

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0108.13
(1262 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
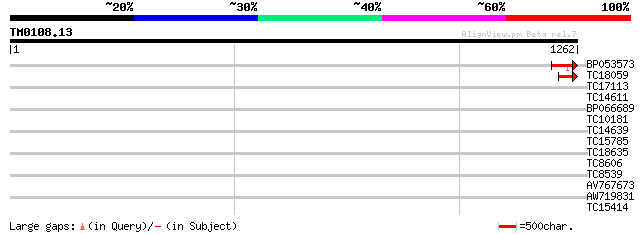
Score E
Sequences producing significant alignments: (bits) Value
BP053573 100 1e-21
TC18059 87 2e-17
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 32 0.48
TC14611 similar to UP|Q8LPB2 (Q8LPB2) Glycine-rich RNA binding p... 30 1.8
BP066689 30 2.4
TC10181 homologue to UP|GAPN_NICPL (P93338) NADP-dependent glyce... 29 4.1
TC14639 weakly similar to GB|AAM91068.1|22136564|AY129482 AT5g11... 28 6.9
TC15785 similar to UP|Q94BV7 (Q94BV7) AT4g05020/T32N4_4, partial... 28 6.9
TC18635 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (6%) 28 6.9
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 28 6.9
TC8539 similar to UP|Q94KE3 (Q94KE3) AT3g52990/F8J2_160 (Pyruvat... 28 6.9
AV767673 28 9.0
AW719831 28 9.0
TC15414 weakly similar to UP|O81655 (O81655) Senescence-associat... 28 9.0
>BP053573
Length = 533
Score = 100 bits (249), Expect = 1e-21
Identities = 56/81 (69%), Positives = 56/81 (69%), Gaps = 25/81 (30%)
Frame = -1
Query: 1207 QVKEACRKAYAAEQLAKERLSMMKDDLSIHCRIT-------------------------N 1241
QVKEACRKAYAAEQLAKERLSMMKDDLSIHCRIT N
Sbjct: 521 QVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITVCAQL*YPKLVPPIEQLAYFSLFVQN 342
Query: 1242 LQRPRVRFADHVQEKIIKGRR 1262
LQRPRVRFADHVQEKIIKGRR
Sbjct: 341 LQRPRVRFADHVQEKIIKGRR 279
>TC18059
Length = 379
Score = 87.0 bits (214), Expect = 2e-17
Identities = 42/42 (100%), Positives = 42/42 (100%)
Frame = +1
Query: 1221 LAKERLSMMKDDLSIHCRITNLQRPRVRFADHVQEKIIKGRR 1262
LAKERLSMMKDDLSIHCRITNLQRPRVRFADHVQEKIIKGRR
Sbjct: 1 LAKERLSMMKDDLSIHCRITNLQRPRVRFADHVQEKIIKGRR 126
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 32.3 bits (72), Expect = 0.48
Identities = 13/26 (50%), Positives = 15/26 (57%)
Frame = -2
Query: 868 GGNKEGSVQKGIAYGSKGNNGRKGCY 893
GG G V+ GI G KGN G GC+
Sbjct: 764 GGGGGGGVKPGITIGGKGNKGEGGCF 687
>TC14611 similar to UP|Q8LPB2 (Q8LPB2) Glycine-rich RNA binding protein,
partial (20%)
Length = 760
Score = 30.4 bits (67), Expect = 1.8
Identities = 26/75 (34%), Positives = 29/75 (38%), Gaps = 19/75 (25%)
Frame = +3
Query: 49 GQRRPDSGPRQGGLYSGLGGSSGNDEKR-------------------EPPEEDDGDTLLV 89
G R+P G QG SG GG S +EK+ PP DD D
Sbjct: 114 GGRKPSYGEEQG---SGYGGRSEYEEKKPSYGRSNYEGQEEVEYGRERPPRLDDDDDDNE 284
Query: 90 KYLRKKAKRDDDGGD 104
Y RK K DDG D
Sbjct: 285 GYARK--KYGDDGSD 323
>BP066689
Length = 470
Score = 30.0 bits (66), Expect = 2.4
Identities = 17/49 (34%), Positives = 24/49 (48%)
Frame = +1
Query: 64 SGLGGSSGNDEKREPPEEDDGDTLLVKYLRKKAKRDDDGGDLPRLLTKD 112
+G+ G +E+ E EEDD + +KY R GG +P LL D
Sbjct: 163 NGVEGDDEREEQEEEDEEDDEEEPRLKYQRM-------GGSVPSLLAAD 288
>TC10181 homologue to UP|GAPN_NICPL (P93338) NADP-dependent
glyceraldehyde-3-phosphate dehydrogenase
(Non-phosphorylating glyceraldehyde 3-phosphate
dehydrogenase) (Glyceraldehyde-3-phosphate dehydrogenase
[NADP+]) (Triosephosphate dehydrogenase) , partial (38%)
Length = 582
Score = 29.3 bits (64), Expect = 4.1
Identities = 16/45 (35%), Positives = 27/45 (59%)
Frame = -2
Query: 710 REPNFDLCSAIVDFHSTKIVANDGHDVDIKHMQNSISSLQERGPS 754
R PN LC+ ++DF + K V+N HD H + S++++ E P+
Sbjct: 134 RRPNS*LCNFVIDFFN-KSVSN*FHDQGNLHSRASLTTVGELVPN 3
>TC14639 weakly similar to GB|AAM91068.1|22136564|AY129482
AT5g11740/T22P22_130 {Arabidopsis thaliana;}, partial
(61%)
Length = 507
Score = 28.5 bits (62), Expect = 6.9
Identities = 20/54 (37%), Positives = 28/54 (51%), Gaps = 1/54 (1%)
Frame = -1
Query: 25 NPSSSPPGALTRSRSQLFVHRNRSGQRRPDSGPRQGGLYS-GLGGSSGNDEKRE 77
+P+SS P ++ S + G+ R DSG R+GG S L GSS D+ E
Sbjct: 264 HPASSDPERRSKGESS-DGGKETGGEGRRDSGRRRGGARSRSLSGSSDGDDGGE 106
>TC15785 similar to UP|Q94BV7 (Q94BV7) AT4g05020/T32N4_4, partial (43%)
Length = 854
Score = 28.5 bits (62), Expect = 6.9
Identities = 13/27 (48%), Positives = 19/27 (70%)
Frame = -2
Query: 725 STKIVANDGHDVDIKHMQNSISSLQER 751
S+++V + D +KH QN ISSLQ+R
Sbjct: 343 SSELVFSKSSDSFVKHAQNMISSLQKR 263
>TC18635 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (6%)
Length = 610
Score = 28.5 bits (62), Expect = 6.9
Identities = 22/92 (23%), Positives = 38/92 (40%)
Frame = +3
Query: 584 LMELAKDDPGISKFGHGTPKKDEADMHGKRFELTLSSQPEEASIDGHKIEKGPIHGTVES 643
++ LA+ P I GT +++ ++ E QP++ DGH G S
Sbjct: 60 IVGLARTRPDI----FGTTEEEVSNAVKAEIEKKNDEQPKQVIWDGHTGSIGRTANQAMS 227
Query: 644 NALGNILFNSANELSNGNLPKLTSSQDLPELP 675
++G+ N A+ NLP + P +P
Sbjct: 228 QSIGSEDQNDASNYEGKNLPGPAAPPPRPGMP 323
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 28.5 bits (62), Expect = 6.9
Identities = 33/115 (28%), Positives = 44/115 (37%), Gaps = 6/115 (5%)
Frame = -2
Query: 178 LGFGSSAEKGGLCVDDATMKVTVSP----DLGISSQARDFDRANTDSSDHKQFEEGNDGD 233
LG G+ AE V A +K T+ L + + A + S D + EG DG+
Sbjct: 782 LGGGALAEVVVTLVASAVVKATLLAADLCGLLLINMATTKGETKSQSEDKQHDAEGEDGN 603
Query: 234 CSGKINDFDEEIVVTTPPDAELLGDSKVDGDE--GKGEEDLLVRDAPFTGKAENS 286
D+E GD DGD G+GEEDL D G N+
Sbjct: 602 --------DDE------------GDEDDDGDGPFGEGEEDLSSEDGAGYGNNSNN 498
>TC8539 similar to UP|Q94KE3 (Q94KE3) AT3g52990/F8J2_160 (Pyruvate
kinase-like protein), partial (81%)
Length = 1430
Score = 28.5 bits (62), Expect = 6.9
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = +3
Query: 927 KRLLPLLSNAKKYNSCASVNRHDPKHQKFLDQTPLLPIS 965
+R PLL N + NS + + P H L QTP LP++
Sbjct: 54 QRSTPLLFNHRHRNSSSPI----PPHAAVLSQTPCLPVT 158
>AV767673
Length = 497
Score = 28.1 bits (61), Expect = 9.0
Identities = 25/96 (26%), Positives = 40/96 (41%)
Frame = -1
Query: 617 TLSSQPEEASIDGHKIEKGPIHGTVESNALGNILFNSANELSNGNLPKLTSSQDLPELPM 676
T +SQP A+ + + S++L A ELS +PK TSS
Sbjct: 476 TSNSQPGAAAAAPRAADPS---SSTSSSSLNQHQKEIAEELSTSQVPKATSSLASASTQK 306
Query: 677 QSDVKEVVQECLSAPSVEEPIEKAVVGSKDECSREP 712
+ E+ + SAP+ EP++ S +E + P
Sbjct: 305 SNLTPELTNQ--SAPTEAEPMQIEASSSANENANPP 204
>AW719831
Length = 436
Score = 28.1 bits (61), Expect = 9.0
Identities = 18/68 (26%), Positives = 29/68 (42%)
Frame = +3
Query: 802 SRYPSEGQSSSQLDTDMQDLEENERSNGAIRKSDDAVVLNHVGVSNEVPRPPPENIISKT 861
S +P+ SS + +D+ D N +N +H +E P PPP K
Sbjct: 111 SSHPNSRDSSPRSRSDVIDTTTNNDTNN-----------HHHSSFDEPPPPPPSAAACKV 257
Query: 862 QDMAAHGG 869
+ M ++GG
Sbjct: 258 KLMCSYGG 281
>TC15414 weakly similar to UP|O81655 (O81655) Senescence-associated protein
5, partial (47%)
Length = 448
Score = 28.1 bits (61), Expect = 9.0
Identities = 21/75 (28%), Positives = 33/75 (44%), Gaps = 2/75 (2%)
Frame = -2
Query: 196 MKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKIND--FDEEIVVTTPPDA 253
++V P L I+S + F +S + DG+C + ++ DEE+ P DA
Sbjct: 231 VRVISKPILPITSTRKRFTSTLVGNSKRE------DGECEDEDDEEEHDEEVDPEEP*DA 70
Query: 254 ELLGDSKVDGDEGKG 268
D +GD G G
Sbjct: 69 AARADETGEGDRGGG 25
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.131 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,017,049
Number of Sequences: 28460
Number of extensions: 274229
Number of successful extensions: 1117
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1108
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1116
length of query: 1262
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1161
effective length of database: 2,023,140
effective search space: 2348865540
effective search space used: 2348865540
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0108.13


