

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.9
(341 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
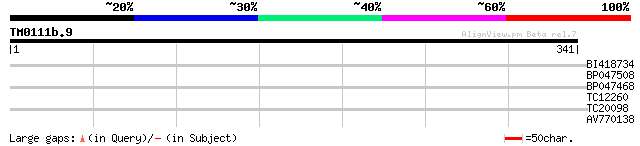
Score E
Sequences producing significant alignments: (bits) Value
BI418734 28 2.9
BP047508 27 6.5
BP047468 27 6.5
TC12260 26 8.4
TC20098 similar to UP|APG8_YEAST (P38182) Autophagy protein 8 [C... 26 8.4
AV770138 26 8.4
>BI418734
Length = 539
Score = 27.7 bits (60), Expect = 2.9
Identities = 13/43 (30%), Positives = 21/43 (48%)
Frame = +2
Query: 73 WISWLASSQVGHVGSFEAAWWSWYPRCDTREYFLVEEIGVGLC 115
W+SWL + G + + +W W R + +FL I V +C
Sbjct: 236 WLSWLVHAY-GLLVWWRLVYWEWVARHSSLRFFLKHLILVVIC 361
>BP047508
Length = 504
Score = 26.6 bits (57), Expect = 6.5
Identities = 9/22 (40%), Positives = 16/22 (71%)
Frame = -3
Query: 66 LYLVFPSWISWLASSQVGHVGS 87
+Y +FPSW ++L+S V +G+
Sbjct: 472 MYFIFPSWTNFLSSPIVSSMGT 407
>BP047468
Length = 502
Score = 26.6 bits (57), Expect = 6.5
Identities = 9/22 (40%), Positives = 16/22 (71%)
Frame = -2
Query: 66 LYLVFPSWISWLASSQVGHVGS 87
+Y +FPSW ++L+S V +G+
Sbjct: 435 MYFIFPSWTNFLSSPIVSSMGT 370
>TC12260
Length = 320
Score = 26.2 bits (56), Expect = 8.4
Identities = 10/20 (50%), Positives = 13/20 (65%), Gaps = 2/20 (10%)
Frame = -1
Query: 81 QVGHVGSFEAAWW--SWYPR 98
++GH+GS AWW W PR
Sbjct: 233 RLGHLGSTGLAWWLSPWLPR 174
>TC20098 similar to UP|APG8_YEAST (P38182) Autophagy protein 8 [Contains:
Apg8FG], partial (71%)
Length = 468
Score = 26.2 bits (56), Expect = 8.4
Identities = 12/34 (35%), Positives = 15/34 (43%)
Frame = -3
Query: 192 FCAYFRYRDDSCRCYCLW*MERLAVVHSHPYFRH 225
F YFR C +C + +H HPYF H
Sbjct: 274 FMNYFRSSSQMCSPHC----KSYKGIHLHPYFPH 185
>AV770138
Length = 483
Score = 26.2 bits (56), Expect = 8.4
Identities = 9/16 (56%), Positives = 12/16 (74%)
Frame = -1
Query: 191 SFCAYFRYRDDSCRCY 206
SFCAY+R+ +C CY
Sbjct: 75 SFCAYWRF*TPTCICY 28
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.358 0.162 0.621
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,791,967
Number of Sequences: 28460
Number of extensions: 134997
Number of successful extensions: 1597
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1574
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1596
length of query: 341
length of database: 4,897,600
effective HSP length: 91
effective length of query: 250
effective length of database: 2,307,740
effective search space: 576935000
effective search space used: 576935000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0111b.9


