

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.10
(1408 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
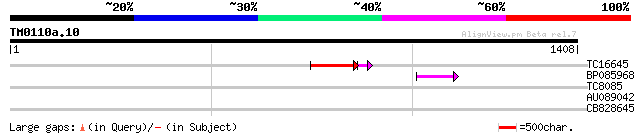
Score E
Sequences producing significant alignments: (bits) Value
TC16645 122 9e-32
BP085968 53 4e-07
TC8085 similar to GB|BAB03033.1|11994717|AP001300 AtDMC1 (meioti... 31 1.5
AU089042 29 4.5
CB828645 29 5.9
>TC16645
Length = 596
Score = 122 bits (307), Expect(2) = 9e-32
Identities = 55/119 (46%), Positives = 79/119 (66%)
Frame = +1
Query: 747 MDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGELYYLRMLL 806
M+AN KY GR LTY EFP F+Y + W P+++ S+GR+ F+ PG GE YY+R+LL
Sbjct: 7 MNANKKYQEGRNLTYAEFPSEFLYHQHTTKWVPKQKRFSLGRLPFIAPGMGENYYMRVLL 186
Query: 807 NVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGGAYTRALF 865
+Q+GC ++ +++V V+ TF ACEA+GLL+DDRE++DGI + G R LF
Sbjct: 187 TMQKGCDSFKSIKTVKGVVYPTFHDACEAMGLLEDDREYVDGISISSEFGSGTQLRKLF 363
Score = 32.7 bits (73), Expect(2) = 9e-32
Identities = 13/37 (35%), Positives = 20/37 (53%)
Frame = +2
Query: 865 FVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKHN 901
F L+ N + P VW ++W +L DGI D ++ N
Sbjct: 362 FTRMLMTNLISRPQEVWRKSWSLLCDGILYDRRKTLN 472
>BP085968
Length = 341
Score = 52.8 bits (125), Expect = 4e-07
Identities = 39/105 (37%), Positives = 52/105 (49%)
Frame = +1
Query: 1010 *SV*GINFCSI*G*WWVFLCQWTWWYGENILVENFNF*VKIRKKKSYSMLLQVA*LLSYF 1069
*SV* N S+ W V W W +G+NI VE F KI + + ++ L
Sbjct: 25 *SV*TNNEFSLVRRWRVLFSIWIWRFGKNICVEYMVFRFKITRTHGFERGIKWYCFLVVA 204
Query: 1070 QVVGLHILFFAFL*MLMRIQYVESFKVVQKLNYFSQPVLLFGMKH 1114
GL I F F+ LM Q V S K +++LNYF + V +GMKH
Sbjct: 205 WRKGLLIQGFLFIYQLMIYQPVISSKDLKRLNYFKRQVRSYGMKH 339
>TC8085 similar to GB|BAB03033.1|11994717|AP001300 AtDMC1 (meiotic
recombination protein)-like protein {Arabidopsis
thaliana;} , partial (8%)
Length = 500
Score = 30.8 bits (68), Expect = 1.5
Identities = 15/31 (48%), Positives = 18/31 (57%)
Frame = +1
Query: 874 MCNPLHVWNETWHVLADGIACDLQRKHNNRG 904
M NP HVWN TW +LA+ I +H RG
Sbjct: 115 MTNPDHVWNITWKLLANDI------QHEYRG 189
>AU089042
Length = 191
Score = 29.3 bits (64), Expect = 4.5
Identities = 21/47 (44%), Positives = 24/47 (50%)
Frame = +1
Query: 1339 FSNKVLMLC*CEI*IYLLVCAMALD*LLHIFDQMLLEGLLFLALMLE 1385
F KV +LC C I +LLV A D LL QM L L L+LE
Sbjct: 7 FLXKVYLLCSCXIXTFLLVYATVQDLLLTTLVQM*LVLPFSLGLILE 147
>CB828645
Length = 527
Score = 28.9 bits (63), Expect = 5.9
Identities = 14/45 (31%), Positives = 25/45 (55%)
Frame = +3
Query: 1187 C*HYHKTCVCFLIQMLQMLMILENLPSGC*MLEMVTWESIMMVNL 1231
C HY K+C C + +L++L++ NL C +L M+ + N+
Sbjct: 312 CSHYSKSCFCVKLGVLEVLLLQLNL---CLILTMLITMKCFLPNI 437
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.353 0.158 0.566
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,928,179
Number of Sequences: 28460
Number of extensions: 415228
Number of successful extensions: 5367
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 3990
Number of HSP's successfully gapped in prelim test: 132
Number of HSP's that attempted gapping in prelim test: 1205
Number of HSP's gapped (non-prelim): 4376
length of query: 1408
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1306
effective length of database: 1,994,680
effective search space: 2605052080
effective search space used: 2605052080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0110a.10


