

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
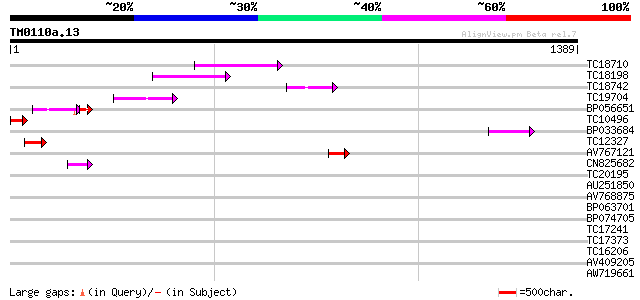
Query= TM0110a.13
(1389 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18710 129 3e-30
TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, p... 89 5e-18
TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%) 59 7e-09
TC19704 weakly similar to UP|Q9SZ87 (Q9SZ87) RNA-directed DNA po... 57 3e-08
BP056651 45 7e-08
TC10496 52 6e-07
BP033684 51 1e-06
TC12327 50 2e-06
AV767121 47 3e-05
CN825682 43 4e-04
TC20195 similar to UP|Q8IQS0 (Q8IQS0) CG32193-PA, partial (5%) 35 0.11
AU251850 35 0.11
AV768875 31 1.5
BP063701 31 1.5
BP074705 30 3.4
TC17241 29 4.4
TC17373 similar to GB|AAO63356.1|28950865|BT005292 At4g16580 {Ar... 29 5.8
TC16206 29 5.8
AV409205 29 5.8
AW719661 29 5.8
>TC18710
Length = 843
Score = 129 bits (324), Expect = 3e-30
Identities = 77/217 (35%), Positives = 111/217 (50%), Gaps = 3/217 (1%)
Frame = +2
Query: 454 GTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRP 513
G DG N F+++ +VK+ V F ++ +N T ++LIPK P +++ FRP
Sbjct: 188 GMDGMNGLFYQKNGDVVKDSVT*AILAFLERAEIPNEINETLVTLIPKVPHPESINQFRP 367
Query: 514 ISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGG 573
IS YK+ISK+ +RL+ P LIS Q F QGRQI D +LI E +N+
Sbjct: 368 ISGCTFLYKVISKIFVARLKDAMPDLISPMQSGFIQGRQIQDNLLIVQEAFHAINRPGAL 547
Query: 574 G---FLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGF 630
G ++KLD K YD ++W FL L F W+ I VS + + +NG
Sbjct: 548 GRNHSIIKLDMNKAYDRVEWKFLESSLLAFGFSTNWVKMIMILVSGVSYNYKINGVVGPK 727
Query: 631 FEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSN 667
++GLRQGDP SP+LF + LS L+ + N
Sbjct: 728 LLPQRGLRQGDPFSPYLFLFTMEVLSLLIQNSFNMGN 838
>TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, partial
(14%)
Length = 592
Score = 89.0 bits (219), Expect = 5e-18
Identities = 64/194 (32%), Positives = 97/194 (49%), Gaps = 2/194 (1%)
Frame = -3
Query: 350 RVRWLREGDRNTSFFHSISKIRKSKKLISQLV-VGGVSIEDPNMIKDAIFSHFQSFFTSE 408
R++ L+ GD NT FFH+ + R+ I ++ G+ +E + +A ++F ++S
Sbjct: 590 RIK*LK*GDHNTRFFHASTIQRRDFNRILKIKDAHGIWVEGQKKVNEAAVNYFSDIYSSS 411
Query: 409 NRLRPKIRCANLSRL-SQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFW 467
+ A + +L + L Q S EI E + S +APG DG N F++Q
Sbjct: 410 PLQYLRECLAPIPKLVTDRLNHKLCAQVSTDEIKEAVFSMGDLKAPGMDGINGLFYQQNR 231
Query: 468 GIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKV 527
IVK DV DF G+L LN T ++LIPK + FRPIS YK+ISK+
Sbjct: 230 EIVKNDVNNAILDFFDHGELPVELNETLVTLIPKIPHAEAIQQFRPISCCSFIYKVISKI 51
Query: 528 LASRLQVVTPLLIS 541
+ +RL+ LIS
Sbjct: 50 IVARLKPDMVNLIS 9
>TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%)
Length = 912
Score = 58.5 bits (140), Expect = 7e-09
Identities = 39/126 (30%), Positives = 66/126 (51%), Gaps = 2/126 (1%)
Frame = -3
Query: 679 LNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQR-- 736
++HL FAD+ ++F I ++ N L F SGL N KS + + V+ET +
Sbjct: 373 ISHLCFADNLMVFSNGYLESIAIINNVLHIFQHLSGLTPNPAKSEVF---ILVKETTKNQ 203
Query: 737 VAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMK 796
+ M G+ G+LP+ LG P +AS + ++D+ K + W + +S +GRL L+
Sbjct: 202 ITNMLGYKEGKLPVR*LGVPQLTTKLKASDCKVLVDRKPSKLQHWTGRSLSYAGRLQLIN 23
Query: 797 ATLCSV 802
+ L S+
Sbjct: 22 SILFSM 5
>TC19704 weakly similar to UP|Q9SZ87 (Q9SZ87) RNA-directed DNA
polymerase-like protein, partial (3%)
Length = 530
Score = 56.6 bits (135), Expect = 3e-08
Identities = 39/158 (24%), Positives = 72/158 (44%), Gaps = 1/158 (0%)
Frame = -3
Query: 254 EKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVRINTIEADLQVTMDR 313
E + W ++ T L K K + EW S + I ++ L+ +
Sbjct: 519 ETIKKAWDNTSETASSHCSNLSGKTTNTKLALTEWHHQNFSRADKEIAELKNQLEHLQNY 340
Query: 314 LASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKS 373
S E E+++ ++ L+ LW R+EE W Q++R++WL+ GD+NT FFH+ + R+
Sbjct: 339 SNSGEDWQEIKQRKMR-LKDLW---RQEELFWSQRSRIKWLKSGDKNTKFFHASTVQRRE 172
Query: 374 KKLISQLV-VGGVSIEDPNMIKDAIFSHFQSFFTSENR 410
+ I ++ + G + I + + + S NR
Sbjct: 171 RNRIERIKDMQGTWVRSQLAINRDVPDFYPDIYKSTNR 58
>BP056651
Length = 564
Score = 45.1 bits (105), Expect(2) = 7e-08
Identities = 34/127 (26%), Positives = 52/127 (40%), Gaps = 6/127 (4%)
Frame = +1
Query: 56 MDYVSVDAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVD 115
MD+ + A AGGL+ +W P+ + +E F+G L+ I+NVY P
Sbjct: 73 MDWRMIPAKERAGGLLCVWKPSVFTVEECFLGEGFLG--LKGKWKGVSCVIINVYSPCSR 246
Query: 116 SERYDFFNLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQ------MGVESSFSDFVAD 169
E+ + L M GDFN++ ER G G F+DF+ +
Sbjct: 247 GEKRQLWEDLKALRRRLEEDIWCMAGDFNSVRRSEERRGTHGEGSSGAGEMEEFNDFIDE 426
Query: 170 SNLIDLP 176
N + P
Sbjct: 427 WNWMTSP 447
Score = 29.6 bits (65), Expect(2) = 7e-08
Identities = 16/31 (51%), Positives = 20/31 (63%)
Frame = +3
Query: 172 LIDLPLQSDKFTWYSSRAGGLWSRIDRWLVS 202
L D+PL K+TW S G SR+DR+LVS
Sbjct: 432 LDDIPLVGRKYTWVRSN-GKARSRLDRFLVS 521
>TC10496
Length = 562
Score = 52.0 bits (123), Expect = 6e-07
Identities = 23/41 (56%), Positives = 29/41 (70%)
Frame = +3
Query: 3 WVSWNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSR 43
W WNVRGLG + KR A + L +KPE+ L+QESKLD +R
Sbjct: 435 WAFWNVRGLGSSTKREAAKKILLKIKPEVCLLQESKLDENR 557
>BP033684
Length = 557
Score = 51.2 bits (121), Expect = 1e-06
Identities = 42/112 (37%), Positives = 53/112 (46%)
Frame = -2
Query: 1174 FSGTNLSARKRYGISMC*GLFGG*KPIGRTAPSIRINSAQTSAISKFLGRFIFLDQRHGI 1233
FS N KR+G S+ * FGG K GR AP+I NS TSA + D + G
Sbjct: 460 FSRAN*LLLKRFGTSIS*QFFGGSKQFGRNAPTISTNSTGTSACYPSRN*YPNKD*KGGH 281
Query: 1234 PLQLEP*SLMLMVPLRGIRGLVAWGVSFVITRIVSLVIFQLMLGLDGLLRLR 1285
PL S M L+ IR LV GV +V T L Q + GL++L+
Sbjct: 280 PLAEVFSSSTSMERLKVIRALVV*GVFYVTTEAWFLASSQSIWVTGGLMKLK 125
>TC12327
Length = 568
Score = 50.1 bits (118), Expect = 2e-06
Identities = 22/54 (40%), Positives = 37/54 (67%)
Frame = +1
Query: 36 ESKLDSSRVKTVEAFANAVKMDYVSVDAVGSAGGLISLWNPASYAMESAVKNVR 89
ESKLD +R KT+ ++A ++ M++ V A+G AGGL+SLW +++ + K+ R
Sbjct: 1 ESKLDENRKKTIVSWAKSIGMEFEFVPAIGVAGGLVSLWRKSTFKVVQVKKHQR 162
>AV767121
Length = 525
Score = 46.6 bits (109), Expect = 3e-05
Identities = 19/51 (37%), Positives = 33/51 (64%)
Frame = +2
Query: 781 WNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQGG 831
W K +S GR+ L+++ L ++PLF ++ FK P+ V +++R+FL GG
Sbjct: 11 WKQKTVSFGGRIFLIQSVLTALPLFFLSFFKLPIGVGKSCVRLMRNFLWGG 163
>CN825682
Length = 626
Score = 42.7 bits (99), Expect = 4e-04
Identities = 27/67 (40%), Positives = 40/67 (59%), Gaps = 5/67 (7%)
Frame = -3
Query: 141 GDFNAILNEGERIGIQMGVESS---FSDFVADSNLIDLPLQSDKFTWYSSRAGGLWS--R 195
GDFN IL+ E+ G + + F DF+ +NL+DL L+ +FTW S+ GL + R
Sbjct: 288 GDFNEILHHQEKEGARDQDPTRIQVFRDFLDGANLMDLELKGCRFTWLSNLRNGLVTKER 109
Query: 196 IDRWLVS 202
+DR LV+
Sbjct: 108 LDRVLVN 88
>TC20195 similar to UP|Q8IQS0 (Q8IQS0) CG32193-PA, partial (5%)
Length = 591
Score = 34.7 bits (78), Expect = 0.11
Identities = 15/28 (53%), Positives = 18/28 (63%)
Frame = +2
Query: 3 WVSWNVRGLGRAEKRRAVVEALSGLKPE 30
W +WNVR LG KR V +A+ LKPE
Sbjct: 98 WFTWNVRRLGGTTKREIVKKAIPPLKPE 181
>AU251850
Length = 343
Score = 34.7 bits (78), Expect = 0.11
Identities = 22/75 (29%), Positives = 36/75 (47%), Gaps = 3/75 (4%)
Frame = +2
Query: 4 VSWNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKL---DSSRVKTVEAFANAVKMDYVS 60
+SWN RGLG A+ + P++V + E+KL ++ ++ +N + +D V
Sbjct: 47 LSWNCRGLGSPRAVEALQRLVRKEIPQLVFLMETKLKKFETEKISLGFGLSNCLIVDCVG 226
Query: 61 VDAVGSAGGLISLWN 75
D GGL WN
Sbjct: 227 -DEKERKGGLAMWWN 268
>AV768875
Length = 232
Score = 30.8 bits (68), Expect = 1.5
Identities = 18/50 (36%), Positives = 25/50 (50%), Gaps = 2/50 (4%)
Frame = +1
Query: 141 GDFNAILNEGERIGIQMGVESS--FSDFVADSNLIDLPLQSDKFTWYSSR 188
GD N IL + G + + F+ + D NLIDL + KFTW+ R
Sbjct: 70 GDMNEILFPSQVRGGEFHPNRAQRFATVLDDCNLIDLGMVGGKFTWFRKR 219
>BP063701
Length = 547
Score = 30.8 bits (68), Expect = 1.5
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = -1
Query: 1 RRWVSWNVRGLGRAEKRRA 19
R+WV WN+RGLG KR++
Sbjct: 121 RKWVIWNMRGLGGLXKRKS 65
>BP074705
Length = 379
Score = 29.6 bits (65), Expect = 3.4
Identities = 15/42 (35%), Positives = 25/42 (58%)
Frame = -1
Query: 4 VSWNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVK 45
V W+ RGL +A+ + V AL+ LK +I V+ ++ +VK
Sbjct: 172 VVWDARGLAQAKAKDTKVRALTRLKKKIYYVKNGQIYFYQVK 47
>TC17241
Length = 605
Score = 29.3 bits (64), Expect = 4.4
Identities = 17/44 (38%), Positives = 25/44 (56%)
Frame = -2
Query: 634 EKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGL 677
+KG + G F F+LCV G+ C++NQ ++ FSG GL
Sbjct: 295 QKGRQDG-----F*FDLCVLGIMCIMNQ-INSYTFSGGDTSEGL 182
>TC17373 similar to GB|AAO63356.1|28950865|BT005292 At4g16580 {Arabidopsis
thaliana;}, partial (15%)
Length = 857
Score = 28.9 bits (63), Expect = 5.8
Identities = 16/51 (31%), Positives = 25/51 (48%)
Frame = +3
Query: 37 SKLDSSRVKTVEAFANAVKMDYVSVDAVGSAGGLISLWNPASYAMESAVKN 87
S L +R+ + +NAV D D V S GG+++L PA + +N
Sbjct: 252 SNLKFNRISMATSASNAVLGDVYVDDLVSSCGGVLNLSKPAYAYFKERARN 404
>TC16206
Length = 557
Score = 28.9 bits (63), Expect = 5.8
Identities = 12/36 (33%), Positives = 20/36 (55%)
Frame = +2
Query: 767 WEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSV 802
WE +L+ +++ RR N F L R L + T C++
Sbjct: 137 WETLLEAMKKARRLNNHVFSMLRARCPLEEKTACAI 244
>AV409205
Length = 426
Score = 28.9 bits (63), Expect = 5.8
Identities = 10/23 (43%), Positives = 17/23 (73%)
Frame = -2
Query: 766 FWEPMLDKLRRKRRTWNSKFISL 788
+WE M+ ++R+RR W+SK +L
Sbjct: 125 YWENMMQIIKRRRRMWHSKHRNL 57
>AW719661
Length = 263
Score = 28.9 bits (63), Expect = 5.8
Identities = 20/59 (33%), Positives = 33/59 (55%), Gaps = 4/59 (6%)
Frame = -3
Query: 324 RKNRLTILEG-LWQEYRKEEYRWRQK-ARVRW--LREGDRNTSFFHSISKIRKSKKLIS 378
+KNR ++L+G +W R+ RW ++ A W REG+R SI+ +K + L+S
Sbjct: 168 KKNRSSVLDGGIWTGRRR*PSRWEERMASAGWKDWREGERR---MRSIAAEKKPENLMS 1
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.334 0.147 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,391,684
Number of Sequences: 28460
Number of extensions: 401358
Number of successful extensions: 3088
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 3032
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3082
length of query: 1389
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1287
effective length of database: 1,994,680
effective search space: 2567153160
effective search space used: 2567153160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0110a.13


