

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
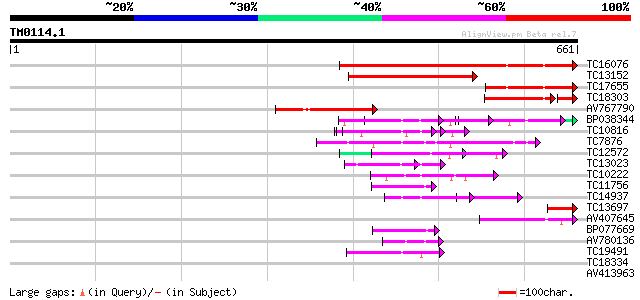
Query= TM0114.1
(661 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG... 308 2e-84
TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial... 154 6e-38
TC17655 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG... 131 3e-31
TC18303 similar to GB|AAG32022.1|11141605|AF277458 LEUNIG {Arabi... 103 4e-26
AV767790 104 5e-23
BP038344 70 1e-12
TC10816 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subun... 60 1e-09
TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-bi... 53 2e-07
TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory pro... 51 7e-07
TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragm... 47 7e-06
TC10222 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Ar... 45 5e-05
TC11756 weakly similar to GB|AAH01494.1|16306637|BC001494 U5 snR... 44 6e-05
TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5,... 43 1e-04
TC13697 similar to GB|AAG32022.1|11141605|AF277458 LEUNIG {Arabi... 43 1e-04
AV407645 42 2e-04
BP077669 41 5e-04
AV780136 41 5e-04
TC19491 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Ar... 41 7e-04
TC18334 weakly similar to UP|Q9LHN3 (Q9LHN3) Emb|CAB63739.1 (AT3... 40 0.001
AV413963 37 0.013
>TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG
{Arabidopsis thaliana;} , partial (24%)
Length = 1048
Score = 308 bits (788), Expect = 2e-84
Identities = 144/277 (51%), Positives = 200/277 (71%)
Frame = +3
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V CHFSSDGK++ASAG +KKV +WNM+ Q ++ + H +I+DVRF+P S+ AT+S
Sbjct: 3 VTCCHFSSDGKMLASAGDDKKVVLWNMDTLQTESTPEQHKSVISDVRFRPNSSQLATASI 182
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D+SVRLWDA+NP + GH +MSLDFHP++ D+ C DS + IR WN+ S C
Sbjct: 183 DKSVRLWDAANPTYCVQEYNGHSSAIMSLDFHPKKTDIFCFCDSANEIRYWNITSSSCTR 362
Query: 505 TTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFL 564
++GGS QVRFQP+ GQ+L+ A ++I DVE+D ++ L+GH + V SICWD +G+FL
Sbjct: 363 VSKGGSSQVRFQPRIGQVLSAASD*VVSIFDVESDRQIYTLQGHPEPVNSICWDVNGDFL 542
Query: 565 ASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGS 624
ASVS + +IWS S G+CI EL S GN+F SC+FHP Y +LL+IGG+ SLE W+ + +
Sbjct: 543 ASVSPNLVKIWSLTS-GECIQELSSSGNQFYSCVFHPSYSTLLVIGGFSSLELWNMAD-N 716
Query: 625 KTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
K+ +I+AH+ +I+ LA SPV ++ASASHDNSVK+WK
Sbjct: 717 KSMTISAHESVISALAQSPVTGMVASASHDNSVKLWK 827
>TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial (60%)
Length = 453
Score = 154 bits (388), Expect = 6e-38
Identities = 70/150 (46%), Positives = 100/150 (66%)
Frame = +2
Query: 396 VMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASN 455
++AS GH+KK +W ++ + A+ + HS LITDVRF P ATSSFD++V++WD N
Sbjct: 2 LLASGGHDKKAVLWYTDSLKQKATLEEHSALITDVRFSPSMPRLATSSFDKTVKVWDVDN 181
Query: 456 PIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRF 515
P SL TGH VMSLDFHP + DL+CS DS+ IR W++N C ++GG+ ++RF
Sbjct: 182 PGYSLRTFTGHSASVMSLDFHPNKEDLICSCDSDGEIRYWSINNGSCARVSKGGTTKMRF 361
Query: 516 QPQYGQLLATAIGNSITIIDVETDSHLHDL 545
QP+ G+ LA A N ++I+DVET + + L
Sbjct: 362 QPRLGRYLAAAAENIVSILDVETXACRYSL 451
>TC17655 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG
{Arabidopsis thaliana;} , partial (9%)
Length = 741
Score = 131 bits (330), Expect = 3e-31
Identities = 60/107 (56%), Positives = 82/107 (76%)
Frame = +2
Query: 555 ICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQS 614
ICWD +G+FLAS+S+DS ++WS A+ G CI EL+S GN F SC+FHP Y +LL++GGYQS
Sbjct: 2 ICWDTNGDFLASLSQDSVKVWSLAT-GDCIHELNSSGNMFHSCVFHPSYSTLLVVGGYQS 178
Query: 615 LEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
LE W+ E +K +I AH+ +I+ LA SP+ ++AS SHD SVK+WK
Sbjct: 179 LELWNMAE-NKCMTIPAHECVISSLAQSPLTGMVASTSHDKSVKIWK 316
Score = 32.3 bits (72), Expect = 0.25
Identities = 21/95 (22%), Positives = 41/95 (43%), Gaps = 4/95 (4%)
Frame = +2
Query: 485 SSDSNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSH 541
+S S D +++W++ +C+H F P Y LL S+ + ++ ++
Sbjct: 32 ASLSQDSVKVWSLATGDCIHELNSSGNMFHSCVFHPSYSTLLVVGGYQSLELWNM-AENK 208
Query: 542 LHDLKGHDKDVLSICWDRSGNFLASVSED-SARIW 575
+ H+ + S+ +AS S D S +IW
Sbjct: 209 CMTIPAHECVISSLAQSPLTGMVASTSHDKSVKIW 313
>TC18303 similar to GB|AAG32022.1|11141605|AF277458 LEUNIG {Arabidopsis
thaliana;} , partial (12%)
Length = 571
Score = 103 bits (258), Expect(2) = 4e-26
Identities = 49/85 (57%), Positives = 63/85 (73%), Gaps = 2/85 (2%)
Frame = +1
Query: 554 SICWDRSGNFLASVSEDSARIWS--AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGG 611
S+CWD SG LASVSEDS R+W+ SDG+C+ EL G+KF SC+FHP Y SLL+IG
Sbjct: 10 SVCWDPSGELLASVSEDSVRVWNLGTGSDGECVHELSCNGSKFHSCVFHPTYPSLLVIGC 189
Query: 612 YQSLEAWSPTEGSKTWSIAAHQGLI 636
YQS+E W+ E +KT +++AH GLI
Sbjct: 190 YQSMELWNIAE-NKTMTLSAHDGLI 261
Score = 31.6 bits (70), Expect(2) = 4e-26
Identities = 14/23 (60%), Positives = 17/23 (73%)
Frame = +3
Query: 639 LADSPVGELIASASHDNSVKVWK 661
LA S V L+ASASHD +K+WK
Sbjct: 270 LAVSTVNGLVASASHDKFIKLWK 338
>AV767790
Length = 617
Score = 104 bits (259), Expect = 5e-23
Identities = 54/119 (45%), Positives = 77/119 (64%)
Frame = +3
Query: 311 ITESTRPVESAQDCADAKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGF 370
+T ST ++ + D D+NVESFLS +D D R + F LKR + + + +K F
Sbjct: 270 LTSSTNQLDDMEHFGDVGSLDDNVESFLSQDD--GDGR-DLFGTLKRNPSEHATDASKAF 440
Query: 371 SLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITD 429
S E+G + KS +KV+ CHFSSDGK++ASAGH++KV +WNME Q ++ D HS +ITD
Sbjct: 441 SFSEVGSIRKSNSKVVCCHFSSDGKLLASAGHDQKVVLWNMETLQTESTPDAHSLIITD 617
>BP038344
Length = 525
Score = 69.7 bits (169), Expect = 1e-12
Identities = 39/130 (30%), Positives = 66/130 (50%), Gaps = 8/130 (6%)
Frame = +2
Query: 384 KVLTCH--------FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPG 435
K LT H FS+DG ++ASA +K + IW+ HS I+D+ +
Sbjct: 80 KTLTAHDAAVACVKFSNDGTLLASASLDKTLIIWSSATLSLLHRLTGHSEGISDLAWSSD 259
Query: 436 STMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLW 495
S ++S DR++R+WDA+ + L GH V ++F+P + + + S ++ +R+W
Sbjct: 260 SHYICSASDDRTLRIWDATGG-DCVKTLRGHTHAVFCVNFNP-QSNYIVSGSFDETVRVW 433
Query: 496 NVNQSECMHT 505
V C+HT
Sbjct: 434 EVKTGRCIHT 463
Score = 67.8 bits (164), Expect = 5e-12
Identities = 35/154 (22%), Positives = 77/154 (49%), Gaps = 3/154 (1%)
Frame = +2
Query: 414 FQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSL 473
+++ + H + V+F T+ A++S D+++ +W ++ + L LTGH E + L
Sbjct: 68 YRHLKTLTAHDAAVACVKFSNDGTLLASASLDKTLIIWSSAT-LSLLHRLTGHSEGISDL 244
Query: 474 DFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQ---VRFQPQYGQLLATAIGNS 530
+ +CS+ + +R+W+ +C+ T RG + V F PQ +++ + +
Sbjct: 245 AWSSDS-HYICSASDDRTLRIWDATGGDCVKTLRGHTHAVFCVNFNPQSNYIVSGSFDET 421
Query: 531 ITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFL 564
+ + +V+T +H + H V S+ ++R G +
Sbjct: 422 VRVWEVKTGRCIHTIIAHTMPVTSVHFNRDGTLI 523
Score = 45.4 bits (106), Expect = 3e-05
Identities = 37/133 (27%), Positives = 62/133 (45%), Gaps = 4/133 (3%)
Frame = +2
Query: 520 GQLLATA-IGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSA 577
G LLA+A + ++ I T S LH L GH + + + W +++ S S+D + RIW
Sbjct: 134 GTLLASASLDKTLIIWSSATLSLLHRLTGHSEGISDLAWSSDSHYICSASDDRTLRIWD- 310
Query: 578 ASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGY--QSLEAWSPTEGSKTWSIAAHQGL 635
A+ G C+ L + F+P +S I+ G +++ W G +I AH
Sbjct: 311 ATGGDCVKTLRGHTHAVFCVNFNP--QSNYIVSGSFDETVRVWEVKTGRCIHTIIAHTMP 484
Query: 636 IAGLADSPVGELI 648
+ + + G LI
Sbjct: 485 VTSVHFNRDGTLI 523
Score = 42.0 bits (97), Expect = 3e-04
Identities = 36/142 (25%), Positives = 56/142 (39%), Gaps = 4/142 (2%)
Frame = +2
Query: 524 ATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSAR-IWSAASDG- 581
A A+ NS + HL L HD V + + G LAS S D IWS+A+
Sbjct: 23 AAAMANSSDEPPYKPYRHLKTLTAHDAAVACVKFSNDGTLLASASLDKTLIIWSSATLSL 202
Query: 582 --KCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGL 639
+ G + + S H + ++L W T G ++ H + +
Sbjct: 203 LHRLTGHSEGISDLAWSSDSH----YICSASDDRTLRIWDATGGDCVKTLRGHTHAVFCV 370
Query: 640 ADSPVGELIASASHDNSVKVWK 661
+P I S S D +V+VW+
Sbjct: 371 NFNPQSNYIVSGSFDETVRVWE 436
>TC10816 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subunit-like
protein, partial (20%)
Length = 979
Score = 60.1 bits (144), Expect = 1e-09
Identities = 31/109 (28%), Positives = 57/109 (51%)
Frame = +1
Query: 389 HFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSV 448
HF + S G + K+ +WN + + + H I V+F S ++S D+++
Sbjct: 412 HFHHSQPLFVSGGDDYKIKVWNYKLHRCLFTLLGHLDYIRTVQFHHESPWIVSASDDQTI 591
Query: 449 RLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNV 497
R+W+ + + +LTGH+ VM FHPR DL+ S+ + +R+W++
Sbjct: 592 RIWNWQSR-TCISVLTGHNHYVMCALFHPRE-DLVVSASLDQTVRVWDI 732
Score = 51.2 bits (121), Expect = 5e-07
Identities = 34/155 (21%), Positives = 65/155 (41%), Gaps = 28/155 (18%)
Frame = +1
Query: 382 INKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFAT 441
++ + T F + + SA ++ + IWN ++ + H+H + F P + +
Sbjct: 517 LDYIRTVQFHHESPWIVSASDDQTIRIWNWQSRTCISVLTGHNHYVMCALFHPREDLVVS 696
Query: 442 SSFDRSVRLWDASNPIRSLG--------------------------ILTGHDEQVMSLDF 475
+S D++VR+WD + R +L GHD V F
Sbjct: 697 ASLDQTVRVWDIGSLKRKNASPADDILRLSQMNTDLFGGVDAVVKYVLEGHDRGVNWASF 876
Query: 476 HPRRMDLLCSSDSNDVIRLWNVNQSEC--MHTTRG 508
HP + L+ S+ + ++LW +N ++ + T RG
Sbjct: 877 HP-ALPLIVSAADDRQVKLWRMNDTKAWEVDTLRG 978
Score = 48.5 bits (114), Expect = 3e-06
Identities = 35/172 (20%), Positives = 71/172 (40%), Gaps = 14/172 (8%)
Frame = +1
Query: 379 HKSINKVLTCHFSSDGKVMASAGHEKKVFI-----------WNMENFQYFASADTHSHLI 427
H+ K+LT + +V + H K+ +I W+ D H +
Sbjct: 223 HQQSKKMLTKFETKSNRVKGLSFHPKRPWILASLHSGVIQLWDYRMGTLIDKFDEHDGPV 402
Query: 428 TDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSD 487
V F +F + D +++W+ R L L GH + + ++ FH ++ +SD
Sbjct: 403 RGVHFHHSQPLFVSGGDDYKIKVWNYKLH-RCLFTLLGHLDYIRTVQFHHESPWIVSASD 579
Query: 488 SNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDV 536
+ IR+WN C+ G + V F P+ +++ ++ ++ + D+
Sbjct: 580 -DQTIRIWNWQSRTCISVLTGHNHYVMCALFHPREDLVVSASLDQTVRVWDI 732
Score = 33.5 bits (75), Expect = 0.11
Identities = 30/151 (19%), Positives = 60/151 (38%), Gaps = 1/151 (0%)
Frame = +1
Query: 511 KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
K + F P+ +LA+ I + D + + HD V + + S S +D
Sbjct: 277 KGLSFHPKRPWILASLHSGVIQLWDYRMGTLIDKFDEHDGPVRGVHFHHSQPLFVSGGDD 456
Query: 571 -SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSI 629
++W+ +C+ L + ++ FH ++ Q++ W+ + +
Sbjct: 457 YKIKVWNYKLH-RCLFTLLGHLDYIRTVQFHHESPWIVSASDDQTIRIWNWQSRTCISVL 633
Query: 630 AAHQGLIAGLADSPVGELIASASHDNSVKVW 660
H + P +L+ SAS D +V+VW
Sbjct: 634 TGHNHYVMCALFHPREDLVVSASLDQTVRVW 726
>TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1379
Score = 52.8 bits (125), Expect = 2e-07
Identities = 62/275 (22%), Positives = 114/275 (40%), Gaps = 14/275 (5%)
Frame = +3
Query: 358 ISATCSRNENKGFSLQEIGCLHKS-INKVLTCHFSSDGKVMASAGHEKKVFIWNM--ENF 414
+S T R +G L+ H + + T +SD ++ +A +K + +W++ E+
Sbjct: 48 VSQTEKRKMAEGLVLRGTMRAHTDQVTAIATPIDNSD--MIVTASRDKSIILWHLTKEDK 221
Query: 415 QYFASADT---HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVM 471
Y HSH + DV + S+D +RLWD + S GH + V+
Sbjct: 222 TYGVPRRRLTGHSHFVQDVVLSSDGQFALSGSWDGELRLWDLAAG-TSARRFVGHTKDVL 398
Query: 472 SLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQ------VRFQPQYGQ--LL 523
S+ F ++ S+ + I+LWN EC +T + VRF P Q ++
Sbjct: 399 SVAFSIDNRQIV-SASRDRTIKLWN-TLGECKYTIQDNDAHSDWVSCVRFSPSTLQPTIV 572
Query: 524 ATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGKC 583
+ + ++ + ++ + L GH V ++ G+ AS +D + ++GK
Sbjct: 573 SASWDRTVKVWNLTNCKLRNTLAGHSGYVNTVAVSPDGSLCASGGKDGVILLWDLAEGKR 752
Query: 584 IGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAW 618
+ L G+ + F P R L S++ W
Sbjct: 753 LYSL-DAGSIIHALCFSPN-RYWLCAATETSIKIW 851
Score = 32.0 bits (71), Expect = 0.33
Identities = 48/231 (20%), Positives = 79/231 (33%), Gaps = 6/231 (2%)
Frame = +3
Query: 436 STMFATSSFDRSVRLWDASNPIRSLGI----LTGHDEQVMSLDFHPRRMDLLCSSDSNDV 491
S M T+S D+S+ LW + ++ G+ LTGH V D++ SSD
Sbjct: 153 SDMIVTASRDKSIILWHLTKEDKTYGVPRRRLTGHSHFVQ---------DVVLSSD---- 293
Query: 492 IRLWNVNQSECMHTTRGGSKQVRFQPQYGQL-LATAIGNSITIIDVETDSHLHDLKGHDK 550
GQ L+ + + + D+ + GH K
Sbjct: 294 ----------------------------GQFALSGSWDGELRLWDLAAGTSARRFVGHTK 389
Query: 551 DVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLII 609
DVLS+ + + S S D + ++W+ + K + + + + SC+
Sbjct: 390 DVLSVAFSIDNRQIVSASRDRTIKLWNTLGECKYTIQDNDAHSDWVSCV----------- 536
Query: 610 GGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+SP+ T I SAS D +VKVW
Sbjct: 537 -------RFSPSTLQPT---------------------IVSASWDRTVKVW 605
Score = 27.3 bits (59), Expect = 8.0
Identities = 25/122 (20%), Positives = 42/122 (33%), Gaps = 6/122 (4%)
Frame = +3
Query: 545 LKGHDKDVLSICW--DRSGNFLASVSEDSARIWSAASDGKCIG----ELHSVGNKFQSCI 598
++ H V +I D S + + + S +W + K G L + Q +
Sbjct: 102 MRAHTDQVTAIATPIDNSDMIVTASRDKSIILWHLTKEDKTYGVPRRRLTGHSHFVQDVV 281
Query: 599 FHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVK 658
+ L L W G+ H + +A S I SAS D ++K
Sbjct: 282 LSSDGQFALSGSWDGELRLWDLAAGTSARRFVGHTKDVLSVAFSIDNRQIVSASRDRTIK 461
Query: 659 VW 660
+W
Sbjct: 462 LW 467
>TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory protein tup1
homolog - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (51%)
Length = 770
Score = 50.8 bits (120), Expect = 7e-07
Identities = 41/167 (24%), Positives = 77/167 (45%), Gaps = 9/167 (5%)
Frame = +3
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
HS +T V F ++ +SS+D R+WDAS ++ + V + F P +
Sbjct: 36 HSDPVTAVDFNRDGSLIVSSSYDGLCRIWDASTGHCMKTLIDDENPPVSFVKFSPNAKFI 215
Query: 483 LCSSDSNDVIRLWNVNQSECMHTTRG--GSK---QVRFQPQYGQ-LLATAIGNSITIIDV 536
L + N+ +RLWN + + T G SK F G+ ++ + + I + ++
Sbjct: 216 LVGTLDNN-LRLWNYSTGRFLKTYTGHVNSKYCISSTFSTTNGKYVVGGSEDHGIYLWEL 392
Query: 537 ETDSHLHDLKGHDKDVLSICWDRSGNFLAS---VSEDSARIWSAASD 580
+T + L+GH V+S+ + N +AS ++ + +IW+ D
Sbjct: 393 QTRKIVQKLEGHSDTVVSVSCHPTENMIASGALGNDKTVKIWTQQKD 533
Score = 42.4 bits (98), Expect = 2e-04
Identities = 34/155 (21%), Positives = 62/155 (39%), Gaps = 6/155 (3%)
Frame = +3
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASA-DTHSHLITDVRFQPGSTMFATSS 443
V F+ DG ++ S+ ++ IW+ + D + ++ V+F P + +
Sbjct: 48 VTAVDFNRDGSLIVSSSYDGLCRIWDASTGHCMKTLIDDENPPVSFVKFSPNAKFILVGT 227
Query: 444 FDRSVRLWDASNPIRSLGILTGH--DEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSE 501
D ++RLW+ S R L TGH + +S F + + I LW + +
Sbjct: 228 LDNNLRLWNYSTG-RFLKTYTGHVNSKYCISSTFSTTNGKYVVGGSEDHGIYLWELQTRK 404
Query: 502 CMHTTRGGSK---QVRFQPQYGQLLATAIGNSITI 533
+ G S V P + + A+GN T+
Sbjct: 405 IVQKLEGHSDTVVSVSCHPTENMIASGALGNDKTV 509
>TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragment),
partial (18%)
Length = 591
Score = 47.4 bits (111), Expect = 7e-06
Identities = 31/118 (26%), Positives = 55/118 (46%)
Frame = +1
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
SSDG +A A E + I + N ++ S +T + P + +S R +R+
Sbjct: 187 SSDGSFIACACGES-IKIVDSANASIRSTLQGDSESVTALALSPDDNLLFSSGHSRQIRV 363
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRG 508
WD S ++ + GH+ VM + HP LL + ++ + +W+V+ C H +G
Sbjct: 364 WDLST-LKCVRSWKGHEGPVMCMSCHPSG-GLLATGGADRKVLVWDVDGGYCTHFFKG 531
Score = 42.4 bits (98), Expect = 2e-04
Identities = 22/87 (25%), Positives = 37/87 (42%)
Frame = +1
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
S D ++ S+GH +++ +W++ + S H + + P + AT DR V +
Sbjct: 310 SPDDNLLFSSGHSRQIRVWDLSTLKCVRSWKGHEGPVMCMSCHPSGGLLATGGADRKVLV 489
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHP 477
WD GH V + FHP
Sbjct: 490 WDVDGGY-CTHFFKGHGGVVSCVMFHP 567
Score = 31.2 bits (69), Expect = 0.56
Identities = 24/100 (24%), Positives = 37/100 (37%), Gaps = 3/100 (3%)
Frame = +1
Query: 505 TTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG 561
T +G S+ V P L ++ I + D+ T + KGH+ V+ + SG
Sbjct: 268 TLQGDSESVTALALSPDDNLLFSSGHSRQIRVWDLSTLKCVRSWKGHEGPVMCMSCHPSG 447
Query: 562 NFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHP 601
LA+ D + G C G +FHP
Sbjct: 448 GLLATGGADRKVLVWDVDGGYCTHFFKGHGGVVSCVMFHP 567
Score = 29.6 bits (65), Expect = 1.6
Identities = 22/100 (22%), Positives = 38/100 (38%)
Frame = +1
Query: 561 GNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSP 620
G+F+A +S +I +A+ L + P L G + + W
Sbjct: 196 GSFIACACGESIKIVDSAN-ASIRSTLQGDSESVTALALSPDDNLLFSSGHSRQIRVWDL 372
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ S H+G + ++ P G L+A+ D V VW
Sbjct: 373 STLKCVRSWKGHEGPVMCMSCHPSGGLLATGGADRKVLVW 492
>TC10222 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;}, partial (55%)
Length = 557
Score = 44.7 bits (104), Expect = 5e-05
Identities = 47/173 (27%), Positives = 71/173 (40%), Gaps = 23/173 (13%)
Frame = +1
Query: 421 DTHSHLITDVRFQPGST----MFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFH 476
+ H + V + P + + T S D +VRLW + + TGH V S+ H
Sbjct: 43 NAHDDSVWAVTWAPATANRPPLLLTGSLDETVRLWRSDELVLER-TNTGHCLGVASVAAH 219
Query: 477 PRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQV---RFQPQYGQLLATAIGNS--- 530
P + SS + +R+++V+ + + T +V RF P+ G LLA A G S
Sbjct: 220 PLG-SIAASSSLDSFVRVFDVDSNATLATLEAPPSEVWQMRFDPK-GALLAVAGGGSASV 393
Query: 531 -------------ITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
++I E S D G K VLS+ W G +A S D
Sbjct: 394 KLWDTSTWELVATLSIPRPEGGSKPTDKSGSKKFVLSVAWSPDGKRIACGSMD 552
>TC11756 weakly similar to GB|AAH01494.1|16306637|BC001494 U5 snRNP-specific
40 kDa protein (hPrp8-binding) {Homo sapiens;} , partial
(43%)
Length = 719
Score = 44.3 bits (103), Expect = 6e-05
Identities = 20/75 (26%), Positives = 39/75 (51%)
Frame = +1
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDL 482
H +I ++F P T+ A+ S DR + LW+ ++ +L GH V+ L + +
Sbjct: 301 HQSVIYTMKFNPAGTVIASGSHDREIFLWNVHGECKNFMVLKGHKNAVLDLHWTTDGTQI 480
Query: 483 LCSSDSNDVIRLWNV 497
+ S+ + +R+W+V
Sbjct: 481 V-SASPDKTVRVWDV 522
Score = 39.3 bits (90), Expect = 0.002
Identities = 35/141 (24%), Positives = 66/141 (45%), Gaps = 7/141 (4%)
Frame = +1
Query: 462 ILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH--TTRGGSKQV---RFQ 516
+LTGH + ++ F+P + S ++ LWNV+ EC + +G V +
Sbjct: 289 LLTGHQSVIYTMKFNPAGTVIASGSHDREIF-LWNVH-GECKNFMVLKGHKNAVLDLHWT 462
Query: 517 PQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG-NFLASVSED-SARI 574
Q+++ + ++ + DVET + + H V S C R G + S S+D +A++
Sbjct: 463 TDGTQIVSASPDKTVRVWDVETGKQVKKMVEHLSYVNSCCPSRRGPPLVVSGSDDGTAKL 642
Query: 575 WSAASDGKCIGELHSVGNKFQ 595
W D + G + + +K+Q
Sbjct: 643 W----DMRQRGSIQTFPDKYQ 693
Score = 36.6 bits (83), Expect = 0.013
Identities = 20/71 (28%), Positives = 37/71 (51%), Gaps = 1/71 (1%)
Frame = +1
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTH-SHLITDVRFQPGSTMFAT 441
N VL H+++DG + SA +K V +W++E + H S++ + + G + +
Sbjct: 436 NAVLDLHWTTDGTQIVSASPDKTVRVWDVETGKQVKKMVEHLSYVNSCCPSRRGPPLVVS 615
Query: 442 SSFDRSVRLWD 452
S D + +LWD
Sbjct: 616 GSDDGTAKLWD 648
Score = 31.6 bits (70), Expect = 0.43
Identities = 12/32 (37%), Positives = 19/32 (58%)
Frame = +1
Query: 629 IAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ HQ +I + +P G +IAS SHD + +W
Sbjct: 292 LTGHQSVIYTMKFNPAGTVIASGSHDREIFLW 387
>TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5, TAC
clone:K8K14 (AT5g67320/K8K14_4), partial (19%)
Length = 1089
Score = 43.1 bits (100), Expect = 1e-04
Identities = 28/107 (26%), Positives = 55/107 (51%), Gaps = 2/107 (1%)
Frame = +1
Query: 438 MFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNV 497
+ A++SFD +V+LWD + + L GH + V S+ F P + L S + I +W++
Sbjct: 52 VLASASFDSTVKLWDVEVG-KLIYSLNGHRDGVYSVAFSPNG-EYLASGSPDKSIHIWSL 225
Query: 498 NQSECM--HTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHL 542
+ + + +T GG +V + + ++ A N++ ++D S L
Sbjct: 226 KEGKIVKTYTGSGGIFEVCWNKEGDKIAACFANNTVCVLDFRM*SSL 366
Score = 41.2 bits (95), Expect = 5e-04
Identities = 24/77 (31%), Positives = 39/77 (50%), Gaps = 1/77 (1%)
Frame = +1
Query: 522 LLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASD 580
L + + +++ + DVE ++ L GH V S+ + +G +LAS S D S IWS +
Sbjct: 55 LASASFDSTVKLWDVEVGKLIYSLNGHRDGVYSVAFSPNGEYLASGSPDKSIHIWS-LKE 231
Query: 581 GKCIGELHSVGNKFQSC 597
GK + G F+ C
Sbjct: 232 GKIVKTYTGSGGIFEVC 282
Score = 36.6 bits (83), Expect = 0.013
Identities = 17/56 (30%), Positives = 29/56 (51%)
Frame = +1
Query: 396 VMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLW 451
V+ASA + V +W++E + S + H + V F P A+ S D+S+ +W
Sbjct: 52 VLASASFDSTVKLWDVEVGKLIYSLNGHRDGVYSVAFSPNGEYLASGSPDKSIHIW 219
>TC13697 similar to GB|AAG32022.1|11141605|AF277458 LEUNIG {Arabidopsis
thaliana;} , partial (4%)
Length = 453
Score = 43.1 bits (100), Expect = 1e-04
Identities = 19/34 (55%), Positives = 25/34 (72%)
Frame = +2
Query: 628 SIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+++AH GLI LA S V L+ASASHD +K+WK
Sbjct: 2 TLSAHDGLITALAVSTVNGLVASASHDKFIKLWK 103
>AV407645
Length = 413
Score = 42.4 bits (98), Expect = 2e-04
Identities = 33/119 (27%), Positives = 54/119 (44%), Gaps = 5/119 (4%)
Frame = +2
Query: 548 HDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNK-FQSCIFHPGYRS 605
HD +V + + +G +LAS S D +A IW +G + S K F S + P +
Sbjct: 5 HDDEVWLVQFSHNGKYLASASNDRTAIIWEVGINGLSLKHRLSGHQKAFSSISWSPNDQE 184
Query: 606 LLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADS---PVGELIASASHDNSVKVWK 661
LL G +++ W + G + ++ AGL P G+ I S D S+ +W+
Sbjct: 185 LLTCGVEEAIRRWDVSTGK---CLQIYEKAGAGLISCTWFPSGKYILSGLSDKSICMWE 352
Score = 35.8 bits (81), Expect = 0.023
Identities = 28/92 (30%), Positives = 40/92 (43%), Gaps = 4/92 (4%)
Frame = +2
Query: 520 GQLLATAIGNSITII---DVETDSHLHDLKGHDKDVLSICWDRSGNFLASVS-EDSARIW 575
G+ LA+A + II + S H L GH K SI W + L + E++ R W
Sbjct: 44 GKYLASASNDRTAIIWEVGINGLSLKHRLSGHQKAFSSISWSPNDQELLTCGVEEAIRRW 223
Query: 576 SAASDGKCIGELHSVGNKFQSCIFHPGYRSLL 607
S GKC+ G SC + P + +L
Sbjct: 224 D-VSTGKCLQIYEKAGAGLISCTWFPSGKYIL 316
Score = 35.8 bits (81), Expect = 0.023
Identities = 27/121 (22%), Positives = 50/121 (41%), Gaps = 2/121 (1%)
Frame = +2
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIW--NMENFQYFASADTHSHLITDVRFQPGSTMFA 440
++V FS +GK +ASA +++ IW + H + + + P
Sbjct: 11 DEVWLVQFSHNGKYLASASNDRTAIIWEVGINGLSLKHRLSGHQKAFSSISWSPNDQELL 190
Query: 441 TSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQS 500
T + ++R WD S + L I ++S + P +L S S+ I +W ++
Sbjct: 191 TCGVEEAIRRWDVSTG-KCLQIYEKAGAGLISCTWFPSGKYIL-SGLSDKSICMWELDGK 364
Query: 501 E 501
E
Sbjct: 365 E 367
>BP077669
Length = 488
Score = 41.2 bits (95), Expect = 5e-04
Identities = 24/78 (30%), Positives = 38/78 (47%)
Frame = -2
Query: 424 SHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLL 483
+H I + P T AT S D +VRLWD ++ R L GH + V ++ L+
Sbjct: 319 NHYILKCKIAPNCTQLATCSSDSTVRLWDLTSQCRLQKTLNGHKKWVWDCTYNSDSSYLV 140
Query: 484 CSSDSNDVIRLWNVNQSE 501
+S S+ +LW+ + E
Sbjct: 139 TAS-SDLTAKLWDCARGE 89
>AV780136
Length = 558
Score = 41.2 bits (95), Expect = 5e-04
Identities = 21/71 (29%), Positives = 38/71 (52%)
Frame = -1
Query: 435 GSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRL 494
G T+ + ++ +R+WD + ++L L GH + + +L L S S+ +IRL
Sbjct: 540 GGTVLVSGGTEKVIRVWDPRSGSKTLK-LRGHADNIRALLLDSTGRYCL-SGSSDSMIRL 367
Query: 495 WNVNQSECMHT 505
W++ Q C+HT
Sbjct: 366 WDIGQPRCLHT 334
Score = 36.2 bits (82), Expect = 0.017
Identities = 17/57 (29%), Positives = 27/57 (46%), Gaps = 1/57 (1%)
Frame = -1
Query: 605 SLLIIGGYQS-LEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++L+ GG + + W P GSKT + H I L G S S D+ +++W
Sbjct: 534 TVLVSGGTEKVIRVWDPRSGSKTLKLRGHADNIRALLLDSTGRYCLSGSSDSMIRLW 364
>TC19491 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;}, partial (39%)
Length = 527
Score = 40.8 bits (94), Expect = 7e-04
Identities = 31/120 (25%), Positives = 54/120 (44%), Gaps = 5/120 (4%)
Frame = +1
Query: 393 DGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWD 452
D +++ +A + V +++ E + H+ + V P AT S DR+VRLWD
Sbjct: 52 DPRLLFTASDDGNVHMYDAEGKALVGTMSGHASWVLCVDVSPDGGAIATGSSDRTVRLWD 231
Query: 453 ASNPIRSLGILTGHDEQVMSLDFHPR-----RMDLLCSSDSNDVIRLWNVNQSECMHTTR 507
+ S+ ++ H +QV + F P R L S + I L++ + S H T+
Sbjct: 232 LAMR-ASVQTMSNHTDQVWGVAFRPPGGTDVRGGRLASVSDDKSISLYDYS*S*FRHCTQ 408
Score = 32.0 bits (71), Expect = 0.33
Identities = 22/86 (25%), Positives = 40/86 (45%), Gaps = 6/86 (6%)
Frame = +1
Query: 375 IGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQ- 433
+G + + VL S DG +A+ ++ V +W++ + H+ + V F+
Sbjct: 124 VGTMSGHASWVLCVDVSPDGGAIATGSSDRTVRLWDLAMRASVQTMSNHTDQVWGVAFRP 303
Query: 434 PGST-----MFATSSFDRSVRLWDAS 454
PG T A+ S D+S+ L+D S
Sbjct: 304 PGGTDVRGGRLASVSDDKSISLYDYS 381
>TC18334 weakly similar to UP|Q9LHN3 (Q9LHN3) Emb|CAB63739.1
(AT3g18860/MCB22_3), partial (21%)
Length = 595
Score = 40.4 bits (93), Expect = 0.001
Identities = 38/139 (27%), Positives = 57/139 (40%), Gaps = 7/139 (5%)
Frame = +2
Query: 382 INKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVR--FQPGSTMF 439
+N F S GH+ K +Y + H H DVR G+
Sbjct: 44 LNSAFVRSFLFSSSRSLSQGHDFK---------EYQLRCELHGHE-DDVRGICVCGNDGI 193
Query: 440 ATSSFDRSVRLWDASN-PIRSLGILTGHDEQVMSLDFHPRRMDL----LCSSDSNDVIRL 494
ATSS D++VR W N S +L GH V L + P +L + S + ++ +
Sbjct: 194 ATSSRDKTVRFWSPENRKFVSSKVLVGHSSFVGPLVWIPPNPELPQGGVASGGMDTLVLV 373
Query: 495 WNVNQSECMHTTRGGSKQV 513
W+++ E HT +G QV
Sbjct: 374 WDLSTGEKAHTLKGHQLQV 430
Score = 32.7 bits (73), Expect = 0.19
Identities = 33/128 (25%), Positives = 54/128 (41%), Gaps = 10/128 (7%)
Frame = +2
Query: 544 DLKGHDKDVLSICWDRSGNFLASVSEDSARIWSA-----ASDGKCIGELHSVGNKFQSCI 598
+L GH+ DV IC + S + + R WS S +G VG +
Sbjct: 134 ELHGHEDDVRGICVCGNDGIATSSRDKTVRFWSPENRKFVSSKVLVGHSSFVG----PLV 301
Query: 599 FHPGYRSL----LIIGGYQSLE-AWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASH 653
+ P L + GG +L W + G K ++ HQ + +A G+++ SAS
Sbjct: 302 WIPPNPELPQGGVASGGMDTLVLVWDLSTGEKAHTLKGHQLQVTSIAFDD-GDVV-SASV 475
Query: 654 DNSVKVWK 661
D +++ WK
Sbjct: 476 DCTLRRWK 499
>AV413963
Length = 313
Score = 36.6 bits (83), Expect = 0.013
Identities = 19/78 (24%), Positives = 37/78 (47%), Gaps = 4/78 (5%)
Frame = +2
Query: 423 HSHLITDVRFQPGSTMFATSSFDRSVRLWDASN----PIRSLGILTGHDEQVMSLDFHPR 478
H + V + T A+ S D++ R+W ++ + L GH + V L + P+
Sbjct: 74 HKKKVHSVAWNCIGTKLASGSVDQTARIWHVEQHGHGKVKDIE-LKGHTDSVDQLCWDPK 250
Query: 479 RMDLLCSSDSNDVIRLWN 496
DL+ ++ + +RLW+
Sbjct: 251 HADLIATASGDKTVRLWD 304
Score = 33.5 bits (75), Expect = 0.11
Identities = 18/42 (42%), Positives = 22/42 (51%), Gaps = 1/42 (2%)
Frame = +2
Query: 541 HLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDG 581
H + GH K V S+ W+ G LAS S D +ARIW G
Sbjct: 53 HSREYSGHKKKVHSVAWNCIGTKLASGSVDQTARIWHVEQHG 178
Score = 31.6 bits (70), Expect = 0.43
Identities = 15/38 (39%), Positives = 23/38 (60%), Gaps = 2/38 (5%)
Frame = +2
Query: 544 DLKGHDKDVLSICWD-RSGNFLASVSED-SARIWSAAS 579
+LKGH V +CWD + + +A+ S D + R+W A S
Sbjct: 200 ELKGHTDSVDQLCWDPKHADLIATASGDKTVRLWDARS 313
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.130 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,080,667
Number of Sequences: 28460
Number of extensions: 179696
Number of successful extensions: 1111
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 1014
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1088
length of query: 661
length of database: 4,897,600
effective HSP length: 96
effective length of query: 565
effective length of database: 2,165,440
effective search space: 1223473600
effective search space used: 1223473600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0114.1


