = 48/84 (56%)
Frame = +1
Query: 332 HTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQ 391
+T QNG +ERK RTI NM+R+LL + +P SF A+ + ++LN P + N+ P +
Sbjct: 19 YTPQQNGVSERKNRTILNMVRSLLTMSGLPKSFLPEAVMWSLHILNRSPTLVVQNMMPEE 198
Query: 392 LLYRRDPSYTHLRVFGCLCYPLVP 415
R P+ H R+ CL Y VP
Sbjct: 199 AWSGRQPAVDHFRISRCLAYAHVP 270
>AV769206
Length = 536
Score = 60.5 bits (145), Expect = 1e-09
Identities = 42/109 (38%), Positives = 56/109 (50%)
Frame = -3
Query: 421 KLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFATMPTPPPSTYDCF 480
KL S CVF+GY ++H+GYKC D S R I IS+ V+F E RFP+ ++ P T
Sbjct: 510 KLSLHSKECVFIGYSINHKGYKCLDQSGR-IYISKDVLFHEHRFPYPSL--FPDETTSST 340
Query: 481 SDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSP 529
S P S P + PPA+ S S+ SA+ PSS +P
Sbjct: 339 SAEFFP------LSTIPIVSHTTPPALASSGF---SSLGSATLPSSFNP 220
>AV767042
Length = 444
Score = 50.8 bits (120), Expect(2) = 6e-08
Identities = 25/33 (75%), Positives = 27/33 (81%)
Frame = -3
Query: 643 QIAGVDCDETFSPVVKPATIRTVLTIALSGLGP 675
Q AGVDC++TFSPVVKPA IRTV TIALS P
Sbjct: 349 QTAGVDCNKTFSPVVKPAPIRTVFTIALSRSWP 251
Score = 23.9 bits (50), Expect(2) = 6e-08
Identities = 10/12 (83%), Positives = 11/12 (91%)
Frame = -1
Query: 672 GLGPFTS*MFRM 683
GLGPF S*MFR+
Sbjct: 261 GLGPFPS*MFRI 226
>AV764934
Length = 406
Score = 54.7 bits (130), Expect = 8e-08
Identities = 32/84 (38%), Positives = 47/84 (55%), Gaps = 1/84 (1%)
Frame = -2
Query: 169 PSVPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKHVRLPFVS 228
PS A + S WHSRLGHP L+ + S+ ++Y ++ +C GK RLP
Sbjct: 249 PSSVSAFVTLSTWHSRLGHPHLDNLKRVLSSCNVSYSPKDTVEFRTACCLGKAHRLPSQM 70
Query: 229 SNNVTVMPFDILHSDLW-TSPVLS 251
S T +PF++++SDLW +PV S
Sbjct: 69 S-TTTYLPFELIYSDLWGPAPVCS 1
>TC13393 similar to UP|Q9FMU6 (Q9FMU6) Mitochondrial phosphate translocator
(AT5g14040/MUA22_4), partial (8%)
Length = 541
Score = 53.1 bits (126), Expect = 2e-07
Identities = 44/150 (29%), Positives = 57/150 (37%), Gaps = 3/150 (2%)
Frame = +1
Query: 467 ATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSS 526
+T P PPP+ + +SP SP SV PP+ P T T S A+P +S
Sbjct: 73 STFPNPPPT---------------RTSSPSSSPHSVPPPSTPSKSFPITPTGSLAAPAAS 207
Query: 527 PSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLA---LSDPN 583
S SPP P P + PR + PT S PK P ++ L PN
Sbjct: 208 SSRTSPPSPAPSLSP----------NPRS------SSSSPTSSASPKTPMISSPGLRSPN 339
Query: 584 WKSAMQSEFDALIRNKTWDLVPRPSDVNII 613
S S A PSD+ I+
Sbjct: 340 TDSTTPSSMQA-----------SPSDLTIL 396
>BI418438
Length = 431
Score = 52.8 bits (125), Expect = 3e-07
Identities = 30/82 (36%), Positives = 42/82 (50%), Gaps = 9/82 (10%)
Frame = +1
Query: 463 RFPFATMPTPPPSTYDCFSD-PIH-PSIIHQWTSPDPSP-------TSVVPPAVTPSQLP 513
++P P P PS Y FS P+ PS+ +SP P P T++ PP +P+ P
Sbjct: 94 KYPLNPPPPPSPSQYPPFSPCPLSSPSLFSSSSSPSPPPPTATTTTTTIPPPPPSPTPPP 273
Query: 514 TTSTSSSASPPSSPSPPSPPQP 535
+S A PP+SPS +PP P
Sbjct: 274 RQKSSKPARPPASPSNANPPSP 339
Score = 45.1 bits (105), Expect = 6e-05
Identities = 27/70 (38%), Positives = 36/70 (50%)
Frame = +1
Query: 468 TMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSP 527
T+P PPPS P + P SP++ PP+ P+ L T SSS++PPS P
Sbjct: 235 TIPPPPPSP-----TPPPRQKSSKPARPPASPSNANPPS--PTSLLTQPLSSSSNPPSPP 393
Query: 528 SPPSPPQPVP 537
PP+ P P P
Sbjct: 394 PPPTSPPPNP 423
>TC9116 similar to UP|Q9ZW85 (Q9ZW85) 3-isopropylmalate dehydratase, small
subunit, partial (25%)
Length = 426
Score = 51.6 bits (122), Expect = 7e-07
Identities = 34/76 (44%), Positives = 40/76 (51%), Gaps = 2/76 (2%)
Frame = +1
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPS-PTSVVPPAVTPSQLPT-TSTSSSAS 522
P +P PPP+ HPS +S S PT PP+++PS LPT TST SSA
Sbjct: 187 PTLLLPPPPPNPQP------HPSTASATSSATTSTPTRSSPPSISPSSLPTPTSTRSSAP 348
Query: 523 PPSSPSPPSPPQPVPP 538
PSS SPP P PP
Sbjct: 349 TPSSASPP----PTPP 384
>TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein {Danio
rerio;} , partial (5%)
Length = 678
Score = 50.1 bits (118), Expect = 2e-06
Identities = 38/138 (27%), Positives = 56/138 (40%), Gaps = 13/138 (9%)
Frame = +1
Query: 451 IIISRHVIFDETRFPFAT----MPTPPPSTYDCFSDPIHPSIIHQWT--SPDPSPTSVVP 504
+I ++ E FP + P PPP Y + P P ++ T +P P P++ +
Sbjct: 250 VIAGSAIMSGEASFPCSLPSPPQPPPPPPKYGVTTIPPSPPLLMSATDRAPLPPPSTHLT 429
Query: 505 PAVTPSQLPTTSTSSSASPPSSPSPPSPPQ-------PVPPIRTMATRSMRGIYKPRKLF 557
TPS + +T + PS P PPSPP P PP+ + L
Sbjct: 430 SLTTPSPPSSRTTITQTPLPSPPQPPSPPPKYGVATIPPPPLPMSSIDRTPLPLLSTPLT 609
Query: 558 NLSVTIDDPTISPLPKNP 575
+L PT+SP P P
Sbjct: 610 SLDAPPPTPTLSPPPPPP 663
Score = 40.4 bits (93), Expect = 0.002
Identities = 28/87 (32%), Positives = 35/87 (40%), Gaps = 2/87 (2%)
Frame = +1
Query: 454 SRHVIFDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLP 513
S +T P P PP Y + P P P P S + TP L
Sbjct: 454 SSRTTITQTPLPSPPQPPSPPPKYGVATIP-----------PPPLPMSSIDR--TPLPLL 594
Query: 514 TTSTSSSASPPSSP--SPPSPPQPVPP 538
+T +S +PP +P SPP PP P PP
Sbjct: 595 STPLTSLDAPPPTPTLSPPPPPPPTPP 675
Score = 38.9 bits (89), Expect = 0.004
Identities = 39/147 (26%), Positives = 52/147 (34%), Gaps = 32/147 (21%)
Frame = +1
Query: 470 PTPPPSTYDCFSDPIHPSI------IHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSAS- 522
P PPPS I + + SP P P PP++ P+ P S + ++
Sbjct: 91 PPPPPSAPFIAGSAIISKVSKSVGDVSLPCSPSPPPPQPPPPSLPPTTPPPASVIAGSAI 270
Query: 523 -------PPSSPSPPSPP-----------QPVPPIRTMATRSMRGIYKPRKLFNLSVTID 564
P S PSPP PP P PP+ AT R P S+T
Sbjct: 271 MSGEASFPCSLPSPPQPPPPPPKYGVTTIPPSPPLLMSATD--RAPLPPPSTHLTSLTTP 444
Query: 565 DP-------TISPLPKNPKLALSDPNW 584
P T +PLP P+ P +
Sbjct: 445 SPPSSRTTITQTPLPSPPQPPSPPPKY 525
>AV419448
Length = 382
Score = 48.5 bits (114), Expect = 6e-06
Identities = 32/88 (36%), Positives = 41/88 (46%), Gaps = 14/88 (15%)
Frame = +2
Query: 465 PFATMPTPPPSTYDCFSDPIHPSI-----IHQWTS--------PDPSPTSVVPPAVTPSQ 511
P ++ P P PS S P+HP WTS P +P++ PP
Sbjct: 32 PSSSHPPPSPSQPSLPSPPLHPPTPPSPTASSWTSASAPPPSSPTATPSAPTPPLSAALS 211
Query: 512 LPTTSTSS-SASPPSSPSPPSPPQPVPP 538
L +T+TSS S SP S+PS PP P PP
Sbjct: 212 LASTATSSPSPSPISNPSASLPPTPPPP 295
Score = 29.6 bits (65), Expect = 2.7
Identities = 25/75 (33%), Positives = 35/75 (46%), Gaps = 8/75 (10%)
Frame = +2
Query: 467 ATMPTPPPSTYDCFSDPIHPSIIHQWT---SPDPSPTS----VVPPAVTPSQLPT-TSTS 518
A P+ P +T + P+ ++ T SP PSP S +PP P T +STS
Sbjct: 143 APPPSSPTATPSAPTPPLSAALSLASTATSSPSPSPISNPSASLPPTPPPPPTKTPSSTS 322
Query: 519 SSASPPSSPSPPSPP 533
SS + SSP+ P
Sbjct: 323 SSPAATSSPAARVDP 367
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 48.1 bits (113), Expect = 7e-06
Identities = 27/75 (36%), Positives = 32/75 (42%)
Frame = +1
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPP 524
P P+PPP D F P P + P PSP ++P P P + PP
Sbjct: 505 PNPFQPSPPPLLPDPFQPPSPPPLFPNPFQP-PSPPPLIPNPFQPPPSPPPFIPNPFQPP 681
Query: 525 SSPSPPSPPQPVPPI 539
S PPSP P PPI
Sbjct: 682 PSKQPPSPLFPFPPI 726
Score = 42.0 bits (97), Expect = 5e-04
Identities = 26/75 (34%), Positives = 32/75 (42%), Gaps = 6/75 (8%)
Frame = +1
Query: 470 PTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSP-- 527
P+PPP + F P P +I P PSP +P P Q P + S P P
Sbjct: 559 PSPPPLFPNPFQPPSPPPLIPNPFQPPPSPPPFIP---NPFQPPPSKQPPSPLFPFPPIV 729
Query: 528 ----SPPSPPQPVPP 538
+PP PP P PP
Sbjct: 730 IPGLTPPPPPPPPPP 774
Score = 34.7 bits (78), Expect = 0.083
Identities = 28/77 (36%), Positives = 30/77 (38%), Gaps = 1/77 (1%)
Frame = +1
Query: 465 PFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPP 524
PF P+PPP + F P PS PSP PP V P P PP
Sbjct: 625 PFQPPPSPPPFIPNPFQPP--PS------KQPPSPLFPFPPIVIPGLTPPPPP-----PP 765
Query: 525 SSPS-PPSPPQPVPPIR 540
P PP PP PP R
Sbjct: 766 PPPFFPPFPPLFPPPGR 816
Score = 32.7 bits (73), Expect = 0.31
Identities = 26/80 (32%), Positives = 33/80 (40%), Gaps = 4/80 (5%)
Frame = +1
Query: 464 FPFATMPTP--PPSTYDCFSDPIHPSIIHQWTSPD--PSPTSVVPPAVTPSQLPTTSTSS 519
FP +P P PP + F P P I + + P P+P PP + P S
Sbjct: 400 FPPPLIPNPFQPPLVPNPFQPP--PLIPNPFQPPPLIPNPFQPSPPPLLPDPFQPPSP-- 567
Query: 520 SASPPSSPSPPSPPQPVPPI 539
PP P+P PP P P I
Sbjct: 568 ---PPLFPNPFQPPSPPPLI 618
>AW719831
Length = 436
Score = 48.1 bits (113), Expect = 7e-06
Identities = 32/84 (38%), Positives = 40/84 (47%), Gaps = 5/84 (5%)
Frame = +1
Query: 468 TMPTPPPSTYDCFSDPIHP-----SIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSAS 522
T T PPST P P S P+PT+ P + + ST+SS+S
Sbjct: 190 TTTTTPPSTNPHLLLPPPPHARSSSCAATAAKSSPAPTTTTSPTSPATPRSSPSTASSSS 369
Query: 523 PPSSPSPPSPPQPVPPIRTMATRS 546
PPSSPS S P P+PPI +T S
Sbjct: 370 PPSSPS--SLPSPIPPISASSTNS 435
>BP066067
Length = 505
Score = 47.8 bits (112), Expect = 9e-06
Identities = 31/104 (29%), Positives = 47/104 (44%), Gaps = 1/104 (0%)
Frame = -2
Query: 22 QAHAASTSPVPTDIATAMHTMTLAPPDSTWYMDTGASSHVASSPGNLTSYSNLSHLNQKL 81
Q H P +A +T+ +W D+GAS HV + P +L + Q +
Sbjct: 384 QQHLQLPQSSPAPMAFVSNTVAAP*TTHSW*ADSGASHHVTNDPQHLMESITMPGQEQ-V 208
Query: 82 YVGSGQGIPIQGSGDTTLPTS-HKHKPLTLSHVLHTPQIVKNLI 124
YV SGQG+PI G T+ + + +L H P I KN++
Sbjct: 207 YVCSGQGLPILSIGFTSFSSPLISNTKFSLKDFFHLPTITKNIV 76
>TC15031 weakly similar to UP|Q69107 (Q69107) LAT protein, partial (27%)
Length = 997
Score = 47.4 bits (111), Expect = 1e-05
Identities = 30/83 (36%), Positives = 42/83 (50%), Gaps = 6/83 (7%)
Frame = +3
Query: 470 PTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSP 529
P+ PPS+ FS + S ++P+PSP S A PS ++S+SSS P+S
Sbjct: 492 PSSPPSSSSLFSPSVSSSTFV--SNPNPSPISKASTASGPSVSSSSSSSSSGPSPNSSGY 665
Query: 530 P------SPPQPVPPIRTMATRS 546
P SPP P+ PI +T S
Sbjct: 666 PSSAAGTSPPSPIKPISAESTSS 734
>TC10928 weakly similar to PIR|G96668|G96668 protein F1N19.7 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 590
Score = 47.4 bits (111), Expect = 1e-05
Identities = 24/78 (30%), Positives = 41/78 (51%), Gaps = 1/78 (1%)
Frame = -3
Query: 472 PPPSTYDCFSDPIHPSIIHQWTSP-DPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPP 530
P + ++ S P+ P + H + P DP+ ++ P P+ P+ + S++A PP P PP
Sbjct: 426 PHHTHHNHISLPLSPHVSHSRSDPTDPTHSAPPTPPTRPNTSPSQTPSAAAPPPPPPPPP 247
Query: 531 SPPQPVPPIRTMATRSMR 548
+ P P+ R+ A R R
Sbjct: 246 AAPSPLSETRSGAPRHAR 193
Score = 36.6 bits (83), Expect = 0.022
Identities = 31/96 (32%), Positives = 41/96 (42%), Gaps = 7/96 (7%)
Frame = -3
Query: 447 SHRKIIISRHVIF---DETRFPFATMPTPP--PSTYDCFSDPIHPSIIHQWTSPDPSPTS 501
+H + +S HV D T + PTPP P+T PS +P P P
Sbjct: 408 NHISLPLSPHVSHSRSDPTDPTHSAPPTPPTRPNT--------SPSQTPSAAAPPPPPP- 256
Query: 502 VVPPAVTPSQLPTTSTSSS--ASPPSSPSPPSPPQP 535
PP PS L T + + A PP +PP+PP P
Sbjct: 255 --PPPAAPSPLSETRSGAPRHARPPPLYAPPAPPSP 154
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.329 0.140 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,387,219
Number of Sequences: 28460
Number of extensions: 487917
Number of successful extensions: 14781
Number of sequences better than 10.0: 987
Number of HSP's better than 10.0 without gapping: 6413
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 10154
length of query: 1082
length of database: 4,897,600
effective HSP length: 100
effective length of query: 982
effective length of database: 2,051,600
effective search space: 2014671200
effective search space used: 2014671200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0114.10


