

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0106.15
(120 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
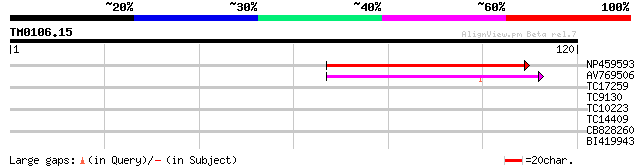
NP459593 Pge1 protein [Lotus japonicus] 61 6e-11
AV769506 37 9e-04
TC17259 weakly similar to UP|Q94KL5 (Q94KL5) BEL1-like homeobox ... 28 0.42
TC9130 homologue to UP|Q8LLE2 (Q8LLE2) BEL1-related homeotic pro... 27 0.71
TC10223 26 2.1
TC14409 similar to PIR|A84668|A84668 Argonaute (AGO1)-like prote... 25 4.6
CB828260 25 4.6
BI419943 24 7.9
>NP459593 Pge1 protein [Lotus japonicus]
Length = 633
Score = 60.8 bits (146), Expect = 6e-11
Identities = 26/43 (60%), Positives = 33/43 (76%)
Frame = +1
Query: 68 VFDWAQLVGKNNDFVVIMIRSDYGTLNGKESLTLGCEHGGKFK 110
+ DWA+ VGK N +VVI+IRSDYG+ K +TLGCE GGK+K
Sbjct: 487 LIDWARCVGKENGYVVIVIRSDYGSAKRKPLITLGCERGGKYK 615
>AV769506
Length = 345
Score = 37.0 bits (84), Expect = 9e-04
Identities = 15/47 (31%), Positives = 28/47 (58%), Gaps = 1/47 (2%)
Frame = +1
Query: 68 VFDWAQLVGKNNDFVVIMIRSDYGTLNGKES-LTLGCEHGGKFKPYK 113
+ +W + N +V ++ +SDYG +++ + LGCE GK+ PY+
Sbjct: 61 LINWVHGIAIENGYVTLITKSDYGGNESRKAYVMLGCEKHGKYVPYR 201
>TC17259 weakly similar to UP|Q94KL5 (Q94KL5) BEL1-like homeobox 4, partial
(6%)
Length = 552
Score = 28.1 bits (61), Expect = 0.42
Identities = 13/28 (46%), Positives = 14/28 (49%)
Frame = +1
Query: 92 TLNGKESLTLGCEHGGKFKPYKGKFDIR 119
T G SLTLG H G P K F +R
Sbjct: 313 TTAGDVSLTLGLRHAGNVPPEKSPFSLR 396
>TC9130 homologue to UP|Q8LLE2 (Q8LLE2) BEL1-related homeotic protein 13
(Fragment), partial (5%)
Length = 728
Score = 27.3 bits (59), Expect = 0.71
Identities = 16/44 (36%), Positives = 19/44 (42%)
Frame = +1
Query: 76 GKNNDFVVIMIRSDYGTLNGKESLTLGCEHGGKFKPYKGKFDIR 119
G D +IR +GT G SLTLG H G F +R
Sbjct: 235 GGGGDIGSTLIR--FGTTAGDVSLTLGLRHAGNTPEKTNSFSVR 360
>TC10223
Length = 957
Score = 25.8 bits (55), Expect = 2.1
Identities = 10/28 (35%), Positives = 18/28 (63%)
Frame = +2
Query: 12 VAEGDHIKVDRVKEVDNEVDDMKEWGHI 39
++E + R+K+V NEV +K+W +I
Sbjct: 116 ISETGSVDSTRLKQVQNEVSLLKQWKNI 199
>TC14409 similar to PIR|A84668|A84668 Argonaute (AGO1)-like protein
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (9%)
Length = 674
Score = 24.6 bits (52), Expect = 4.6
Identities = 15/59 (25%), Positives = 25/59 (41%)
Frame = -3
Query: 50 IDDMNALSHVVMKFQNRQVFDWAQLVGKNNDFVVIMIRSDYGTLNGKESLTLGCEHGGK 108
+DD++ LS M + N WA +G + V+++ Y L T C G+
Sbjct: 171 LDDVSDLSSNFMNWPNWVAAKWA*HIGATTEIAVVLL*YTYDRL-----CTSSCSSSGE 10
>CB828260
Length = 557
Score = 24.6 bits (52), Expect = 4.6
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = +3
Query: 91 GTLNGKESLTLGCEHGGKFKPYK 113
G + K + LGCE GK+ PY+
Sbjct: 162 GNGSRKAYVMLGCEKHGKYVPYR 230
>BI419943
Length = 495
Score = 23.9 bits (50), Expect = 7.9
Identities = 9/22 (40%), Positives = 14/22 (62%)
Frame = -1
Query: 60 VMKFQNRQVFDWAQLVGKNNDF 81
V+ FQN + F W L+ +N +F
Sbjct: 207 VLAFQNFE*FKWFPLISENPEF 142
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.139 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,620,672
Number of Sequences: 28460
Number of extensions: 15378
Number of successful extensions: 69
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 69
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 69
length of query: 120
length of database: 4,897,600
effective HSP length: 79
effective length of query: 41
effective length of database: 2,649,260
effective search space: 108619660
effective search space used: 108619660
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 49 (23.5 bits)
Lotus: description of TM0106.15


