

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
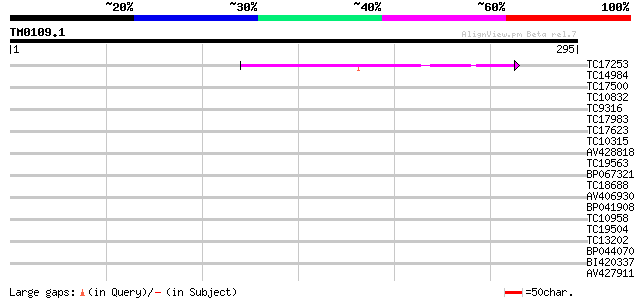
Query= TM0109.1
(295 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17253 similar to UP|Q8SKU2 (Q8SKU2) Tic62 protein precursor, p... 50 3e-07
TC14984 similar to GB|AAM62447.1|21553354|AY085214 glycine-rich ... 36 0.009
TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase t... 32 0.13
TC10832 weakly similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-ric... 31 0.22
TC9316 similar to UP|Q94F86 (Q94F86) Early light inducible prote... 31 0.22
TC17983 similar to UP|Q9XEX5 (Q9XEX5) Cytidine deaminase , part... 31 0.22
TC17623 similar to UP|Q93YH8 (Q93YH8) HMG I/Y like protein, part... 31 0.22
TC10315 similar to UP|P93166 (P93166) SCOF-1, partial (42%) 31 0.28
AV428818 30 0.48
TC19563 29 0.83
BP067321 29 0.83
TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%) 29 0.83
AV406930 29 1.1
BP041908 29 1.1
TC10958 similar to PIR|T47938|T47938 RING finger protein - Arabi... 28 1.4
TC19504 similar to UP|Q9SV89 (Q9SV89) Isoleucine-tRNA ligase-lik... 28 1.4
TC13202 28 1.4
BP044070 28 1.8
BI420337 27 3.1
AV427911 26 7.0
>TC17253 similar to UP|Q8SKU2 (Q8SKU2) Tic62 protein precursor, partial (9%)
Length = 793
Score = 50.4 bits (119), Expect = 3e-07
Identities = 49/149 (32%), Positives = 73/149 (48%), Gaps = 4/149 (2%)
Frame = +2
Query: 121 VTAEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTED 180
V A E +S +NVA+ TE +S ++VAA TE ES +V TE ES ++V S T +
Sbjct: 125 VAAPTEVESTGDNVAASTEVESTGDNVAASTEVESTGSNVAAPTEVESTGDNVAASTTVE 304
Query: 181 --VGEM-EKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEK 237
VG + EKN S ++ ++ + PP P AP + +SA EM+
Sbjct: 305 STVGNVDEKNPSPDSNSSNSPYPVYSDFKPPTSPSPN----APNTSTSKSAVDVVPEMDA 472
Query: 238 NESLTTDVASLSLGDDAL-ASPPPQPKTR 265
S A LS+ D++ P+PK+R
Sbjct: 473 ASS--NGPAQLSIADESKDEEHAPEPKSR 553
>TC14984 similar to GB|AAM62447.1|21553354|AY085214 glycine-rich RNA binding
protein 7 {Arabidopsis thaliana;} , partial (19%)
Length = 483
Score = 35.8 bits (81), Expect = 0.009
Identities = 30/92 (32%), Positives = 39/92 (41%)
Frame = +2
Query: 189 SLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASL 248
S TT ASLSL SPPPQP P P + + S ++ A+
Sbjct: 92 SPTTAPASLSL------SPPPQPSN-----PPPPPPLPPPEMPSSPATLKSSPSSATATA 238
Query: 249 SLGDDALASPPPQPKTRRKLAAPQPAAPQPAV 280
+ + SPPP P T K + P P P+ AV
Sbjct: 239 APSTKSTTSPPP-PSTL*KSSTPAPTPPRAAV 331
Score = 28.5 bits (62), Expect = 1.4
Identities = 22/52 (42%), Positives = 26/52 (49%), Gaps = 1/52 (1%)
Frame = +2
Query: 240 SLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRS-ARIKTTP 290
S TT ASLSL SPPPQP P P P P + S A +K++P
Sbjct: 92 SPTTAPASLSL------SPPPQPSN----PPPPPPLPPPEMPSSPATLKSSP 217
>TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase type 2C,
complete
Length = 1325
Score = 32.0 bits (71), Expect = 0.13
Identities = 25/75 (33%), Positives = 31/75 (41%), Gaps = 8/75 (10%)
Frame = -3
Query: 207 PPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLS----LGDDALASPPPQP 262
PPPQ T P P +RR + +K S T +LS SPPP+P
Sbjct: 561 PPPQADTAAPGTPPPPELRR-------DRQKQRSSRTSSRTLSPRSPRPPSPSCSPPPKP 403
Query: 263 ----KTRRKLAAPQP 273
K RR +AP P
Sbjct: 402 NRTAKRRRLASAPDP 358
>TC10832 weakly similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-rich protein,
partial (30%)
Length = 539
Score = 31.2 bits (69), Expect = 0.22
Identities = 24/90 (26%), Positives = 41/90 (44%), Gaps = 3/90 (3%)
Frame = +3
Query: 206 SPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTR 265
SPP + + P+ RS+ ++ + ++ S ++ +S S G SPPPQ +
Sbjct: 129 SPPSGSVSGSSSRSSSPSSSRSSS-RSRSLSRSRSFSSSSSSPSRGGSRSRSPPPQRRKS 305
Query: 266 RKLAAPQPAAPQPAVRR---SARIKTTPKK 292
AA + +P P+ R S R P+K
Sbjct: 306 PVEAARRGRSPPPSKRASPPSKRASPPPRK 395
>TC9316 similar to UP|Q94F86 (Q94F86) Early light inducible protein,
partial (84%)
Length = 831
Score = 31.2 bits (69), Expect = 0.22
Identities = 21/95 (22%), Positives = 42/95 (44%), Gaps = 6/95 (6%)
Frame = +1
Query: 195 ASLSLGDDALASPPPQPKTRRKL------VAPQPAVRRSARIKTGEMEKNESLTTDVASL 248
+S++ ++SP + TR ++ P++RR+A ++ M + E + +
Sbjct: 106 SSMAAMQSIISSPMIRMTTRSRVNQFTTPAVYMPSLRRNATLRVRSMAEGEPKEQPIEPV 285
Query: 249 SLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRS 283
+ SPPP P P+P AP+ + + S
Sbjct: 286 TSASPVTPSPPPPP--------PKPRAPKVSTKFS 366
>TC17983 similar to UP|Q9XEX5 (Q9XEX5) Cytidine deaminase , partial (42%)
Length = 519
Score = 31.2 bits (69), Expect = 0.22
Identities = 17/61 (27%), Positives = 32/61 (51%), Gaps = 4/61 (6%)
Frame = +2
Query: 238 NESLTTDVASLSL--GDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIKTT--PKKF 293
+ES + +AS S + +S P P T ++++ P P+ P P ++ K + P++F
Sbjct: 206 SESTSNSLASPSTTPSTPSSSSSPTSPSTTKQISTPSPSPPPPVATAASSSKNSAEPRRF 385
Query: 294 K 294
K
Sbjct: 386 K 388
>TC17623 similar to UP|Q93YH8 (Q93YH8) HMG I/Y like protein, partial (19%)
Length = 624
Score = 31.2 bits (69), Expect = 0.22
Identities = 27/110 (24%), Positives = 43/110 (38%), Gaps = 6/110 (5%)
Frame = +1
Query: 162 EETENESLTEDVNESLTEDVGEMEKNESLTTDVASLSLGDDA--LASPPPQPKTRRKLVA 219
E+ + LT+ LT + ++ N L D S L A + PPP+ A
Sbjct: 223 EQVYKDQLTQSHESLLTHHLKRLKTNGMLVMDKKSYKLPGSAPPILPPPPENIAAGAGAA 402
Query: 220 PQPAVRRSARIKTGEMEKNESLTTDVASL----SLGDDALASPPPQPKTR 265
P PA R R + + ++ T +A + L P QP+T+
Sbjct: 403 PSPASRPRGRPRKVQPPQHLPQTLTLAGVQLQPQLPPQQQVQPEAQPQTQ 552
>TC10315 similar to UP|P93166 (P93166) SCOF-1, partial (42%)
Length = 592
Score = 30.8 bits (68), Expect = 0.28
Identities = 27/80 (33%), Positives = 32/80 (39%), Gaps = 8/80 (10%)
Frame = +3
Query: 207 PPPQPKTRRKLVAPQPAVRRSARIKTGEM--------EKNESLTTDVASLSLGDDALASP 258
PPP P R QP+V R +R + G K+ SL+ SL A A P
Sbjct: 165 PPPPPHPSRSTTT-QPSVTRRSRGQNGNAPSDLVHAPRKSTSLSASSCSL-----AAAPP 326
Query: 259 PPQPKTRRKLAAPQPAAPQP 278
PP P R QP P P
Sbjct: 327 PPLPPLR---CNRQPLLPVP 377
>AV428818
Length = 417
Score = 30.0 bits (66), Expect = 0.48
Identities = 17/70 (24%), Positives = 29/70 (41%)
Frame = +3
Query: 205 ASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKT 264
A+PP P+T +P P+ +++ T S T S +++S P T
Sbjct: 198 AAPPLSPRTSTSTASPSPSPSPNSKTATTSPPSTNSSGTSPPSALTSATSVSSRNPSCST 377
Query: 265 RRKLAAPQPA 274
R +P P+
Sbjct: 378 ARPECSPTPS 407
>TC19563
Length = 538
Score = 29.3 bits (64), Expect = 0.83
Identities = 20/62 (32%), Positives = 35/62 (56%), Gaps = 7/62 (11%)
Frame = +3
Query: 150 ETENESLTKDVPEETENESLTEDVNESLTE---DVGEMEKN----ESLTTDVASLSLGDD 202
E+ENE L ++ E ++ ++ ED+ E L+E +++KN +S ++ S SL DD
Sbjct: 45 ESENEILVQEEKEVSDIKTKIEDIKEHLSELEVKASDLDKNLLSIKSQVQNLDSKSLLDD 224
Query: 203 AL 204
L
Sbjct: 225 LL 230
>BP067321
Length = 513
Score = 29.3 bits (64), Expect = 0.83
Identities = 24/88 (27%), Positives = 36/88 (40%), Gaps = 9/88 (10%)
Frame = +1
Query: 207 PPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALAS--------- 257
PPP P RR+ +P ++++ + + S TT + S S +A AS
Sbjct: 94 PPPPPPLRRQSQSPNTHLQKNPKTRHHYRRTRASSTTTIPSTSSPWNAAASSRQTATGP* 273
Query: 258 PPPQPKTRRKLAAPQPAAPQPAVRRSAR 285
P R AP P++ A RR R
Sbjct: 274 SSGSPACRTSSPAP-PSSSSSATRRPRR 354
>TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%)
Length = 514
Score = 29.3 bits (64), Expect = 0.83
Identities = 24/93 (25%), Positives = 35/93 (36%), Gaps = 10/93 (10%)
Frame = +3
Query: 206 SPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTR 265
S PP P+ R P P + S + S S + +AS PP P +
Sbjct: 156 SAPPSPEERFSGTPPPPPPATGSASTATPTPPTSSRSASPPSPSPANSHMASSPPSPTSA 335
Query: 266 R--KLAAPQPA--------APQPAVRRSARIKT 288
++ P PA AP+ A S+RI +
Sbjct: 336 LSVSVSTPSPAHSPPTSPPAPRSATSTSSRISS 434
>AV406930
Length = 433
Score = 28.9 bits (63), Expect = 1.1
Identities = 22/92 (23%), Positives = 40/92 (42%), Gaps = 8/92 (8%)
Frame = +1
Query: 192 TDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEK-----NESLTTDV- 245
+ +++L+L +PPP P+ P++ ++ I T + + S +
Sbjct: 43 SSISALALAPPHFLAPPPPPQPPNP-----PSISQNLSISTHPITPFHTPLSHSFNLPLP 207
Query: 246 --ASLSLGDDALASPPPQPKTRRKLAAPQPAA 275
AS+ L AL SP P P + L P P++
Sbjct: 208 KPASIPLSFSALPSPSPSPSSSPTLIPPPPSS 303
Score = 27.7 bits (60), Expect = 2.4
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +1
Query: 195 ASLSLGDDALASPPPQPKTRRKLVAPQPAVRR 226
AS+ L AL SP P P + L+ P P+ R
Sbjct: 214 ASIPLSFSALPSPSPSPSSSPTLIPPPPSSSR 309
>BP041908
Length = 487
Score = 28.9 bits (63), Expect = 1.1
Identities = 18/49 (36%), Positives = 23/49 (46%), Gaps = 1/49 (2%)
Frame = +3
Query: 240 SLTTDVASLSLGDD-ALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIK 287
+LT SL+L D A ++PPP T + P P PA SA K
Sbjct: 168 TLTASKISLTLPHDPASSTPPPATSTPTTTSTSPPGEPPPASTNSATSK 314
Score = 27.7 bits (60), Expect = 2.4
Identities = 29/103 (28%), Positives = 39/103 (37%), Gaps = 7/103 (6%)
Frame = +3
Query: 189 SLTTDVASLSLGDDALASPPPQPKTRRKLVA------PQPAVRRSARIKTGEMEKNESLT 242
+LT SL+L D +S PP + + P PA SA K E S
Sbjct: 168 TLTASKISLTLPHDPASSTPPPATSTPTTTSTSPPGEPPPASTNSATSKK-ETSS*SSSP 344
Query: 243 TDVASLSLGDDALASPP-PQPKTRRKLAAPQPAAPQPAVRRSA 284
T + S S L S P P P+ P+ P+P S+
Sbjct: 345 TPLNSSSPSSPHLTSAPWPPPQIPSSPPRKSPSKPKPPTPSSS 473
>TC10958 similar to PIR|T47938|T47938 RING finger protein - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (38%)
Length = 779
Score = 28.5 bits (62), Expect = 1.4
Identities = 12/35 (34%), Positives = 17/35 (48%)
Frame = +2
Query: 258 PPPQPKTRRKLAAPQPAAPQPAVRRSARIKTTPKK 292
PP + RR+L P P P P R+ T P++
Sbjct: 206 PPRHRRPRRRLTGPPPPDPHPYTLRNPHPGTPPRR 310
>TC19504 similar to UP|Q9SV89 (Q9SV89) Isoleucine-tRNA ligase-like protein
, partial (11%)
Length = 491
Score = 28.5 bits (62), Expect = 1.4
Identities = 19/67 (28%), Positives = 27/67 (39%)
Frame = +3
Query: 219 APQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQP 278
+P P RS T SLTT +S + + + P P T + A+ A P
Sbjct: 171 SPSPRTCRSTSSTTAHPSPPASLTTATSSPAQSKTSSPATSP*PATTSRAASAGTATASP 350
Query: 279 AVRRSAR 285
+ RS R
Sbjct: 351 SRTRSTR 371
>TC13202
Length = 431
Score = 28.5 bits (62), Expect = 1.4
Identities = 13/37 (35%), Positives = 18/37 (48%)
Frame = +2
Query: 258 PPPQPKTRRKLAAPQPAAPQPAVRRSARIKTTPKKFK 294
PPP P ++ PQP +P P+ A T +K K
Sbjct: 212 PPPPPSVLQQAWQPQPPSPIPSAMNGATNARTREKAK 322
>BP044070
Length = 289
Score = 28.1 bits (61), Expect = 1.8
Identities = 21/62 (33%), Positives = 28/62 (44%), Gaps = 1/62 (1%)
Frame = +3
Query: 203 ALASPPPQP-KTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQ 261
A++SPPP P + + +P P V S R + S T S S A +SPPP
Sbjct: 108 AVSSPPPPPTRPSAPMASPPPTVSPSPRCSSTRATTXASPT----SPSPPTPASSSPPPT 275
Query: 262 PK 263
K
Sbjct: 276 TK 281
>BI420337
Length = 397
Score = 27.3 bits (59), Expect = 3.1
Identities = 21/69 (30%), Positives = 26/69 (37%)
Frame = +1
Query: 205 ASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKT 264
+SPPP PK K +P P V + K+ T SPPP PK
Sbjct: 181 SSPPPPPKKPYKYPSPPPPVYK---------YKSPPPRTPAYYYK-------SPPPPPKN 312
Query: 265 RRKLAAPQP 273
K +P P
Sbjct: 313 PYKYPSPPP 339
>AV427911
Length = 348
Score = 26.2 bits (56), Expect = 7.0
Identities = 19/68 (27%), Positives = 22/68 (31%)
Frame = +1
Query: 206 SPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTR 265
SPPP PK K +P P V + SPPP PK
Sbjct: 193 SPPPPPKKPYKYPSPPPPVHVYPK-----------------------PTYHSPPPPPKKP 303
Query: 266 RKLAAPQP 273
K +P P
Sbjct: 304 YKYPSPPP 327
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.309 0.127 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,953,516
Number of Sequences: 28460
Number of extensions: 80486
Number of successful extensions: 967
Number of sequences better than 10.0: 105
Number of HSP's better than 10.0 without gapping: 767
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 947
length of query: 295
length of database: 4,897,600
effective HSP length: 90
effective length of query: 205
effective length of database: 2,336,200
effective search space: 478921000
effective search space used: 478921000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0109.1


