

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.4
(467 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
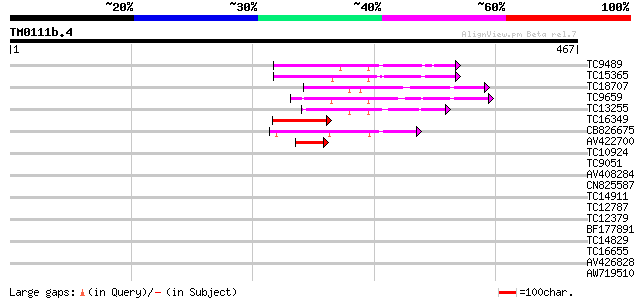
Score E
Sequences producing significant alignments: (bits) Value
TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase... 65 3e-11
TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {... 60 8e-10
TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J... 57 6e-09
TC9659 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATP... 50 6e-07
TC13255 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (41%) 44 6e-05
TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulat... 43 1e-04
CB826675 42 2e-04
AV422700 42 2e-04
TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPas... 40 0.001
TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 ... 39 0.002
AV408284 38 0.003
CN825587 38 0.004
TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), pa... 37 0.007
TC12787 weakly similar to UP|MSP1_YEAST (P28737) MSP1 protein (T... 36 0.012
TC12379 homologue to UP|O04327 (O04327) Cell division protein Ft... 36 0.015
BF177891 35 0.020
TC14829 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulat... 35 0.026
TC16655 homologue to UP|O99018 (O99018) Chloroplast protease pre... 35 0.026
AV426828 35 0.034
AW719510 34 0.045
>TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase subunit
RPT2a, partial (69%)
Length = 916
Score = 64.7 bits (156), Expect = 3e-11
Identities = 53/162 (32%), Positives = 81/162 (49%), Gaps = 8/162 (4%)
Frame = +1
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIY-----D 272
+E+ ++ L E Y G +G +LYG PGTGK+ L A+AN S +
Sbjct: 412 QEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLLAKAVANSTSATFLRVVGSE 591
Query: 273 LDLTALNDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
L L D L L +++ SI+ I++ID + + D + + + T+
Sbjct: 592 LIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDA---VGTKRYDAHSGGEREIQRTMLE 762
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
LLN +DG S G+ ++I+ TN E LDPALLRPGR+D +
Sbjct: 763 LLNQLDGFDSR-GDVKVIL-ATNRIESLDPALLRPGRIDRKI 882
>TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {Arabidopsis
thaliana;}, partial (87%)
Length = 1385
Score = 60.1 bits (144), Expect = 8e-10
Identities = 48/162 (29%), Positives = 81/162 (49%), Gaps = 8/162 (4%)
Frame = +3
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMAN-----YLSYDIYD 272
KE+ ++ ++ E + G A +G LLYGPPGTGK+ L A+A+ ++ +
Sbjct: 357 KEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLLARAVAHHTDCTFIRVSGSE 536
Query: 273 LDLTALNDNKDLKNLLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSG 329
L + + + L M+ SI+ +++ID SI R E + + + T+
Sbjct: 537 LVQKYIGEGSRMVRELFVMAREHAPSIIFMDEID-SIG-SARMESGSGXGDSEVQRTMLE 710
Query: 330 LLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
LLN +DG + + ++ TN + LD ALLRPGR+D +
Sbjct: 711 LLNQLDGFEA--SNKIKVLMATNRIDILDQALLRPGRIDRKI 830
>TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J8.110 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 771
Score = 57.0 bits (136), Expect = 6e-09
Identities = 42/161 (26%), Positives = 78/161 (48%), Gaps = 8/161 (4%)
Frame = +3
Query: 243 RGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTAL------NDNKDLKNL--LLAMSNR 294
RG LL+GPPGTGK+ L A+AN ++ ++ + D K+++ L L A
Sbjct: 306 RGILLFGPPGTGKTMLAXAIANEAGASFINVSMSTITSKWFGEDEKNVRALFSLAAKVAP 485
Query: 295 SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHR 354
+I+ ++++D + R + EA + N+ + DGL + GE+ +++ TN
Sbjct: 486 TIIFVDEVDSMLGQRTRVGEHEAMRKIKNE-----FMTHWDGLLTAPGEQILVLAATNRP 650
Query: 355 ERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHK 395
LD A++R R + + + + E + T L +H+
Sbjct: 651 FDLDEAIIR--RFERRIMVGLPSVENREMILKTLLA*EKHE 767
>TC9659 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATPase,
partial (74%)
Length = 1060
Score = 50.4 bits (119), Expect = 6e-07
Identities = 46/175 (26%), Positives = 81/175 (46%), Gaps = 8/175 (4%)
Frame = +1
Query: 232 EFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMA-----NYLSYDIYDLDLTALNDNKDLKN 286
+F+ + W R +LLYGPPGTGKS L A+A + S DL + +++ L +
Sbjct: 535 QFFTGKRRPW-RAFLLYGPPGTGKSYLAKAVATEADSTFFSVSSSDLGSKWMGESEKLVS 711
Query: 287 LLLAMSNR---SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGE 343
L M+ SI+ +++ID + EA++ + LL + G+ + +
Sbjct: 712 NLFQMARESAPSIIFVDEIDSLCGQRGEGNESEASRR-----IKTELLVQMQGVGN-NDQ 873
Query: 344 ERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFE 398
+ +++ TN LD A+ R R D + + A + + +LG + H L E
Sbjct: 874 KVLVLAATNTPYALDQAIRR--RFDKRIYIPLPDVKARQHMFKVHLGDTPHNLAE 1032
>TC13255 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (41%)
Length = 532
Score = 43.9 bits (102), Expect = 6e-05
Identities = 36/131 (27%), Positives = 66/131 (49%), Gaps = 8/131 (6%)
Frame = +1
Query: 241 WKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTAL-----NDNKDLKNLLLAMSNR- 294
WK G LL+GPPGTGK+ L A+A +++ +++ D++ L +L ++
Sbjct: 34 WK-GILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHH 210
Query: 295 --SILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTN 352
S + +++ID I + E +++ ++ + LL +DGL + E ++ TN
Sbjct: 211 APSTIFLDEIDAIIS----QRGEARSEHEASRRLKTELLIQMDGL-TRTDELVFVLAATN 375
Query: 353 HRERLDPALLR 363
LD A+LR
Sbjct: 376 LPWELDAAMLR 408
>TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulatory subunit
6B homolog, partial (65%)
Length = 901
Score = 42.7 bits (99), Expect = 1e-04
Identities = 22/49 (44%), Positives = 30/49 (60%)
Frame = +3
Query: 217 KKELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANY 265
K+E+ ++ L E Y + G RG LLYGPPGTGK+ L A+AN+
Sbjct: 525 KQEIREAVELPLTHHELYKQIGIDPPRGVLLYGPPGTGKTMLAKAVANH 671
>CB826675
Length = 551
Score = 42.4 bits (98), Expect = 2e-04
Identities = 39/136 (28%), Positives = 62/136 (44%), Gaps = 11/136 (8%)
Frame = +1
Query: 215 GLKK---ELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAM-----ANYL 266
GL+K ELV + + KE + + G +G LLYGPPGTGK+ + A A +L
Sbjct: 148 GLEKQIQELVEAIVLPMTHKERFQKLGVRPPKGVLLYGPPGTGKTLMARACAAQTNATFL 327
Query: 267 SYDIYDLDLTALNDNKDLKNLLLAMSNRS---ILVIEDIDCSIKLPNREEDEEAAKNGDN 323
L + D L ++ I+ I++ID + + D E + + +
Sbjct: 328 KLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDA---IGTKRFDSEVSGDREV 498
Query: 324 KMTLSGLLNVIDGLWS 339
+ T+ LLN +DG S
Sbjct: 499 QRTMLELLNQLDGFSS 546
>AV422700
Length = 472
Score = 42.0 bits (97), Expect = 2e-04
Identities = 17/27 (62%), Positives = 21/27 (76%)
Frame = +2
Query: 236 RTGKAWKRGYLLYGPPGTGKSSLIAAM 262
+ G W RG LLYGPPGTGK+SL+ A+
Sbjct: 290 KLGLKWPRGLLLYGPPGTGKTSLVRAV 370
>TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPase subunit
RPT4a, partial (49%)
Length = 648
Score = 39.7 bits (91), Expect = 0.001
Identities = 20/47 (42%), Positives = 28/47 (59%)
Frame = +2
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMAN 264
+EL ++ L E + R G +G LLYGPPGTGK+ L A+A+
Sbjct: 488 RELRESIELPLMNPELFLRVGIKPPKGVLLYGPPGTGKTLLARAIAS 628
>TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1a) (Regulatory particle triple-A ATPase subunit 1a),
partial (46%)
Length = 867
Score = 38.5 bits (88), Expect = 0.002
Identities = 22/53 (41%), Positives = 35/53 (65%), Gaps = 2/53 (3%)
Frame = +2
Query: 321 GDNKM--TLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHV 371
GDN++ T+ ++N +DG + G ++++ TN + LDPALLRPGR+D V
Sbjct: 161 GDNEVQRTMLEIVNQLDG-FDARGNIKVLM-ATNRPDTLDPALLRPGRLDRKV 313
>AV408284
Length = 425
Score = 38.1 bits (87), Expect = 0.003
Identities = 27/87 (31%), Positives = 47/87 (53%), Gaps = 8/87 (9%)
Frame = +1
Query: 242 KRGYLLYGPPGTGKSSLIAAMAN-----YLSYDIYDLDLTALNDNKDLKNLLLAMSNR-- 294
++G LLYGPPGTGK+ L A+A +++ I +L D + L + +++++
Sbjct: 91 QKGVLLYGPPGTGKTMLAKAIAKESGAVFINVRISNLMSKWFGDAQKLVAAVFSLAHKLQ 270
Query: 295 -SILVIEDIDCSIKLPNREEDEEAAKN 320
+I+ I+++D S R D EA N
Sbjct: 271 PAIIFIDEVD-SFLGQRRTTDHEALLN 348
>CN825587
Length = 663
Score = 37.7 bits (86), Expect = 0.004
Identities = 21/54 (38%), Positives = 30/54 (54%)
Frame = +2
Query: 320 NGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNL 373
N + + T++ LL +DG G I++ TN + LD ALLRPGR D V +
Sbjct: 86 NDEREQTINQLLTEMDGFSGNSGV--IVLAATNRPDVLDSALLRPGRFDRQVTV 241
>TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), partial (32%)
Length = 1418
Score = 37.0 bits (84), Expect = 0.007
Identities = 21/42 (50%), Positives = 27/42 (64%)
Frame = +3
Query: 327 LSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMD 368
LS +L IDGL S ++ II +N + +DPALLRPGR D
Sbjct: 597 LSQMLAEIDGL-SDSSQDLFIIGASNRPDLIDPALLRPGRFD 719
Score = 36.6 bits (83), Expect = 0.009
Identities = 41/170 (24%), Positives = 73/170 (42%)
Frame = +2
Query: 127 VDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
VD S ++++S ++ + ++S KE + N+ LER+K K+ A+
Sbjct: 41 VDKDGSEDVDSSLTSGIIEDNNKSKVSPRIPGKEDVMNA-----LERSK--KRNASALGT 199
Query: 187 YTVEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKRGYL 246
V W G D +KK ++ + L K+ + +G + G L
Sbjct: 200 PKVPNVKWEDVGGLED-------------VKKSILDTVQLPLLHKDLF-ASGLRKRSGVL 337
Query: 247 LYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSI 296
LYGPPGTGK+ L A+A S + + +L N+ + S +++
Sbjct: 338 LYGPPGTGKTLLAKAVATECSLNFLSV------KGPELINMYIGESEKNV 469
>TC12787 weakly similar to UP|MSP1_YEAST (P28737) MSP1 protein (TAT-binding
homolog 4), partial (20%)
Length = 389
Score = 36.2 bits (82), Expect = 0.012
Identities = 21/90 (23%), Positives = 46/90 (50%), Gaps = 8/90 (8%)
Frame = +3
Query: 243 RGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALND------NKDLKNL--LLAMSNR 294
+G LL+GPPGTGK+ L A+A + ++ ++++ K +K + L + +
Sbjct: 84 KGILLFGPPGTGKTMLAKAVATEAGANFINISMSSITSKWFGEGEKYVKAVFSLASKISP 263
Query: 295 SILVIEDIDCSIKLPNREEDEEAAKNGDNK 324
S++ ++++D + + EA + N+
Sbjct: 264 SVIFVDEVDXMLXRRENPGEHEAMRKMXNE 353
>TC12379 homologue to UP|O04327 (O04327) Cell division protein FtsH isolog,
partial (15%)
Length = 505
Score = 35.8 bits (81), Expect = 0.015
Identities = 27/78 (34%), Positives = 39/78 (49%), Gaps = 1/78 (1%)
Frame = +1
Query: 295 SILVIEDIDCSIKLPNREEDEEAAKNGDNK-MTLSGLLNVIDGLWSCCGEERIIIFTTNH 353
S++ I+++D RE G + TL+ LL +DG E I I +TN
Sbjct: 97 SVVFIDELDAV----GRERGLIKGSGGQERDATLNQLLVCLDGFEG--RGEVITIASTNR 258
Query: 354 RERLDPALLRPGRMDMHV 371
+ LDPAL+RPGR D +
Sbjct: 259 PDILDPALVRPGRFDRKI 312
>BF177891
Length = 390
Score = 35.4 bits (80), Expect = 0.020
Identities = 23/90 (25%), Positives = 44/90 (48%), Gaps = 8/90 (8%)
Frame = +1
Query: 243 RGYLLYGPPGTGKSSLIAAM-----ANYLSYDIYDLDLTALNDNKDLKNLLLAMSNR--- 294
+G LL+GPPGTGK+ + A+ AN++S L D + L L + +++
Sbjct: 82 KGILLFGPPGTGKTLMAKALATEAGANFISVSGSTLTSKWFGDAEKLTKALFSFASKLAP 261
Query: 295 SILVIEDIDCSIKLPNREEDEEAAKNGDNK 324
I+ ++++D + + EA + N+
Sbjct: 262 VIIFVDEVDSLLGARGGAFEHEATRRMRNE 351
>TC14829 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulatory subunit
6B homolog, partial (28%)
Length = 662
Score = 35.0 bits (79), Expect = 0.026
Identities = 14/25 (56%), Positives = 18/25 (72%)
Frame = +2
Query: 347 IIFTTNHRERLDPALLRPGRMDMHV 371
+I TN + LDPALLRPGR+D +
Sbjct: 20 VIMATNRADTLDPALLRPGRLDRKI 94
>TC16655 homologue to UP|O99018 (O99018) Chloroplast protease precursor,
partial (39%)
Length = 934
Score = 35.0 bits (79), Expect = 0.026
Identities = 18/46 (39%), Positives = 26/46 (56%)
Frame = +3
Query: 218 KELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMA 263
K+ ++ FL+ E + G +G LL GPPGTGK+ L A+A
Sbjct: 681 KQDFMEVVEFLKKPERFTAVGARIPKGVLLIGPPGTGKTLLAKAIA 818
>AV426828
Length = 401
Score = 34.7 bits (78), Expect = 0.034
Identities = 20/38 (52%), Positives = 24/38 (62%), Gaps = 3/38 (7%)
Frame = +1
Query: 246 LLYGPPGTGKSSLIAAMANYL---SYDIYDLDLTALND 280
LLYGPPGTGK+S I A+A L Y L+L A +D
Sbjct: 121 LLYGPPGTGKTSTILAVARKLYGAQYRNMILELNASDD 234
>AW719510
Length = 358
Score = 34.3 bits (77), Expect = 0.045
Identities = 14/25 (56%), Positives = 18/25 (72%)
Frame = -2
Query: 347 IIFTTNHRERLDPALLRPGRMDMHV 371
++ TN + LDPALLRPGR+D V
Sbjct: 342 VLMATNRPDTLDPALLRPGRLDRKV 268
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,847,528
Number of Sequences: 28460
Number of extensions: 82397
Number of successful extensions: 466
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 462
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 463
length of query: 467
length of database: 4,897,600
effective HSP length: 94
effective length of query: 373
effective length of database: 2,222,360
effective search space: 828940280
effective search space used: 828940280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0111b.4


