

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.15
(184 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
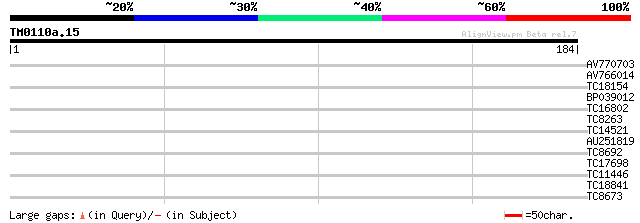
AV770703 35 0.008
AV766014 29 0.55
TC18154 UP|Q40192 (Q40192) RAB11B, complete 28 0.72
BP039012 27 2.7
TC16802 similar to GB|AAP31963.1|30387595|BT006619 At1g01300 {Ar... 26 3.6
TC8263 homologue to UP|Q41038 (Q41038) Type II chlorophyll a/b b... 26 3.6
TC14521 similar to UP|O04894 (O04894) Transaldolase , partial (... 26 4.7
AU251819 25 6.1
TC8692 UP|Q7XY82 (Q7XY82) Photosystem II 12kD extrinsic protein,... 25 6.1
TC17698 25 6.1
TC11446 similar to GB|AAM51367.1|21436395|AY117292 IAA-amino aci... 25 8.0
TC18841 25 8.0
TC8673 similar to UP|Q9LHL9 (Q9LHL9) Kinesin (Centromere protein... 25 8.0
>AV770703
Length = 539
Score = 35.0 bits (79), Expect = 0.008
Identities = 15/49 (30%), Positives = 24/49 (48%)
Frame = +2
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKE 61
H + T R+Y FL++ ++YF I D S L FL +R ++
Sbjct: 197 HRWYFETAGKRSYGFLMEDGYVYFTIVDEGLGNSVVLRFLEHVRDEFRK 343
>AV766014
Length = 391
Score = 28.9 bits (63), Expect = 0.55
Identities = 16/33 (48%), Positives = 19/33 (57%)
Frame = -2
Query: 25 YTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRS 57
YTF + F FAIF H+HT+ FLN RS
Sbjct: 114 YTFHV*YFFYKFAIFTHKHTR*IDFPFLNW*RS 16
>TC18154 UP|Q40192 (Q40192) RAB11B, complete
Length = 1183
Score = 28.5 bits (62), Expect = 0.72
Identities = 18/60 (30%), Positives = 30/60 (50%), Gaps = 1/60 (1%)
Frame = -2
Query: 11 STHSLFSHTVNNRTYTFLIDPPFIY-FAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDS 69
+T S+ S T + +F PFI F F H +T +Q F+N+ + E L+ +N +
Sbjct: 372 NT*SILSCTFFSCRCSFSASTPFIS*FIFFPHGNTSNQKTQFVNKRKCRNMEKLEKMNQT 193
>BP039012
Length = 582
Score = 26.6 bits (57), Expect = 2.7
Identities = 18/58 (31%), Positives = 28/58 (48%), Gaps = 8/58 (13%)
Frame = +3
Query: 44 TKSQTLTFLNRIRSSLKETLDSV---NDSTAPPPLSLQTPFDLI-----LQEILHLDD 93
+ S + T + R SL T ++ NDST PPP +P + LQE+ H+ +
Sbjct: 192 SSSSSTTSFTQWRFSLPTTPNTTTTQNDSTTPPPPPSPSPSTITITIPNLQELFHVSE 365
>TC16802 similar to GB|AAP31963.1|30387595|BT006619 At1g01300 {Arabidopsis
thaliana;}, partial (28%)
Length = 812
Score = 26.2 bits (56), Expect = 3.6
Identities = 12/24 (50%), Positives = 14/24 (58%)
Frame = +1
Query: 57 SSLKETLDSVNDSTAPPPLSLQTP 80
S KETL +N S+ PPP TP
Sbjct: 310 SGSKETLPELNPSSPPPPQLSTTP 381
>TC8263 homologue to UP|Q41038 (Q41038) Type II chlorophyll a/b binding
protein from photosystem I precursor, partial (98%)
Length = 1199
Score = 26.2 bits (56), Expect = 3.6
Identities = 17/39 (43%), Positives = 21/39 (53%)
Frame = +1
Query: 39 FDHRHTKSQTLTFLNRIRSSLKETLDSVNDSTAPPPLSL 77
FDH+HT ++ LT +RI S E L S P PL L
Sbjct: 1 FDHKHTSAKKLTKPHRITVSRGEGL-SWLQRVLPLPLQL 114
>TC14521 similar to UP|O04894 (O04894) Transaldolase , partial (86%)
Length = 1695
Score = 25.8 bits (55), Expect = 4.7
Identities = 18/60 (30%), Positives = 29/60 (48%)
Frame = +2
Query: 119 SATVIANRTSAKGLAAAAATELVELEKDAVEEALVGSAIQIEGFRRVWLDLRSRLSLLHE 178
S T+ +N + A+G+ A L+K V+ + VGS +++EG S L L E
Sbjct: 1145 SRTIDSNASEAEGIYNA-------LQKVGVDWSFVGSQLELEGVESFKKSFDSLLDSLQE 1303
>AU251819
Length = 368
Score = 25.4 bits (54), Expect = 6.1
Identities = 14/48 (29%), Positives = 26/48 (54%)
Frame = +1
Query: 52 LNRIRSSLKETLDSVNDSTAPPPLSLQTPFDLILQEILHLDDGNRSPP 99
+N+ S+K+ S + S++PPP SL + I +++ G+ PP
Sbjct: 124 INKDSHSIKK---SPSSSSSPPPPSLSSSSSSISSSLVNSVAGSNKPP 258
>TC8692 UP|Q7XY82 (Q7XY82) Photosystem II 12kD extrinsic protein, partial
(9%)
Length = 687
Score = 25.4 bits (54), Expect = 6.1
Identities = 19/56 (33%), Positives = 27/56 (47%)
Frame = -3
Query: 35 YFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDSTAPPPLSLQTPFDLILQEILH 90
+ A HRHT S TL FL + +L ++N P QT F+ IL + +H
Sbjct: 478 WHARISHRHTNSPTLLFLQGKKYNLLR--KNLNKQHPLP----QTQFEGILLDDIH 329
>TC17698
Length = 504
Score = 25.4 bits (54), Expect = 6.1
Identities = 10/29 (34%), Positives = 19/29 (65%)
Frame = -3
Query: 33 FIYFAIFDHRHTKSQTLTFLNRIRSSLKE 61
F+YF++FDH + +Q+L + + + KE
Sbjct: 325 FVYFSLFDHEFS-NQSLALAHLLETETKE 242
>TC11446 similar to GB|AAM51367.1|21436395|AY117292 IAA-amino acid hydrolase
{Arabidopsis thaliana;}, partial (8%)
Length = 400
Score = 25.0 bits (53), Expect = 8.0
Identities = 14/46 (30%), Positives = 23/46 (49%)
Frame = +2
Query: 55 IRSSLKETLDSVNDSTAPPPLSLQTPFDLILQEILHLDDGNRSPPA 100
I +S +L + + TA ++ P D + LHL GN +PP+
Sbjct: 152 ITASEHASLHTSHPPTASVLRRVKFPGDTVSLSSLHLRSGNNTPPS 289
>TC18841
Length = 568
Score = 25.0 bits (53), Expect = 8.0
Identities = 15/41 (36%), Positives = 20/41 (48%)
Frame = -1
Query: 50 TFLNRIRSSLKETLDSVNDSTAPPPLSLQTPFDLILQEILH 90
T R+ SL +T + + P P+ T LIL EILH
Sbjct: 409 TCQERLHVSLGKTFEIFLTTVFPFPVRTWTSSPLILSEILH 287
>TC8673 similar to UP|Q9LHL9 (Q9LHL9) Kinesin (Centromere protein) like
heavy chain-like protein, partial (11%)
Length = 819
Score = 25.0 bits (53), Expect = 8.0
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = +2
Query: 127 TSAKGLAAAAATELVELEKDAVE 149
T AKGLA+AAA EL L ++ +
Sbjct: 35 TYAKGLASAAAVELKALSEEVAK 103
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,070,188
Number of Sequences: 28460
Number of extensions: 41164
Number of successful extensions: 256
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 253
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 256
length of query: 184
length of database: 4,897,600
effective HSP length: 85
effective length of query: 99
effective length of database: 2,478,500
effective search space: 245371500
effective search space used: 245371500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0110a.15


