

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0113.13
(359 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
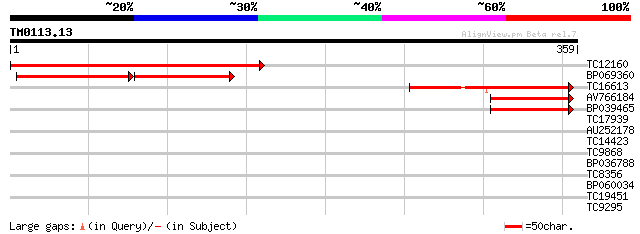
TC12160 weakly similar to UP|Q9SPV4 (Q9SPV4) S-adenosyl-L-methio... 183 4e-47
BP069360 76 2e-27
TC16613 weakly similar to UP|Q9SBK6 (Q9SBK6) Floral nectary-spec... 92 2e-19
AV766184 44 5e-05
BP039465 40 8e-04
TC17939 weakly similar to UP|Q945P2 (Q945P2) AT5g49210/K21P3_8, ... 32 0.16
AU252178 29 1.1
TC14423 similar to UP|Q8VWX2 (Q8VWX2) Gamma-tocopherol methyltra... 27 4.1
TC9868 27 6.9
BP036788 27 6.9
TC8356 similar to GB|AAM64696.1|21592747|AY087138 gamma-tocopher... 27 6.9
BP060034 26 9.0
TC19451 similar to GB|AAM65667.1|21593700|AY088122 J8-like prote... 26 9.0
TC9295 26 9.0
>TC12160 weakly similar to UP|Q9SPV4 (Q9SPV4)
S-adenosyl-L-methionine:salicylic acid carboxyl
methyltransferase, partial (32%)
Length = 520
Score = 183 bits (464), Expect = 4e-47
Identities = 85/161 (52%), Positives = 111/161 (68%)
Frame = +1
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCS 60
M + QVLHMNGG GD SYA NS Q+ + TK + E+++ LY P L +ADLGCS
Sbjct: 34 MDVAQVLHMNGGSGDTSYAKNSLPQQKAISLTKAMREEALTSLYLKKAPRILSIADLGCS 213
Query: 61 SGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK 120
SGPN LMV S ++ TV+ + + + + P +QFF+NDL NDFN FKSL + ++L E
Sbjct: 214 SGPNTLMVLSELVKTVENLCRKMKHKSPEYQFFMNDLPVNDFNNIFKSLDSYKEKLSDEL 393
Query: 121 GHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV 161
G + GPCFF+G PGSFYGR+FP ++HF HSSYSL WLS+V
Sbjct: 394 GAEIGPCFFTGVPGSFYGRIFPTKTLHFVHSSYSLQWLSRV 516
>BP069360
Length = 430
Score = 76.3 bits (186), Expect(2) = 2e-27
Identities = 39/64 (60%), Positives = 43/64 (66%), Gaps = 1/64 (1%)
Frame = +1
Query: 80 SQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFG-PCFFSGTPGSFYG 138
S+ L+ P LNDLF NDFNT F SLP FYK +QEKG+ FG CF S PGSFYG
Sbjct: 238 SRVLDCPVPELVVHLNDLFTNDFNTVFDSLPSFYKTQRQEKGNGFGSTCFVSPVPGSFYG 417
Query: 139 RLFP 142
RLFP
Sbjct: 418 RLFP 429
Score = 62.8 bits (151), Expect(2) = 2e-27
Identities = 29/74 (39%), Positives = 46/74 (61%)
Frame = +3
Query: 5 QVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCSSGPN 64
+VLHMN G + SY NS+ Q ++ T + +++ + C++ P + +ADLGCSSGPN
Sbjct: 12 KVLHMNKGVRETSYVMNSTLQSKIIDVTNPATKKAMVEILCSSRPVRMGIADLGCSSGPN 191
Query: 65 ALMVASNIINTVDA 78
L V + I++ V A
Sbjct: 192TLRVITEIVDVVHA 233
>TC16613 weakly similar to UP|Q9SBK6 (Q9SBK6) Floral nectary-specific
protein, partial (7%)
Length = 920
Score = 91.7 bits (226), Expect = 2e-19
Identities = 49/109 (44%), Positives = 68/109 (61%), Gaps = 5/109 (4%)
Frame = +3
Query: 254 IPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDT-----RAEAVAKY 308
+P YCPT EE++ +IE EGSF +Q L+TI+ W N ++ DDID R +AK
Sbjct: 3 MPLYCPTMEEVKXIIEGEGSFTLQTLKTIQISW--NSHLPDDIDVSVLDNXMRGYLIAKS 176
Query: 309 IRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELANLVMHITK 357
+RAV E +L FG IMDELF RF KI ++ +E L+ L+M++TK
Sbjct: 177 LRAVMEPLLSFAFGNNIMDELFSRFGKKISQIMEMETLQYTTLIMYMTK 323
>AV766184
Length = 376
Score = 43.5 bits (101), Expect = 5e-05
Identities = 22/54 (40%), Positives = 37/54 (67%), Gaps = 1/54 (1%)
Frame = -2
Query: 305 VAKYIRAVAESILKSEFGEAI-MDELFRRFKNKIVKLHGVEKLELANLVMHITK 357
VAK++RAVAE +L S FGEA+ ++E+FRR++ + E+ E N+ + +T+
Sbjct: 354 VAKFMRAVAEPLLISHFGEAVDIEEVFRRYQAILTDRMSKERTEFINVTISMTR 193
>BP039465
Length = 482
Score = 39.7 bits (91), Expect = 8e-04
Identities = 19/53 (35%), Positives = 34/53 (63%)
Frame = -2
Query: 305 VAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELANLVMHITK 357
VA+ +RAVAE +L + FGEAI++ +F R++ + EK + N+ + +T+
Sbjct: 460 VAQSVRAVAEPMLVNHFGEAIIENVFSRYQEILDDRMSKEKTKFVNVTILLTR 302
>TC17939 weakly similar to UP|Q945P2 (Q945P2) AT5g49210/K21P3_8, partial
(61%)
Length = 999
Score = 32.0 bits (71), Expect = 0.16
Identities = 26/98 (26%), Positives = 46/98 (46%), Gaps = 2/98 (2%)
Frame = +2
Query: 257 YCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDTRAEAVAKYIRAVAESI 316
Y +E+R+ EE ++ RLE R D + + +E R +A IRA +I
Sbjct: 329 YAKQVKELRKEYIEE--MELMRLEKQRKDEARREALRVATEERKRLKAQEAEIRAQQRNI 502
Query: 317 LKSEFGEAIMDELFRRFK--NKIVKLHGVEKLELANLV 352
+ +F E ++ E + + + VK+H +K E L+
Sbjct: 503 EQQQFRETLLKERAEKLEYWRRKVKMHEEKKAEKNELL 616
>AU252178
Length = 350
Score = 29.3 bits (64), Expect = 1.1
Identities = 22/80 (27%), Positives = 40/80 (49%), Gaps = 1/80 (1%)
Frame = +2
Query: 239 ASENLIEKAKLDAFNIPSYCPTAEEIRQVIE-EEGSFDIQRLETIRTDWIKNINVSDDID 297
++E ++EK K + Y EE+ + I EEGS + + E T + S++ D
Sbjct: 83 SAEAVVEKIKAAVQKLVPY--NVEELSEDISGEEGSLESENEEGWSTYSDVKSDASENED 256
Query: 298 EDTRAEAVAKYIRAVAESIL 317
+ AEAV + I+A + ++
Sbjct: 257 AEASAEAVVEKIKAAVQKLV 316
>TC14423 similar to UP|Q8VWX2 (Q8VWX2) Gamma-tocopherol methyltransferase,
partial (59%)
Length = 1239
Score = 27.3 bits (59), Expect = 4.1
Identities = 20/72 (27%), Positives = 35/72 (47%), Gaps = 3/72 (4%)
Frame = +1
Query: 197 FLGSRSCELLPGGAMVI-TLIGRDEN--NELINAWVVIGMVLNDMASENLIEKAKLDAFN 253
F+G + PGG ++I T RD E + W +NL+++ DAF
Sbjct: 679 FVGELARVAAPGGTIIIVTWCHRDLGPAEESLQPW-----------EQNLLKRI-CDAFY 822
Query: 254 IPSYCPTAEEIR 265
+P++C TA+ ++
Sbjct: 823 LPAWCSTADYVK 858
>TC9868
Length = 540
Score = 26.6 bits (57), Expect = 6.9
Identities = 16/50 (32%), Positives = 23/50 (46%), Gaps = 5/50 (10%)
Frame = +1
Query: 113 YKRLQQEKGHKFGPCFF-----SGTPGSFYGRLFPNDSVHFFHSSYSLHW 157
+K QQ KG K SG +YG +F + S +F H +LH+
Sbjct: 196 FKNYQQSKGTKVPTTLTFVLDGSGITKDYYGLIFLSHSFYF*HLPKNLHF 345
>BP036788
Length = 559
Score = 26.6 bits (57), Expect = 6.9
Identities = 22/67 (32%), Positives = 35/67 (51%), Gaps = 4/67 (5%)
Frame = +3
Query: 23 SFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCSSG----PNALMVASNIINTVDA 78
+F R+V+ +H +D Y A LK+AD+GC G P A + A+ + VDA
Sbjct: 333 AFVRSVLC--RHFKKDP----YSAKPLEGLKIADVGCGGGILSEPLARLGAT--VTGVDA 488
Query: 79 VSQNLNL 85
V +N+ +
Sbjct: 489 VEKNIKI 509
>TC8356 similar to GB|AAM64696.1|21592747|AY087138 gamma-tocopherol
methyltransferase {Arabidopsis thaliana;} , partial
(30%)
Length = 642
Score = 26.6 bits (57), Expect = 6.9
Identities = 24/99 (24%), Positives = 48/99 (48%)
Frame = +2
Query: 241 ENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDT 300
+NL+++ DAF +P++C TA+ ++ ++E DI+ DW + V+
Sbjct: 23 QNLLKRI-CDAFYLPAWCSTADYVK-LLESHSLQDIK-----SADW--SPFVAPFWPAVI 175
Query: 301 RAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
R+ K + ++ S +K+ G M + FK ++K
Sbjct: 176 RSAFTWKGLTSLLRSGMKTIKGALAMPLMIEGFKKGVIK 292
>BP060034
Length = 364
Score = 26.2 bits (56), Expect = 9.0
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +3
Query: 332 RFKNKIVKLHGVEKLELANLVMHIT 356
++++ IV H VEKLE+ +HIT
Sbjct: 228 KWQHAIVDFHHVEKLEVGGHAVHIT 302
>TC19451 similar to GB|AAM65667.1|21593700|AY088122 J8-like protein
{Arabidopsis thaliana;}, partial (35%)
Length = 480
Score = 26.2 bits (56), Expect = 9.0
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +3
Query: 43 LYCATFPSCLKVADLGCS 60
L C+T +C K A++GCS
Sbjct: 297 LLCSTIQTCAKAANVGCS 350
>TC9295
Length = 543
Score = 26.2 bits (56), Expect = 9.0
Identities = 13/45 (28%), Positives = 24/45 (52%)
Frame = -2
Query: 172 IYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLI 216
I+L S +V+++Y ++ LFL S+SC + G + +I
Sbjct: 143 IFLPPGS*VSVYESYIYSIKQQVDLFLSSQSCVIGAGDQLFPAII 9
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.137 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,750,600
Number of Sequences: 28460
Number of extensions: 77004
Number of successful extensions: 392
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 389
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 390
length of query: 359
length of database: 4,897,600
effective HSP length: 91
effective length of query: 268
effective length of database: 2,307,740
effective search space: 618474320
effective search space used: 618474320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0113.13


