

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
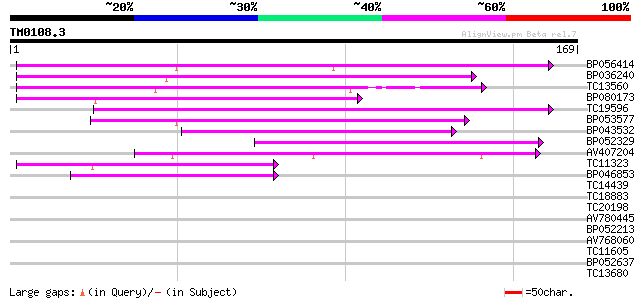
Query= TM0108.3
(169 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP056414 97 1e-21
BP036240 82 5e-17
TC13560 74 2e-14
BP080173 52 7e-08
TC19596 49 3e-07
BP053577 49 4e-07
BP043532 47 2e-06
BP052329 42 5e-05
AV407204 42 7e-05
TC11323 40 2e-04
BP046853 40 2e-04
TC14439 similar to UP|Q9FIC1 (Q9FIC1) Gb|AAF19567.1, partial (49%) 37 0.002
TC18883 35 0.007
TC20198 similar to UP|Q9ZPU0 (Q9ZPU0) At2g13980 protein, partial... 35 0.007
AV780445 34 0.011
BP052213 33 0.020
AV768060 31 0.097
TC11605 29 0.48
BP052637 28 0.82
TC13680 similar to UP|Q84V02 (Q84V02) Vacuolar membrane ATPase s... 27 1.4
>BP056414
Length = 606
Score = 97.4 bits (241), Expect = 1e-21
Identities = 60/164 (36%), Positives = 83/164 (50%), Gaps = 4/164 (2%)
Frame = -3
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQGFCRRL*S---SNPLLAELWAI 59
W PP G +K NVDG++ +A CGGV+RD G+W +GF R L EL A+
Sbjct: 562 WTRPPEGWFKVNVDGALSGDLQASCGGVVRDAAGSWSKGFARSLGVLRWHXAFFVELMAV 383
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIP-PYSEIVRHIRRSIPHGMKVVFDL 118
+TAVD + +IIE+DS + + + P Y EI I++ I +V
Sbjct: 382 QTAVDFIMSWDIPQVIIESDSQQVVELLQNFNASSPHQYGEIAWDIQQKISVHGSIVITS 203
Query: 119 IPREANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDSMTV 162
PREAN +AD LA+ FG H P EC LL +D M++
Sbjct: 202 TPREANFLADYLAKVGLSLPFGTHSFDAPFGECHSLLQKDLMSI 71
>BP036240
Length = 567
Score = 82.0 bits (201), Expect = 5e-17
Identities = 49/140 (35%), Positives = 74/140 (52%), Gaps = 3/140 (2%)
Frame = -3
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQGFCRR---L*SSNPLLAELWAI 59
W P G KF+VDG++ + A CGGVLRD +W++GFCR L N L EL AI
Sbjct: 562 WTAPDEGWTKFDVDGALSRDLLASCGGVLRDSRRHWIKGFCRNMGDLTYHNVFLVELSAI 383
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLI 119
++AV++A+ +I+E+D+ EA ++S P+ + I R +VF
Sbjct: 382 QSAVEIALSMDLQHVIVESDALEAIHLLSSENVHAHPFGSLAASILRMQRDHGSIVFQHT 203
Query: 120 PREANMVADCLAQNAHVFNF 139
PREAN + + VF++
Sbjct: 202 PREANSLVTIWQEWVIVFHW 143
>TC13560
Length = 571
Score = 73.6 bits (179), Expect = 2e-14
Identities = 51/146 (34%), Positives = 77/146 (51%), Gaps = 6/146 (4%)
Frame = +2
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQGF---CRRL*SSNPLLAELWAI 59
W+ P G K NVDG+V +A C GVLR+ DG+W++GF S+N L EL A+
Sbjct: 146 WIKPDAGWVKINVDGAVSNCLQASCRGVLRNADGSWLKGFRWNFGTFSSANVFLTELMAV 325
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEI---VRHIRRSIPHGMKVVF 116
+TAV+ A+ + +I+E+DS E + S YS+I + H++R HG +VF
Sbjct: 326 KTAVEAAMSLDLARVIVESDSLEVVDFLHSTDLGSHRYSQIALDILHLQRE--HG-ALVF 496
Query: 117 DLIPREANMVADCLAQNAHVFNFGVH 142
R N + D +A+ +G H
Sbjct: 497 QYALR-GNSIVDYIAKFGLSLPYGEH 571
>BP080173
Length = 382
Score = 51.6 bits (122), Expect = 7e-08
Identities = 34/104 (32%), Positives = 51/104 (48%), Gaps = 1/104 (0%)
Frame = -2
Query: 3 WLLPPVGSWKFNVDGSVWQTRE-AGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W +P G K N+DGS + A CGGV+ D +G ++ G RL AELW I
Sbjct: 345 WPVPMEGWIKINIDGSFAPSSNCAACGGVMWDHEGTFITGASVRLGCYFINHAELWGIFH 166
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIR 105
+A RG I++E+DS+ + V P + +V+ I+
Sbjct: 165 GAKIAQARGFKKILVESDSSFVVSLVNQGCSPSKPATPLVKDIK 34
>TC19596
Length = 499
Score = 49.3 bits (116), Expect = 3e-07
Identities = 35/137 (25%), Positives = 57/137 (41%)
Frame = +1
Query: 26 GCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDSAEAHA 85
G GG++RD G W+ GF + L AE+ A++ + L E D +
Sbjct: 49 GMGGIVRDAHGAWISGFYAGSLGGDALRAEIAALKHGLTLLWNAHVRRATCEVDCLDIVE 228
Query: 86 AVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMVADCLAQNAHVFNFGVHLLH 145
A+ + + + + IR + V +PREAN ADCLA + L
Sbjct: 229 ALENDRYQFHALASELLDIRLLLDRDWTVTLAYVPREANAAADCLAGLGASLLCPLTCLE 408
Query: 146 HPLMECRYLLYQDSMTV 162
P E + +L +D + +
Sbjct: 409 SPPQELQPILARDLLAL 459
>BP053577
Length = 463
Score = 48.9 bits (115), Expect = 4e-07
Identities = 38/114 (33%), Positives = 51/114 (44%), Gaps = 1/114 (0%)
Frame = +3
Query: 25 AGCGGVLRDCDGNWVQGFCRRL*S-SNPLLAELWAIRTAVDLAVQRGCSPIIIETDSAEA 83
AG G V R+CDG + C S S+PLLAE +R + LA G I +ETD +
Sbjct: 12 AGLGMVARNCDGEVMASACSSPVSISSPLLAEALGLRWTMQLATDLGFRRITLETDCLQL 191
Query: 84 HAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMVADCLAQNAHVF 137
S + S I+ R I V + R N VAD LA+N+ +
Sbjct: 192 FNTWRSQEEGRSYLSSIISDCRILISAFDYVSVSFVRRTGNTVADFLARNSDTY 353
>BP043532
Length = 465
Score = 47.0 bits (110), Expect = 2e-06
Identities = 27/82 (32%), Positives = 40/82 (47%)
Frame = -2
Query: 52 LLAELWAIRTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHG 111
L AELW + + A+ RG S I+IE DSA + P++ P +VR +
Sbjct: 446 LHAELWGLVHGLRFALGRGFSKILIEADSAGTIEFLNKGCPVVHPCFPLVREFHHLVGQN 267
Query: 112 MKVVFDLIPREANMVADCLAQN 133
+ + + RE N VAD A+N
Sbjct: 266 CYIHWKHVYREVNFVADSFAKN 201
>BP052329
Length = 441
Score = 42.0 bits (97), Expect = 5e-05
Identities = 29/86 (33%), Positives = 42/86 (48%)
Frame = -3
Query: 74 IIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMVADCLAQN 133
+IIE+DS E + + S +++I I R + VF PRE+N +AD L +
Sbjct: 352 VIIESDSLEVVSLLHSPDVGEHHFAQIALEIIRMQNDHGEHVFQHTPRESNSMADYLVKI 173
Query: 134 AHVFNFGVHLLHHPLMECRYLLYQDS 159
+G H PL EC LL +DS
Sbjct: 172 GIDLPYGPHWFGSPLGECHSLLLKDS 95
>AV407204
Length = 432
Score = 41.6 bits (96), Expect = 7e-05
Identities = 35/125 (28%), Positives = 54/125 (43%), Gaps = 4/125 (3%)
Frame = -2
Query: 38 WVQGFCRRL*--SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDSAEAHAAVTS-AQPLI 94
WV GF + +S LAE A+R + LA G ++ +D E A+ ++
Sbjct: 428 WVNGFISQYLEAASCSFLAEALALRDGLRLAWDNGTRKLVCNSDCKELIDAIADPSRASF 249
Query: 95 PPYSEIVRHIRRSIPHGMKVVFDLIPREANMVADCLAQNAHVFNF-GVHLLHHPLMECRY 153
+ ++R I + +V R+ NMVADCLA+ + G L P E
Sbjct: 248 HAHGWVLREIHALMTRNWRVELFWCCRDVNMVADCLAKRGSLLPVAGYSALATPSPELEV 69
Query: 154 LLYQD 158
LL +D
Sbjct: 68 LLLKD 54
>TC11323
Length = 657
Score = 40.4 bits (93), Expect = 2e-04
Identities = 30/79 (37%), Positives = 38/79 (47%), Gaps = 1/79 (1%)
Frame = +2
Query: 3 WLLPPVGSWKFNVDGSVWQTR-EAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
WL P G +K N DGS R AG GG+LRD N++ GF L + + L AI
Sbjct: 5 WLKPVHGRFKLNFDGSSLGNRGNAGGGGLLRDGSSNFIFGFSIFLAVAQIMKLSLCAILE 184
Query: 62 AVDLAVQRGCSPIIIETDS 80
+ + G I IE DS
Sbjct: 185GLLVCKPLGYDGIGIECDS 241
>BP046853
Length = 612
Score = 40.0 bits (92), Expect = 2e-04
Identities = 20/62 (32%), Positives = 36/62 (57%)
Frame = -3
Query: 19 VWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIET 78
V+ +A G ++RD + G ++ S+PL AE AIR ++LA GC+ +++E+
Sbjct: 418 VYHGPQAALGVLIRDKYSTLLSGTHAKIYCSSPLTAEACAIREGLNLAYNLGCTEVVLES 239
Query: 79 DS 80
D+
Sbjct: 238 DN 233
>TC14439 similar to UP|Q9FIC1 (Q9FIC1) Gb|AAF19567.1, partial (49%)
Length = 2120
Score = 36.6 bits (83), Expect = 0.002
Identities = 22/50 (44%), Positives = 31/50 (62%)
Frame = +3
Query: 119 IPREANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDSMTVVASLSL 168
IPRE N VAD +A++A V G LL P + + LL +DS++ + S SL
Sbjct: 1950 IPREGNYVADFMAKSAPV-QLGHRLLREPDPDLQVLLMKDSLSAL*SFSL 2096
>TC18883
Length = 821
Score = 35.0 bits (79), Expect = 0.007
Identities = 33/111 (29%), Positives = 52/111 (46%), Gaps = 4/111 (3%)
Frame = +1
Query: 26 GCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDSAEAHA 85
G GG+ R+ D + F + + AE+WAI A+ + V R + IE+DS+ A
Sbjct: 10 GIGGIFRNSDNEILGFFSKHVGEGFAFEAEVWAIFEALTICVNRQLRNLTIESDSSLAVG 189
Query: 86 AVT--SAQP--LIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMVADCLAQ 132
V S +P L+ ++I + +K +F REAN D LA+
Sbjct: 190 WVNNRSXRPWKLLNKLNQIDLWVAEVNCLSVKHIF----REANGEVDKLAK 330
>TC20198 similar to UP|Q9ZPU0 (Q9ZPU0) At2g13980 protein, partial (13%)
Length = 630
Score = 35.0 bits (79), Expect = 0.007
Identities = 17/35 (48%), Positives = 20/35 (56%)
Frame = +3
Query: 2 CWLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDG 36
CW PP G +K N D SV + AG V+RDC G
Sbjct: 90 CWPPPPDGVFKLNFDTSVLASS*AGLSLVVRDCAG 194
>AV780445
Length = 504
Score = 34.3 bits (77), Expect = 0.011
Identities = 18/38 (47%), Positives = 22/38 (57%), Gaps = 1/38 (2%)
Frame = +1
Query: 3 WLLPPVGSWKFNVDGSVW-QTREAGCGGVLRDCDGNWV 39
W P G+ K NVDGS Q R G GG++RD G W+
Sbjct: 391 WQPPRQGTVKLNVDGSGREQDRCMGAGGLIRDDQGKWL 504
>BP052213
Length = 375
Score = 33.5 bits (75), Expect = 0.020
Identities = 18/45 (40%), Positives = 26/45 (57%), Gaps = 3/45 (6%)
Frame = -3
Query: 124 NMVADCLAQNAHVFNFGVHLLHHPL---MECRYLLYQDSMTVVAS 165
N VAD LA AH G+H+L HPL + C L++++ + V S
Sbjct: 259 NQVADELASIAHGLELGIHVLDHPLQGVLSCFSLMFRECPSRVYS 125
>AV768060
Length = 416
Score = 31.2 bits (69), Expect = 0.097
Identities = 24/95 (25%), Positives = 39/95 (40%)
Frame = -3
Query: 43 CRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVR 102
CRR PL+AE R A++LA G ++ +D + A P + I+
Sbjct: 312 CRR----EPLIAEAMCFRWALNLAANWGLDSLVFSSDCQQLVKAFHCPSE-FPSLAHIML 148
Query: 103 HIRRSIPHGMKVVFDLIPREANMVADCLAQNAHVF 137
+ + F R+ N VA LA +H++
Sbjct: 147 DCTQLVNSFRNFSFCFEYRQTNRVAHALAAESHLY 43
>TC11605
Length = 546
Score = 28.9 bits (63), Expect = 0.48
Identities = 21/62 (33%), Positives = 29/62 (45%), Gaps = 1/62 (1%)
Frame = +3
Query: 3 WLLPPVGSWKFNVDGSV-WQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W P KFNVDGS +G GG+LR+ + + F + L AE+ AI
Sbjct: 348 WCPPEANILKFNVDGSARGSPGISGAGGILRNAESAVLGRFSKPLGVLCAYQAEVKAILL 527
Query: 62 AV 63
A+
Sbjct: 528 AL 533
>BP052637
Length = 473
Score = 28.1 bits (61), Expect = 0.82
Identities = 17/54 (31%), Positives = 26/54 (47%), Gaps = 7/54 (12%)
Frame = -2
Query: 17 GSVWQTREAGCG-------GVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAV 63
G V + + GCG GVLRD WV GF + + +A+L A+ ++
Sbjct: 424 GRVMR*WQHGCGLKCYWGSGVLRDKVXRWVSGFAMAFGTGDAFMAKLQALHESL 263
>TC13680 similar to UP|Q84V02 (Q84V02) Vacuolar membrane ATPase subunit c'',
partial (18%)
Length = 536
Score = 27.3 bits (59), Expect = 1.4
Identities = 16/37 (43%), Positives = 18/37 (48%), Gaps = 5/37 (13%)
Frame = -1
Query: 112 MKVVFDLIP-----REANMVADCLAQNAHVFNFGVHL 143
+ VVFD RE N V D LA +FGVHL
Sbjct: 506 LMVVFDFCRVSHDFRETNKVVDALAHRGQSNSFGVHL 396
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.141 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,053,422
Number of Sequences: 28460
Number of extensions: 64930
Number of successful extensions: 435
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 424
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 427
length of query: 169
length of database: 4,897,600
effective HSP length: 84
effective length of query: 85
effective length of database: 2,506,960
effective search space: 213091600
effective search space used: 213091600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0108.3


