

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111a.1
(863 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
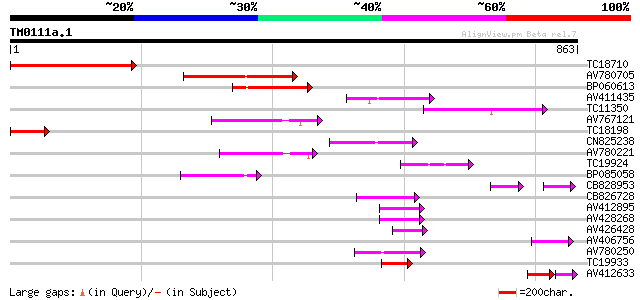
Score E
Sequences producing significant alignments: (bits) Value
TC18710 193 1e-49
AV780705 182 2e-46
BP060613 132 2e-31
AV411435 112 2e-25
TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3,... 86 3e-17
AV767121 84 1e-16
TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, p... 73 2e-13
CN825238 66 2e-11
AV780221 62 5e-10
TC19924 56 2e-08
BP085058 55 5e-08
CB828953 42 8e-08
CB826728 53 2e-07
AV412895 53 2e-07
AV428268 52 5e-07
AV426428 51 7e-07
AV406756 51 7e-07
AV780250 49 3e-06
TC19933 48 8e-06
AV412633 44 1e-05
>TC18710
Length = 843
Score = 193 bits (490), Expect = 1e-49
Identities = 97/192 (50%), Positives = 126/192 (65%)
Frame = +2
Query: 1 SFFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRL 60
+F + +P INET+V L+PKV PE+I Q RPIS C FLYKVI+KI V +LK M L
Sbjct: 266 AFLERAEIPNEINETLVTLIPKVPHPESINQFRPISGCTFLYKVISKIFVARLKDAMPDL 445
Query: 61 ATQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKAC 120
+ QS FI GRQIQDN+LI EAFH + IKL+MNKAYDR+EW FL++
Sbjct: 446 ISPMQSGFIQGRQIQDNLLIVQEAFHAINRPGALGRNHSIIKLDMNKAYDRVEWKFLESS 625
Query: 121 LTSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMS 180
L +FGFS WV IM V+G S+ +K+NG V +++PQRG+RQG P SPYLF+ E +S
Sbjct: 626 LLAFGFSTNWVKMIMILVSGVSYNYKINGVVGPKLLPQRGLRQGDPFSPYLFLFTMEVLS 805
Query: 181 HLLLKAQEEGRI 192
L+ + G +
Sbjct: 806 LLIQNSFNMGNL 841
>AV780705
Length = 524
Score = 182 bits (462), Expect = 2e-46
Identities = 84/175 (48%), Positives = 120/175 (68%), Gaps = 2/175 (1%)
Frame = -2
Query: 265 SILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVLDKIDGWKEKLLNQAGKEVLIKAV 324
++L M WD PG YLGLPA WG++++++L WI+ +V +K++GWKE LLNQAGKEVLIKA+
Sbjct: 523 NLLHMPIWDDPGHYLGLPAIWGRNKSHSLVWIEEKVKEKLEGWKETLLNQAGKEVLIKAI 344
Query: 325 LQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKERGIHWKDWHFIS--KAKCKGGLG 382
+QA+ +Y M+I+ FP+ FC ++ A +A FWW+S G E + + +K KGG+G
Sbjct: 343 IQAIPSYAMTIVHFPKTFCNSLYALVADFWWKSQG-EGAVEYTGAVGTK*PLSKEKGGVG 167
Query: 383 FKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNSSFLQSGRTARNSWGWQ 437
F+D T NLA LA+QAWR+ T P+ALWVR +K LYFP + + W W+
Sbjct: 166 FRDLRTQNLAFLARQAWRVLTNPEALWVRVMKSLYFPTPALCLLLWDGTHPWMWR 2
>BP060613
Length = 378
Score = 132 bits (332), Expect = 2e-31
Identities = 61/123 (49%), Positives = 78/123 (62%)
Frame = +1
Query: 339 PQNFCQAICASIARFWWRSPGKERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQA 398
PQ+ Q S + WW S G RG+HW+ W +++ K +GGLGFKDF N ALLAKQA
Sbjct: 1 PQDLXQGSMLSGCQVWWSSKGT-RGVHWRKWDLLTELKDEGGLGFKDFEIQNQALLAKQA 177
Query: 399 WRIYTQPDALWVRTLKGLYFPNSSFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGS 458
WRI PDALWV+ LK LYFP+ FLQ+ + SW W S+L G+E L + G W +G G
Sbjct: 178 WRILHNPDALWVQILKALYFPHHDFLQTTKRTGASWVWSSLLHGRELLLKQGQWQLGSGE 357
Query: 459 EVK 461
V+
Sbjct: 358 TVQ 366
>AV411435
Length = 426
Score = 112 bits (281), Expect = 2e-25
Identities = 58/139 (41%), Positives = 79/139 (56%), Gaps = 5/139 (3%)
Frame = +3
Query: 513 VFQIPLSLTQVPDKLCWPHTKDGEYTVKTGYQVA----YSLKEQARGPQPHILNDPIWNQ 568
+ Q P S+T D+L WP GEY+VK+GY+ ++L A + +W+
Sbjct: 3 IMQTPFSITGSEDRLFWPFIPSGEYSVKSGYRAVKATQFNLSNLASSSSHP--SSFLWHT 176
Query: 569 IWGIQVP*KIKMFLWKACNEALPVKASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTR 628
IWG QVP K++ FLW+A N A+PVK +L RR++ D C ICN+ E + H LL C WTR
Sbjct: 177 IWGAQVPKKVRSFLWRAANNAIPVKRNLKRRNMGRDDFCPICNKGPEDINHALLACEWTR 356
Query: 629 AVWLSSQLH-VQHLHQGSL 646
AVW L+ V H G L
Sbjct: 357 AVWFGCPLNFVVHEVHGML 413
>TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3, P1 clone:
MIG10, partial (18%)
Length = 1118
Score = 85.9 bits (211), Expect = 3e-17
Identities = 59/194 (30%), Positives = 89/194 (45%), Gaps = 6/194 (3%)
Frame = -3
Query: 631 WLSSQLHVQHLHQGSLTVKDWLTQLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQ 690
W S L G+L W + + +S + L +++ LW IWK RN FQ
Sbjct: 1116 WFGSPLQWSIEPGGNLRFDRWFVERSQFILNHSLNHEEDLATLSFTLWAIWKNRNRVIFQ 937
Query: 691 QRPPDPLGTTIRIRASM--LNGMEDNTPSPIPTQGRKPNYQAR----WDVPPPWTLKLNV 744
P+P+ T +IR N M + + T+ R N + W PP KLN
Sbjct: 936 LEEPNPVNTIFQIRHMQQDANLMSTSQSNKQTTRERNINVTSARDRSWRPPPLGVYKLNS 757
Query: 745 DATYFKETCKGSIAVVVFDEKGTNVLSHSKTIWAASPLAAEALAVREALLLAMNLELPKI 804
DA++ + ++ VV+ D G + + A SP+ +EALA+RE L+LA +L L +I
Sbjct: 756 DASWKAPSPSCTVGVVIRDNLGLLISGTASRPLAPSPIVSEALALREGLILARSLGLDRI 577
Query: 805 YLVSDCLNLVSLIR 818
SDC L+ R
Sbjct: 576 LSESDCQPLIEACR 535
>AV767121
Length = 525
Score = 84.0 bits (206), Expect = 1e-16
Identities = 52/174 (29%), Positives = 81/174 (45%), Gaps = 5/174 (2%)
Frame = +2
Query: 307 WKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKERGIHW 366
WK+K ++ G+ LI++VL A+ + +S + P ++ + F W E I
Sbjct: 11 WKQKTVSFGGRIFLIQSVLTALPLFFLSFFKLPIGVGKSCVRLMRNFLWGGSENENKIA* 190
Query: 367 KDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNSSFLQS 426
W + K K GGLG KD T N ALL K WR T+PD+LW R ++ Q
Sbjct: 191 VKWTDVCKPKELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRVIEA---------QP 343
Query: 427 GRTARNSWGWQSIL----QGKEFLKEAGL-WIIGDGSEVKVWKDKWVSDQVLQA 475
+ S W IL + + +GL ++G+G + K W + W+ +L A
Sbjct: 344 DHYSCGSSWWNDILSLCPEDADGWFSSGLKKLVGEGDQTKFWSEDWLGSGLLSA 505
>TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, partial
(14%)
Length = 592
Score = 73.2 bits (178), Expect = 2e-13
Identities = 34/59 (57%), Positives = 43/59 (72%)
Frame = -3
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRL 60
FF G LP +NET+V L+PK+ E I Q RPISCC+F+YKVI+KI+V +LKP M L
Sbjct: 191 FFDHGELPVELNETLVTLIPKIPHAEAIQQFRPISCCSFIYKVISKIIVARLKPDMVNL 15
>CN825238
Length = 692
Score = 66.2 bits (160), Expect = 2e-11
Identities = 37/134 (27%), Positives = 65/134 (47%), Gaps = 1/134 (0%)
Frame = +3
Query: 488 LILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEYTVKTGYQVAY 547
L++P+ ++WN+ + FP++ A + IPL + W +++G ++V++ Y
Sbjct: 297 LLIPNARRWNVQLIDQLFPKETAAVIKSIPLGWRTTRNSCLWRWSRNGAFSVRSAYYNIQ 476
Query: 548 SLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFRRHIAPDPVC 607
KE + W ++W VP K+K F ++ +P + +L RR IA C
Sbjct: 477 VAKEMQTSSS--AASSFPWKKLWRTPVPNKMKHFAYRVARNIIPCRENLQRRGIAVQNEC 650
Query: 608 FICN-QDEESVYHL 620
CN Q ES+ HL
Sbjct: 651 PTCNHQIPESMSHL 692
>AV780221
Length = 461
Score = 61.6 bits (148), Expect = 5e-10
Identities = 42/157 (26%), Positives = 68/157 (42%), Gaps = 8/157 (5%)
Frame = +3
Query: 320 LIKAVLQAMSAYTMSIIRFPQN-FCQAICASIARFWWRSPGKERGIHWKDWHFISKAKCK 378
LI++VL A+ + + + P F + F+ R + I W W + + K +
Sbjct: 9 LIRSVLIALPLFHLPFFKLPTGGFYTNVNN**DPFFGREVEGGKKIAWVKWSTVCRPKDE 188
Query: 379 GGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNSSFLQSGRTARNSWGWQS 438
GGLG ++ N ALL K WR+ + D LW + L Y +++ W +
Sbjct: 189 GGLGVQNLGLFNKALLGKWRWRMLKERDGLWYKVLLIKY--------KNAIPQSASSWWN 344
Query: 439 ILQGKEFLKEAGLWI-------IGDGSEVKVWKDKWV 468
L F G W+ IG+G+EVK W + W+
Sbjct: 345 DLHSTCFEDGGGGWMQRGLCRRIGEGTEVKFWNENWL 455
>TC19924
Length = 525
Score = 56.2 bits (134), Expect = 2e-08
Identities = 38/112 (33%), Positives = 57/112 (49%), Gaps = 1/112 (0%)
Frame = -2
Query: 595 SLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWLTQ 654
+L +R + P+C IC E+V H + C TR V L SQ + S L++
Sbjct: 344 NLHKRRVLNSPLCPICFAAPETVEHAIFHCNGTRLVGLGSQFQ-WNTSDASRFDSGLLSK 168
Query: 655 L-VIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRA 705
+ +RL+E E+ + + + LW IWK RN F+ + PDP GT IR +A
Sbjct: 167 IECLRLRE--ENFVQECSSLGCLLWEIWKTRNITVFEGKLPDPTGTLIRAQA 18
>BP085058
Length = 452
Score = 55.1 bits (131), Expect = 5e-08
Identities = 37/124 (29%), Positives = 54/124 (42%)
Frame = +2
Query: 260 KHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVLDKIDGWKEKLLNQAGKEV 319
+ ++ SI + D+ GKY+ P G + RV+ ++ K LLN+ +
Sbjct: 83 RDRMVSITSIPFTDNIGKYMSFPIHQGHV*KEDFVFTLDRVISRLAC*KAYLLNKPARLT 262
Query: 320 LIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKERGIHWKDWHFISKAKCKG 379
L K VL A+S Y M + PQ+ C IC + F W + GI W I K
Sbjct: 263 LAKPVLSAISMYVMQLNWLPQSICDQICTVVRHFVWEN---VNGIPLVAWETIQSRKSMV 433
Query: 380 GLGF 383
GL F
Sbjct: 434 GLIF 445
>CB828953
Length = 554
Score = 42.0 bits (97), Expect(2) = 8e-08
Identities = 18/49 (36%), Positives = 29/49 (58%)
Frame = -2
Query: 813 LVSLIRERRTDWKVEAVLRDIQD*RNLFRVCGFFWVPRTRNLCAHYVAK 861
LV R+ + ++ +++ DI + R F+ CGF W+PR N AH +AK
Sbjct: 388 LVEACRDGKGQGEIHSIVADILNLRRHFQWCGFSWIPRNSNKVAHEIAK 242
Score = 32.0 bits (71), Expect(2) = 8e-08
Identities = 19/51 (37%), Positives = 26/51 (50%)
Frame = -3
Query: 732 WDVPPPWTLKLNVDATYFKETCKGSIAVVVFDEKGTNVLSHSKTIWAASPL 782
W P P LKLNVDA++ + SI + V DE G V + ++ S L
Sbjct: 540 WRPPLPRMLKLNVDASWKASSPLCSIGIAVRDEAGLLVAGSASKAFSHSAL 388
>CB826728
Length = 509
Score = 53.1 bits (126), Expect = 2e-07
Identities = 30/98 (30%), Positives = 48/98 (48%), Gaps = 2/98 (2%)
Frame = +1
Query: 529 WPHTKDGEYTVKTGYQ-VAYSLKEQARGPQPHI-LNDPIWNQIWGIQVP*KIKMFLWKAC 586
WP + DG Y+ K+GY+ + K P + IW +W + K +W+AC
Sbjct: 214 WPGSSDGWYSSKSGYEFIRLENKSLLPSTSPAAAVPSLIWKTVWSASSLPRCKEVMWRAC 393
Query: 587 NEALPVKASLFRRHIAPDPVCFICNQDEESVYHLLLGC 624
LPV+++L RR + DP ++E+ H+LL C
Sbjct: 394 AGYLPVRSALKRRGLDVDPSSPWSGLEDEAEVHVLLTC 507
>AV412895
Length = 405
Score = 52.8 bits (125), Expect = 2e-07
Identities = 25/72 (34%), Positives = 38/72 (52%), Gaps = 3/72 (4%)
Frame = +1
Query: 563 DPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFRR---HIAPDPVCFICNQDEESVYH 619
DP++N IW + P IK F+W+ + + +L +R H A D +C +C EES H
Sbjct: 64 DPVFN*IWMVPAPLNIKAFVWRVLLGRIQTRDNLLKRQVIHNALDAICPLCGLAEESGSH 243
Query: 620 LLLGCPWTRAVW 631
LL C + +W
Sbjct: 244 LLFSCAESMLIW 279
>AV428268
Length = 429
Score = 51.6 bits (122), Expect = 5e-07
Identities = 24/69 (34%), Positives = 34/69 (48%)
Frame = +3
Query: 563 DPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFRRHIAPDPVCFICNQDEESVYHLLL 622
D I+ +W ++VP WK + K +L RR + VC +C+ DEES HLL
Sbjct: 60 DNIFKLLWSVKVPSNAIALSWKVLINRVQTKVNLNRRGVVISNVCPLCSLDEESTDHLLF 239
Query: 623 GCPWTRAVW 631
CP +W
Sbjct: 240 SCPIV*RIW 266
>AV426428
Length = 413
Score = 51.2 bits (121), Expect = 7e-07
Identities = 23/54 (42%), Positives = 31/54 (56%)
Frame = +3
Query: 583 WKACNEALPVKASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQL 636
W+AC LPVKASL R +A DP+C C ++ H +L CP +W +S L
Sbjct: 96 WRACLGVLPVKASLHARGLAEDPICPRCLAAPKTADHAVLFCP*VHPIWFASSL 257
>AV406756
Length = 428
Score = 51.2 bits (121), Expect = 7e-07
Identities = 26/63 (41%), Positives = 35/63 (55%)
Frame = +1
Query: 795 LAMNLELPKIYLVSDCLNLVSLIRERRTDWKVEAVLRDIQD*RNLFRVCGFFWVPRTRNL 854
LA +L+L K SDC L+SL+ + DW + A+L DIQ+ R+ F WV R N
Sbjct: 1 LASSLQLGKCIFASDCKELISLVTSFKMDWSISAILLDIQELRS*LSEPKFVWVGRGSNY 180
Query: 855 CAH 857
AH
Sbjct: 181 QAH 189
>AV780250
Length = 494
Score = 49.3 bits (116), Expect = 3e-06
Identities = 32/110 (29%), Positives = 49/110 (44%), Gaps = 1/110 (0%)
Frame = +3
Query: 525 DKLCWPHTKDGEYTVKTGYQVAYSLKEQARGPQPHILN-DPIWNQIWGIQVP*KIKMFLW 583
D L WPHT +GEYT K+GY V + A L P+ +W + + +
Sbjct: 174 DVLAWPHTSNGEYTSKSGYAVVRNKVVHALPSSSTALQVSPV---VWKAPTLPRCRELIG 344
Query: 584 KACNEALPVKASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLS 633
+A + LP +++L R + D +C C ES +LL P W +
Sbjct: 345 RAFLKILPTRSALRARGMDVDELCPSCETHRESPAQVLLFVPVVSKWWFA 494
>TC19933
Length = 472
Score = 47.8 bits (112), Expect = 8e-06
Identities = 22/47 (46%), Positives = 30/47 (63%)
Frame = -3
Query: 566 WNQIWGIQVP*KIKMFLWKACNEALPVKASLFRRHIAPDPVCFICNQ 612
W+ I +V K+K FLWK C+ A+P+K +L RR A D C IC+Q
Sbjct: 143 WSSI*KAKVSNKVKHFLWKLCHNAIPLKGNLRRRGCATDGKCPICDQ 3
>AV412633
Length = 405
Score = 44.3 bits (103), Expect(2) = 1e-05
Identities = 19/42 (45%), Positives = 28/42 (66%)
Frame = -3
Query: 788 AVREALLLAMNLELPKIYLVSDCLNLVSLIRERRTDWKVEAV 829
A+REAL+LA NL+L K SDC L+SL+ + DW + ++
Sbjct: 400 ALREALVLASNLQLDKCIFASDCKELISLVTSFKMDWSISSM 275
Score = 22.3 bits (46), Expect(2) = 1e-05
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = -2
Query: 832 DIQD*RNLFRVCGFFWVPRTRNLCAHYVAKQH 863
DIQ+ R+ F WV R N AH +A+ +
Sbjct: 275 DIQELRSRLSEPKFVWVGRGSNQQAHDLAQSN 180
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,075,510
Number of Sequences: 28460
Number of extensions: 288117
Number of successful extensions: 1430
Number of sequences better than 10.0: 71
Number of HSP's better than 10.0 without gapping: 1404
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1417
length of query: 863
length of database: 4,897,600
effective HSP length: 98
effective length of query: 765
effective length of database: 2,108,520
effective search space: 1613017800
effective search space used: 1613017800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0111a.1


