

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111a.4
(74 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
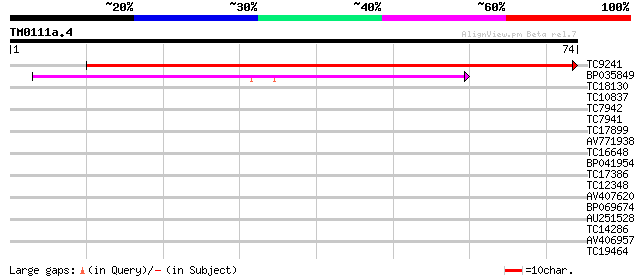
Score E
Sequences producing significant alignments: (bits) Value
TC9241 similar to GB|AAP68270.1|31711828|BT008831 At1g53290 {Ara... 131 3e-32
BP035849 44 6e-06
TC18130 27 0.55
TC10837 26 1.6
TC7942 similar to UP|Q8LPE6 (Q8LPE6) Epoxide hydrolase (Fragmen... 25 2.1
TC7941 similar to UP|O49857 (O49857) Epoxide hydrolase , partia... 25 2.1
TC17899 similar to UP|AAR23727 (AAR23727) At5g19260, partial (9%) 25 2.7
AV771938 25 2.7
TC16648 similar to PIR|T10625|T10625 reticuline oxidase homolog ... 25 2.7
BP041954 25 2.7
TC17386 similar to UP|Q9LU46 (Q9LU46) DEAD-box protein abstrakt,... 25 3.5
TC12348 weakly similar to UP|Q8RY61 (Q8RY61) AT4g26990/F10M23_33... 23 7.9
AV407620 23 7.9
BP069674 23 7.9
AU251528 23 7.9
TC14286 homologue to UP|O65357 (O65357) Aquaporin 2, complete 23 7.9
AV406957 23 7.9
TC19464 similar to UP|Q84LR1 (Q84LR1) Root-specific metal transp... 23 7.9
>TC9241 similar to GB|AAP68270.1|31711828|BT008831 At1g53290 {Arabidopsis
thaliana;}, partial (19%)
Length = 589
Score = 131 bits (329), Expect = 3e-32
Identities = 56/64 (87%), Positives = 62/64 (96%)
Frame = +1
Query: 11 IGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQMESCTQSPT 70
IGAWMLAMNVNHENNHELCAP+CT+TS+AVWDIPKCSGLCNPEK+MLELHQ ESC+QSPT
Sbjct: 1 IGAWMLAMNVNHENNHELCAPDCTATSVAVWDIPKCSGLCNPEKKMLELHQKESCSQSPT 180
Query: 71 AESD 74
E+D
Sbjct: 181LEAD 192
>BP035849
Length = 552
Score = 43.9 bits (102), Expect = 6e-06
Identities = 20/65 (30%), Positives = 35/65 (53%), Gaps = 8/65 (12%)
Frame = -2
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PEC-----TSTSIAVWDIPKCSGLCNPEKR 55
++NEDV++G+W+L + V H + +C P+C + A CSG+C +R
Sbjct: 395 YANEDVSLGSWLLGLEVEHVDERSMCCGTPPDCDWKARSGNMCAASFDWSCSGICKSVER 216
Query: 56 MLELH 60
M ++H
Sbjct: 215 MKDIH 201
>TC18130
Length = 669
Score = 27.3 bits (59), Expect = 0.55
Identities = 20/54 (37%), Positives = 29/54 (53%), Gaps = 7/54 (12%)
Frame = -3
Query: 18 MNVNHENNHELCAPECT----STSIAVWDI--PK-CSGLCNPEKRMLELHQMES 64
+N+ HE+ LC P CT +TSI+ ++ PK S L + RML +M S
Sbjct: 175 INIKHED*SRLCKPSCTHQTDATSISRMNLLXPKNKSLLLSAHPRMLRQFEMSS 14
>TC10837
Length = 667
Score = 25.8 bits (55), Expect = 1.6
Identities = 11/27 (40%), Positives = 18/27 (65%), Gaps = 1/27 (3%)
Frame = -3
Query: 46 CSGLCNPE-KRMLELHQMESCTQSPTA 71
C+GL + + M ++H++E C QSP A
Sbjct: 266 CTGLAHSKLSSMSQIHKLEGC*QSPCA 186
>TC7942 similar to UP|Q8LPE6 (Q8LPE6) Epoxide hydrolase (Fragment) ,
partial (91%)
Length = 1288
Score = 25.4 bits (54), Expect = 2.1
Identities = 11/37 (29%), Positives = 18/37 (47%)
Frame = +2
Query: 13 AWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGL 49
+WM+ + V N LC P C ++ + PKC +
Sbjct: 311 SWMVCLLVPTRQNQSLCLPWCYFPTLPCKE-PKCQNI 418
>TC7941 similar to UP|O49857 (O49857) Epoxide hydrolase , partial (48%)
Length = 610
Score = 25.4 bits (54), Expect = 2.1
Identities = 8/22 (36%), Positives = 13/22 (58%)
Frame = +1
Query: 13 AWMLAMNVNHENNHELCAPECT 34
+WM+ + V N LC P+C+
Sbjct: 427 SWMVCLLVPTRQNQSLCLPQCS 492
>TC17899 similar to UP|AAR23727 (AAR23727) At5g19260, partial (9%)
Length = 603
Score = 25.0 bits (53), Expect = 2.7
Identities = 10/30 (33%), Positives = 16/30 (53%)
Frame = +2
Query: 42 DIPKCSGLCNPEKRMLELHQMESCTQSPTA 71
++P+CS C P K+ + + CT S A
Sbjct: 311 ELPRCSFRCLPRKQRIITERNNVCTPSAEA 400
>AV771938
Length = 582
Score = 25.0 bits (53), Expect = 2.7
Identities = 18/65 (27%), Positives = 30/65 (45%)
Frame = +2
Query: 8 DVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQMESCTQ 67
+V + ++A VNH NNH+ + E ++ +A W GL ++ L Q
Sbjct: 203 EVVENSQIMAETVNHLNNHD--SKEKEASPLAAWLKEMKEGLDVVRHKIQSLTAKVKEGQ 376
Query: 68 SPTAE 72
PTA+
Sbjct: 377 YPTAD 391
>TC16648 similar to PIR|T10625|T10625 reticuline oxidase homolog F21C20.180
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(19%)
Length = 483
Score = 25.0 bits (53), Expect = 2.7
Identities = 10/22 (45%), Positives = 11/22 (49%)
Frame = -1
Query: 44 PKCSGLCNPEKRMLELHQMESC 65
P+C LCN K L Q SC
Sbjct: 252 PQCQDLCNSNKHSLSWTQKGSC 187
>BP041954
Length = 457
Score = 25.0 bits (53), Expect = 2.7
Identities = 9/22 (40%), Positives = 13/22 (58%)
Frame = -1
Query: 14 WMLAMNVNHENNHELCAPECTS 35
W+L + + H N L AP C+S
Sbjct: 142 WLLLLIIQHSQNPSLPAPLCSS 77
>TC17386 similar to UP|Q9LU46 (Q9LU46) DEAD-box protein abstrakt, partial
(19%)
Length = 507
Score = 24.6 bits (52), Expect = 3.5
Identities = 9/28 (32%), Positives = 14/28 (49%)
Frame = -2
Query: 19 NVNHENNHELCAPECTSTSIAVWDIPKC 46
N+ +NNH C S+ + +WD C
Sbjct: 320 NLQIQNNHVYCLQLPCSSDVLIWDNHGC 237
>TC12348 weakly similar to UP|Q8RY61 (Q8RY61) AT4g26990/F10M23_330, partial
(17%)
Length = 447
Score = 23.5 bits (49), Expect = 7.9
Identities = 12/25 (48%), Positives = 14/25 (56%)
Frame = -2
Query: 2 RMFSNEDVTIGAWMLAMNVNHENNH 26
R S ED I +L MN+N E NH
Sbjct: 272 RRSSMEDSRIDRAILDMNINWEANH 198
>AV407620
Length = 414
Score = 23.5 bits (49), Expect = 7.9
Identities = 8/18 (44%), Positives = 12/18 (66%)
Frame = -2
Query: 1 FRMFSNEDVTIGAWMLAM 18
FR+ SN D ++G W L +
Sbjct: 215 FRLISNIDFSVGLWKLPL 162
>BP069674
Length = 513
Score = 23.5 bits (49), Expect = 7.9
Identities = 13/37 (35%), Positives = 19/37 (51%)
Frame = +3
Query: 35 STSIAVWDIPKCSGLCNPEKRMLELHQMESCTQSPTA 71
ST + + D+P +G E+ LH + TQ PTA
Sbjct: 297 STILLIDDLPMTNGKTAFERLKYCLHLLVHSTQIPTA 407
>AU251528
Length = 388
Score = 23.5 bits (49), Expect = 7.9
Identities = 10/32 (31%), Positives = 15/32 (46%)
Frame = +3
Query: 31 PECTSTSIAVWDIPKCSGLCNPEKRMLELHQM 62
P C T + + PK + +P ELHQ+
Sbjct: 246 PSCLLTGLEAQNAPKGCRVLDPAATPFELHQV 341
>TC14286 homologue to UP|O65357 (O65357) Aquaporin 2, complete
Length = 1278
Score = 23.5 bits (49), Expect = 7.9
Identities = 10/25 (40%), Positives = 16/25 (64%)
Frame = -2
Query: 34 TSTSIAVWDIPKCSGLCNPEKRMLE 58
TSTS+A+ +IP C E+++ E
Sbjct: 125 TSTSLAI*EIPFCKTRVKKEEKLSE 51
>AV406957
Length = 439
Score = 23.5 bits (49), Expect = 7.9
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = -3
Query: 16 LAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEK 54
LA+ + H+ H++ P S + V P+CS NP++
Sbjct: 422 LALQI-HQRKHQVENPVVASIFLPV*GEPRCSPQPNPQE 309
>TC19464 similar to UP|Q84LR1 (Q84LR1) Root-specific metal transporter,
partial (22%)
Length = 503
Score = 23.5 bits (49), Expect = 7.9
Identities = 12/36 (33%), Positives = 18/36 (49%)
Frame = -2
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIA 39
F+ D I A + + NH +++ AP C S S A
Sbjct: 298 FAAND*IISANISPTSTNHNSSYLWLAPACRSVSTA 191
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.127 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,770,939
Number of Sequences: 28460
Number of extensions: 28159
Number of successful extensions: 147
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 146
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 146
length of query: 74
length of database: 4,897,600
effective HSP length: 50
effective length of query: 24
effective length of database: 3,474,600
effective search space: 83390400
effective search space used: 83390400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0111a.4


