

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0113.7
(147 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
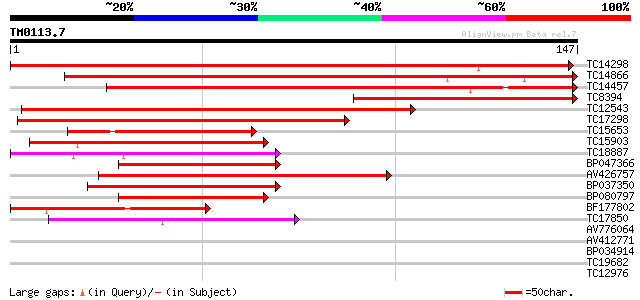
TC14298 homologue to UP|Q8L5W2 (Q8L5W2) bZIP transcription facto... 197 5e-52
TC14866 similar to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor ... 137 8e-34
TC14457 similar to UP|Q8GTZ1 (Q8GTZ1) bZIP transcription factor,... 122 3e-29
TC8394 similar to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor A... 117 8e-28
TC12543 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor,... 82 4e-17
TC17298 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor,... 78 5e-16
TC15653 similar to UP|Q941L5 (Q941L5) bZIP transcription factor ... 59 3e-10
TC15903 similar to UP|Q40724 (Q40724) Transcriptional activator ... 55 5e-09
TC18887 similar to UP|Q94KA6 (Q94KA6) bZIP transcription factor ... 49 3e-07
BP047366 49 5e-07
AV426757 49 5e-07
BP037350 46 3e-06
BP080797 41 7e-05
BF177802 39 3e-04
TC17850 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor,... 39 5e-04
AV776064 36 0.003
AV412771 30 0.13
BP034914 28 0.84
TC19682 similar to UP|O65169 (O65169) Glutaredoxin I, partial (54%) 26 3.2
TC12976 similar to UP|O02448 (O02448) Lox22-otx (Fragment), part... 26 3.2
>TC14298 homologue to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor ATB2,
partial (62%)
Length = 1585
Score = 197 bits (502), Expect = 5e-52
Identities = 105/152 (69%), Positives = 126/152 (82%), Gaps = 6/152 (3%)
Frame = +3
Query: 1 MASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQV 60
MASSSGTSSGSS++QNSGSEE+LQA+MDQRKRKRMISNRESARRSRMRKQKHLD+LA+QV
Sbjct: 612 MASSSGTSSGSSLLQNSGSEEDLQAVMDQRKRKRMISNRESARRSRMRKQKHLDDLASQV 791
Query: 61 AQLRSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFVNATNGVFA 120
QLR ENQQ++TS+N+TTQ+++ ++AENSVL AQMGELS+RLESLNEII+ +NA + F
Sbjct: 792 TQLRKENQQIITSVNITTQQYLSVEAENSVLRAQMGELSNRLESLNEIIEVLNANSAAFG 971
Query: 121 ------ANSFVDPIPSAYYLNHPIMASADILQ 146
+N F S YLN IM SAD+LQ
Sbjct: 972 TTFTEPSNGFFFNPLSMSYLNQQIMPSADMLQ 1067
Score = 25.8 bits (55), Expect = 3.2
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = -2
Query: 2 ASSSGTSSGSSMIQNSGSEEELQALMDQRKR 32
+SSSG S +S +QN SE E+ M + K+
Sbjct: 708 SSSSGPSQPASPLQNQSSEAEMTLKMFRLKK 616
>TC14866 similar to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor ATB2,
partial (60%)
Length = 948
Score = 137 bits (345), Expect = 8e-34
Identities = 74/141 (52%), Positives = 98/141 (69%), Gaps = 8/141 (5%)
Frame = +3
Query: 15 QNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSL 74
Q+SGSE +LQ +MDQRK KR SNRESARRSRMRKQ HL++L +QV QL EN ++LT++
Sbjct: 12 QSSGSEGDLQMVMDQRKNKRKQSNRESARRSRMRKQSHLEDLTSQVTQLTKENGEILTNI 191
Query: 75 NLTTQRFMIIDAENSVLSAQMGELSHRLESLNEIIDFV-------NATNGVFAANSFVDP 127
N+T+Q++ ++ ENS+L AQMGELS RL+SLN+II+ + T G + F
Sbjct: 192 NITSQQYQNVETENSILRAQMGELSQRLQSLNDIINVIKTSAAATTTTTGSYNEREFYQT 371
Query: 128 IPSAY-YLNHPIMASADILQY 147
+ YLN PI ASADI Q+
Sbjct: 372 GALNFSYLNQPITASADIFQW 434
>TC14457 similar to UP|Q8GTZ1 (Q8GTZ1) bZIP transcription factor, partial
(50%)
Length = 645
Score = 122 bits (305), Expect = 3e-29
Identities = 65/131 (49%), Positives = 94/131 (71%), Gaps = 9/131 (6%)
Frame = +1
Query: 26 LMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMIID 85
+MDQ+KRKRM SNRESARRSRMRKQ+HL+ ++AQV QL+ EN Q+ T++ +TTQ ++ ++
Sbjct: 7 VMDQKKRKRMQSNRESARRSRMRKQEHLEGMSAQVEQLKKENNQISTNIGVTTQMYLNVE 186
Query: 86 AENSVLSAQMGELSHRLESLNEIIDFVNATNGV---------FAANSFVDPIPSAYYLNH 136
AEN++L QM ELS+RL+SLNEII ++ ++N F F+D ++ +N
Sbjct: 187 AENAILRVQMAELSNRLQSLNEIIHYIESSNNYLFHEAQETQFNDCGFMDTW-NSLAVNQ 363
Query: 137 PIMASADILQY 147
IMA+AD+L Y
Sbjct: 364 SIMAAADMLMY 396
>TC8394 similar to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor ATB2,
partial (25%)
Length = 569
Score = 117 bits (293), Expect = 8e-28
Identities = 58/58 (100%), Positives = 58/58 (100%)
Frame = +3
Query: 90 VLSAQMGELSHRLESLNEIIDFVNATNGVFAANSFVDPIPSAYYLNHPIMASADILQY 147
VLSAQMGELSHRLESLNEIIDFVNATNGVFAANSFVDPIPSAYYLNHPIMASADILQY
Sbjct: 3 VLSAQMGELSHRLESLNEIIDFVNATNGVFAANSFVDPIPSAYYLNHPIMASADILQY 176
>TC12543 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor, partial
(52%)
Length = 510
Score = 82.0 bits (201), Expect = 4e-17
Identities = 46/102 (45%), Positives = 66/102 (64%)
Frame = +3
Query: 4 SSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQL 63
S +S SS + ++E+ Q L+++RK +RMISNRESARRSRMRKQ+HLDEL +QV L
Sbjct: 159 SPQSSCISSNSTSDEADEQQQGLINERKHRRMISNRESARRSRMRKQRHLDELWSQVVWL 338
Query: 64 RSENQQMLTSLNLTTQRFMIIDAENSVLSAQMGELSHRLESL 105
R+EN Q+L L+ ++ + EN+ L + EL L +
Sbjct: 339 RNENHQLLDKLSHASESHDQVVQENAQLKEEAIELRQMLRDM 464
>TC17298 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor, partial
(39%)
Length = 484
Score = 78.2 bits (191), Expect = 5e-16
Identities = 42/86 (48%), Positives = 58/86 (66%)
Frame = +1
Query: 3 SSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQ 62
SSS SS S+ + +++ ++++RK +RMISNRESARRSRMRKQKHLDEL + V
Sbjct: 226 SSSCISSNSTSDEAEDQQQQQSLIINERKHRRMISNRESARRSRMRKQKHLDELWSMVVC 405
Query: 63 LRSENQQMLTSLNLTTQRFMIIDAEN 88
LR+EN Q+L LN ++ + EN
Sbjct: 406 LRNENHQLLEKLNHLSESHDRVVQEN 483
>TC15653 similar to UP|Q941L5 (Q941L5) bZIP transcription factor BZI-4,
partial (25%)
Length = 511
Score = 58.9 bits (141), Expect = 3e-10
Identities = 31/49 (63%), Positives = 41/49 (83%)
Frame = +2
Query: 16 NSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLR 64
+SGSE A+ D RKRKRM+SNRESARRSRMRKQK L++L A++++L+
Sbjct: 365 SSGSEGGDLAI-DDRKRKRMVSNRESARRSRMRKQKQLEDLTAEMSRLQ 508
>TC15903 similar to UP|Q40724 (Q40724) Transcriptional activator protein
(RITA-1 protein), partial (21%)
Length = 603
Score = 55.1 bits (131), Expect = 5e-09
Identities = 33/67 (49%), Positives = 42/67 (62%), Gaps = 5/67 (7%)
Frame = +1
Query: 6 GTSSGSSMIQN-----SGSEEELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQV 60
GTSSGSS + +G E+ D ++ +R +SNRESARRSR RKQ HL EL QV
Sbjct: 379 GTSSGSSDRSDEDDDEAGPCEQSNNPQDVKRLRRKVSNRESARRSRRRKQAHLAELETQV 558
Query: 61 AQLRSEN 67
+L+ EN
Sbjct: 559 EKLKLEN 579
>TC18887 similar to UP|Q94KA6 (Q94KA6) bZIP transcription factor 6, partial
(64%)
Length = 944
Score = 49.3 bits (116), Expect = 3e-07
Identities = 29/76 (38%), Positives = 45/76 (59%), Gaps = 6/76 (7%)
Frame = +3
Query: 1 MASSSGTSSGSSMIQ---NSGSEEELQALMD---QRKRKRMISNRESARRSRMRKQKHLD 54
M S+G +S S + N G ++ +R+++RMI NRESA RSR RKQ +
Sbjct: 678 MGKSNGDTSSVSPVPYVFNGGLRGRKTGAVEKVIERRQRRMIKNRESAARSRARKQAYTM 857
Query: 55 ELAAQVAQLRSENQQM 70
EL +VA+L+ EN+++
Sbjct: 858 ELEQEVAKLKEENEEL 905
>BP047366
Length = 535
Score = 48.5 bits (114), Expect = 5e-07
Identities = 22/42 (52%), Positives = 32/42 (75%)
Frame = -3
Query: 29 QRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQM 70
+R++KRMI NRESA RSR RKQ + EL +V++L EN+++
Sbjct: 488 ERRQKRMIKNRESAARSRARKQAYTQELEIKVSRLEEENERL 363
>AV426757
Length = 415
Score = 48.5 bits (114), Expect = 5e-07
Identities = 22/76 (28%), Positives = 48/76 (62%)
Frame = +1
Query: 24 QALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQMLTSLNLTTQRFMI 83
+ ++D ++ KR+++NR+SA+RSR+RK +++ EL V L++E + + + +I
Sbjct: 28 ETVVDPKRVKRILANRQSAQRSRVRKLQYISELERSVTTLQTEVSALSPRVAFLDHQRLI 207
Query: 84 IDAENSVLSAQMGELS 99
++ +NS L ++ L+
Sbjct: 208 LNVDNSTLKQRIAALA 255
>BP037350
Length = 495
Score = 45.8 bits (107), Expect = 3e-06
Identities = 22/50 (44%), Positives = 35/50 (70%)
Frame = -2
Query: 21 EELQALMDQRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSENQQM 70
E++ +R++KRMI NRESA RSR RK + +EL +V++L EN+++
Sbjct: 401 EDMVGKTVERRQKRMIKNRESAARSRARKHAYTNEL*NKVSRLEEENERL 252
>BP080797
Length = 498
Score = 41.2 bits (95), Expect = 7e-05
Identities = 19/39 (48%), Positives = 28/39 (71%)
Frame = -2
Query: 29 QRKRKRMISNRESARRSRMRKQKHLDELAAQVAQLRSEN 67
+R+ +R I NRESA RSR RKQ + +EL ++V +L +N
Sbjct: 449 ERRLRRKIKNRESAARSRARKQAYHNELVSKVTRLEQDN 333
>BF177802
Length = 481
Score = 39.3 bits (90), Expect = 3e-04
Identities = 25/54 (46%), Positives = 33/54 (60%), Gaps = 2/54 (3%)
Frame = +1
Query: 1 MASSSGTS--SGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQKH 52
M SS TS + SS + ++ + A DQR RKR++ NRESA RSR RKQ +
Sbjct: 319 MLSSFNTSFDASSSSGEKIAQQDRVYAFSDQR-RKRILKNRESALRSRARKQAY 477
>TC17850 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor, partial
(23%)
Length = 508
Score = 38.5 bits (88), Expect = 5e-04
Identities = 24/68 (35%), Positives = 36/68 (52%), Gaps = 3/68 (4%)
Frame = -1
Query: 11 SSMIQNSGSEEELQALMDQRKRKRMISN---RESARRSRMRKQKHLDELAAQVAQLRSEN 67
SS + N+G+ E L + K + A RSRM+KQKHL+ L ++V +L EN
Sbjct: 307 SSCLDNNGTYAEANELQFR*SAKGSTGE*CLTQFAHRSRMKKQKHLNGLWSKVVRLGIEN 128
Query: 68 QQMLTSLN 75
++ LN
Sbjct: 127 HNLVLKLN 104
>AV776064
Length = 418
Score = 35.8 bits (81), Expect = 0.003
Identities = 16/24 (66%), Positives = 20/24 (82%)
Frame = -1
Query: 29 QRKRKRMISNRESARRSRMRKQKH 52
+R++KRMI NRESA RSR RKQ +
Sbjct: 355 ERRQKRMIKNRESAARSRARKQAY 284
>AV412771
Length = 405
Score = 30.4 bits (67), Expect = 0.13
Identities = 17/42 (40%), Positives = 28/42 (66%)
Frame = +3
Query: 1 MASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESA 42
++ +S SSGS+ G++ + ++RKRKR++SNRESA
Sbjct: 288 LSMASIMSSGSASEGGGGADLHVD---NERKRKRILSNRESA 404
>BP034914
Length = 544
Score = 27.7 bits (60), Expect = 0.84
Identities = 18/57 (31%), Positives = 29/57 (50%), Gaps = 6/57 (10%)
Frame = +3
Query: 4 SSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISN------RESARRSRMRKQKHLD 54
S +SS SS +S S+ + + + K +R S+ RES+RR RK++ D
Sbjct: 69 SPSSSSSSSTASDSDSDSDRSSRRRREKERRRRSDGGSKRERESSRRREKRKKRDRD 239
>TC19682 similar to UP|O65169 (O65169) Glutaredoxin I, partial (54%)
Length = 551
Score = 25.8 bits (55), Expect = 3.2
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = +3
Query: 29 QRKRKRMISNRESARRSRMRKQKHL 53
+RKR R +NR R R+R Q+HL
Sbjct: 108 RRKRDRTATNRSL*FRFRLRSQRHL 182
>TC12976 similar to UP|O02448 (O02448) Lox22-otx (Fragment), partial (3%)
Length = 610
Score = 25.8 bits (55), Expect = 3.2
Identities = 14/50 (28%), Positives = 25/50 (50%)
Frame = +2
Query: 2 ASSSGTSSGSSMIQNSGSEEELQALMDQRKRKRMISNRESARRSRMRKQK 51
A ++ SSG+ + +S + ALM++RKR+ R+ R R +
Sbjct: 311 AEAASFSSGTRGLSSSPTPTPTPALMEERKREGKKKRHGLKRKKRKRMMR 460
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.124 0.319
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,490,171
Number of Sequences: 28460
Number of extensions: 12047
Number of successful extensions: 96
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 94
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 94
length of query: 147
length of database: 4,897,600
effective HSP length: 82
effective length of query: 65
effective length of database: 2,563,880
effective search space: 166652200
effective search space used: 166652200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0113.7


