

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
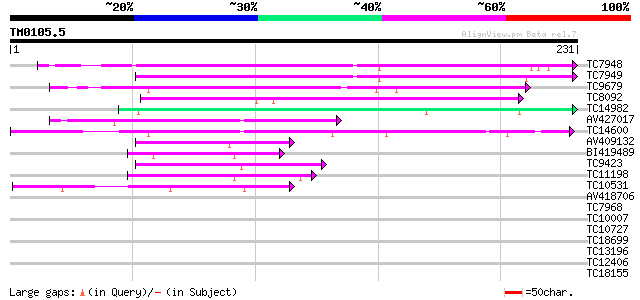
Query= TM0105.5
(231 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC7948 124 2e-29
TC7949 117 2e-27
TC9679 weakly similar to UP|Q8H1B5 (Q8H1B5) Hin1-like protein, p... 85 1e-17
TC8092 similar to UP|Q8MLI6 (Q8MLI6) CG30127-PA, partial (3%) 62 8e-11
TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%) 56 5e-09
AV427017 55 1e-08
TC14600 similar to UP|Q8S8Z8 (Q8S8Z8) Syringolide-induced protei... 52 1e-07
AV409132 51 2e-07
BI419489 45 1e-05
TC9423 similar to UP|Q8S8Z8 (Q8S8Z8) Syringolide-induced protein... 43 5e-05
TC11198 similar to UP|Q9LJT8 (Q9LJT8) Non-race specific disease ... 43 5e-05
TC10531 similar to UP|Q96UV7 (Q96UV7) Endo-1,4-beta-xylanase , ... 40 4e-04
AV418706 36 0.005
TC7968 similar to UP|Q39360 (Q39360) (Clone PIS63-1), partial (15%) 30 0.27
TC10007 30 0.35
TC10727 weakly similar to UP|Q15287 (Q15287) RNA-binding protein... 30 0.35
TC18699 weakly similar to UP|Q9FHE4 (Q9FHE4) Genomic DNA, chromo... 29 0.79
TC13196 28 1.0
TC12406 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, p... 28 1.3
TC18155 homologue to UP|O82161 (O82161) Phi-1 protein, partial (... 28 1.3
>TC7948
Length = 1230
Score = 124 bits (310), Expect = 2e-29
Identities = 77/228 (33%), Positives = 118/228 (50%), Gaps = 8/228 (3%)
Frame = +2
Query: 12 NQHHHLPPPNKIQMNNHKASPSPYGGGGGGSSPKRAACTFITVFLLAVGITLLVLWLVYR 71
+QHHH + + N+HK G + +R C I VFL V +T+L++W + R
Sbjct: 326 HQHHH--NMSMKECNHHK--------GAKRRAFRRVFCC-IVVFLFIVLVTILLIWAILR 472
Query: 72 PHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPIT 131
P KP F + VYA N T P L+ Q + RNPN+ + +YYDR S +V+YR+Q +T
Sbjct: 473 PTKPTFILQDVTVYAFNATVPNFLTTNFQVTLTSRNPNEHIGVYYDRLSTYVTYRSQQVT 652
Query: 132 QQVLLPPLFLEKHSQVSL-SPVIGGTPMPVTVEVANGLAMDESYGVVGLKLVFLGRVKWK 190
+ +PP + + H + + SP + G +PV GL D++ G V + + GRV+WK
Sbjct: 653 YRTAIPPSY-QGHKEFDVWSPFVSGNSVPVAPFNFVGLGQDQANGNVFITVKVDGRVRWK 829
Query: 191 AGGLRTWHHGLYVKCDLLVGL-KKG---FVGQ---VPLLQAQACDVDL 231
G + + LYV+C + L +G FVG+ V Q C V +
Sbjct: 830 VGAFVSGRYHLYVRCPAYISLGSRGNGVFVGENNAVKYQVVQRCSVSV 973
>TC7949
Length = 1183
Score = 117 bits (293), Expect = 2e-27
Identities = 61/188 (32%), Positives = 103/188 (54%), Gaps = 8/188 (4%)
Frame = +1
Query: 52 ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKR 111
I +FL V +T+L++W + +P+KP FT+ VYA N T L++ Q + RNPN +
Sbjct: 268 IVIFLFIVLVTILIIWAILKPNKPTFTLQDVTVYAFNATMANFLTSNFQVTLSARNPNDK 447
Query: 112 VSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVIGGTPMPVTVEVANGLAM 170
+ +YYDR A+++Y++Q +T + +PP + + H ++ + SP + GT +PV L+
Sbjct: 448 IGVYYDRLDAYITYQSQQVTYRTAIPPSY-QGHKEMDVWSPFVYGTNVPVAPYNFVTLSQ 624
Query: 171 DESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLV-------GLKKGFVGQVPLLQ 223
D++ G V + + G+V+WK G T H+ L V+C + G+ G G V
Sbjct: 625 DQANGNVLVTVKVDGKVRWKVGAFVTGHYHLNVRCPAFITFGARSNGVAVGDSGAVKYQL 804
Query: 224 AQACDVDL 231
Q C V +
Sbjct: 805 VQRCSVSV 828
>TC9679 weakly similar to UP|Q8H1B5 (Q8H1B5) Hin1-like protein, partial
(54%)
Length = 710
Score = 84.7 bits (208), Expect = 1e-17
Identities = 63/204 (30%), Positives = 95/204 (45%), Gaps = 8/204 (3%)
Frame = +2
Query: 17 LPPPNKIQMNNHKASPSPYGGGGGGSSPKRAACTFITVF------LLAVGITLLVLWLVY 70
+PPP K H+ S GGGG C F +F L+ +GI +L+ WL+
Sbjct: 98 VPPPKKTY---HRPSR-----GGGGCFECCCGCIFSLIFKLIFTALIIIGIAVLIFWLIV 253
Query: 71 RPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPI 130
RP+ +F V A + N T L+ L N+ +RNPNKR+ IYYD A Y++
Sbjct: 254 RPNVVKFHVTDATLTQFNFTGNNTLNYDLTLNITVRNPNKRLGIYYDYIEARAMYQDVRF 433
Query: 131 TQQVLLPPLFLEKHSQVS-LSPVIGGT-PMPVTVEVANGLAMDESYGVVGLKLVFLGRVK 188
+ F + H + + +S GT +P+ E L D+ GV G+ + +V+
Sbjct: 434 DTEFF--DRFYQGHKKTNVISRQFKGTHVVPLGAEETKELNKDKEDGVYGIDVKMYLKVR 607
Query: 189 WKAGGLRTWHHGLYVKCDLLVGLK 212
+K G +T V C+L V LK
Sbjct: 608 FKLGVFKTKTLKPKVTCELRVPLK 679
>TC8092 similar to UP|Q8MLI6 (Q8MLI6) CG30127-PA, partial (3%)
Length = 1073
Score = 62.0 bits (149), Expect = 8e-11
Identities = 44/161 (27%), Positives = 74/161 (45%), Gaps = 5/161 (3%)
Frame = +3
Query: 54 VFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAAL---QFNVLIR--NP 108
+FL+ +GI V +LV+RP P +++ +V +N TSP SAA +F+V +R N
Sbjct: 327 IFLILLGIAAGVFYLVFRPKAPNYSIQTVSVKGMNLTSPSPSSAAAISPEFDVAVRANNG 506
Query: 109 NKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVTVEVANGL 168
N ++ IYY + S+ + LP + ++ V+ G + + L
Sbjct: 507 NDKIGIYYLKGSSVEMFYRDSRLCGGALPAFYQASNNVTVFKAVLKGDGVELAGSDRTAL 686
Query: 169 AMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLV 209
+ V L + VK K G ++TW + V CD+ V
Sbjct: 687 VNAVAKRSVPLTMKLRAPVKIKVGSVKTWKITVKVDCDVAV 809
>TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%)
Length = 1036
Score = 56.2 bits (134), Expect = 5e-09
Identities = 52/196 (26%), Positives = 79/196 (39%), Gaps = 9/196 (4%)
Frame = +2
Query: 45 KRAACTF-------ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSA 97
+R CTF I + LL +G + +L+YRPH P F+V + LN TS LS+
Sbjct: 179 RRCFCTFCFWLILIILILLLIIGGAGVAFYLLYRPHHPTFSVTSLKLSYLNLTSSSTLSS 358
Query: 98 ALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTP 157
V NPNK+ + ++ +P K + L I +
Sbjct: 359 KFDLTVAATNPNKKNIAFSYLPTSITILSGDVDVGDGTIPTFHHGKKNTTLLKSSISSSG 538
Query: 158 MPVTVEVANGL-AMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCD-LLVGLKKGF 215
+ + A L A +S + LK+ +VK G L+T G+ V CD + V L G
Sbjct: 539 HELDADAAKKLKASMKSKNGLPLKVKLDTKVKATMGKLKTPRVGIRVSCDGIRVTLPTGK 718
Query: 216 VGQVPLLQAQACDVDL 231
C+VD+
Sbjct: 719 KPATASTSNAKCNVDV 766
>AV427017
Length = 429
Score = 55.1 bits (131), Expect = 1e-08
Identities = 35/136 (25%), Positives = 57/136 (41%), Gaps = 17/136 (12%)
Frame = +2
Query: 17 LPPPNKIQMNNHKASPSPYGGGGGG-----------------SSPKRAACTFITVFLLAV 59
+PPP Q ++H+ + + GGGG G + C +T ++
Sbjct: 26 IPPPQ--QKSHHRPTHNDNGGGGCGLCSCFCGCFRCCCGCIANCILGLICKILTTIIIIA 199
Query: 60 GITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKRVSIYYDRF 119
+ + WL+ RP+ +F V A++ T+ L L N+ +RNPN RV +YYD
Sbjct: 200 AVAGFLFWLIVRPNVVKFRVTEASLTEFTYTNNTL-HYNLALNITVRNPNSRVGLYYDSI 376
Query: 120 SAFVSYRNQPITQQVL 135
Y + Q L
Sbjct: 377 ETTAFYHDARFASQNL 424
>TC14600 similar to UP|Q8S8Z8 (Q8S8Z8) Syringolide-induced protein B13-1-9,
partial (76%)
Length = 1052
Score = 51.6 bits (122), Expect = 1e-07
Identities = 56/244 (22%), Positives = 101/244 (40%), Gaps = 14/244 (5%)
Frame = +1
Query: 1 MEEDPNPPKGQNQHHHLPPPNKIQMNNHKASPSPYGGGGGGSSPKRAACTFITVF----- 55
M D P G +PP + + ++H+ + C VF
Sbjct: 124 MANDQQPLNGAYYGPAIPPSGQPRHSHHRG--------------RSCFCCLFGVFWKILI 261
Query: 56 --LLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKRVS 113
++ VG+ +L+ +L+ +P +F V A + N T+ L + N RNPNK+++
Sbjct: 262 TLIVLVGLVILIFYLIVQPRPFKFYVNEAKLTEFNYTNNSL-RYNMVLNFTARNPNKKLN 438
Query: 114 IYYDRFSAFVSYRNQPI-TQQVLLPPLFLEKHSQVSLSPV----IGGTPMPVTVEVANGL 168
IYYD+ A Y + + ++ + + + S SP+ G M + + + L
Sbjct: 439 IYYDKVEALAFYEDSRFASDDKVITHMNSFRQYKKSSSPMSATFSGQHVMVLDSDQLSEL 618
Query: 169 AMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGL--YVKCDLLVGLKKGFVGQVPLLQAQA 226
+ D+S V + + R++++ G +++ L VKCDL V L
Sbjct: 619 SKDKSAHVYDIYVKLHFRIRFRLGDF-IFNNDLKPKVKCDLKVPLDSN--ATFTFFAVTK 789
Query: 227 CDVD 230
CDVD
Sbjct: 790 CDVD 801
>AV409132
Length = 405
Score = 50.8 bits (120), Expect = 2e-07
Identities = 23/76 (30%), Positives = 39/76 (51%), Gaps = 11/76 (14%)
Frame = +1
Query: 52 ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALN-----------TTSPPLLSAALQ 100
+ F+ + + + ++W++ RP KPRF + A V+A N T +P L+ LQ
Sbjct: 157 VVAFIFLILLIIFLIWIILRPTKPRFMLQDATVFAFNLSTQQHTPPLLTPTPNTLTLTLQ 336
Query: 101 FNVLIRNPNKRVSIYY 116
+ NPN R+ +YY
Sbjct: 337 VTLSSHNPNARIGVYY 384
>BI419489
Length = 479
Score = 44.7 bits (104), Expect = 1e-05
Identities = 22/73 (30%), Positives = 37/73 (50%), Gaps = 9/73 (12%)
Frame = +1
Query: 49 CTFITVFLL-----AVGITLLVLWLVYRPHKPRFTVVGAAVYALNTT----SPPLLSAAL 99
C F T+ +L A I L+++YRPH+P F++ + +N T SPP L+
Sbjct: 256 CCFWTILILLFAALAAAIVGAALYVLYRPHRPEFSITNLRIAKMNLTTSADSPPHLTTLF 435
Query: 100 QFNVLIRNPNKRV 112
++ +NPN +
Sbjct: 436 NLTLIAKNPNNHL 474
>TC9423 similar to UP|Q8S8Z8 (Q8S8Z8) Syringolide-induced protein B13-1-9,
partial (53%)
Length = 969
Score = 42.7 bits (99), Expect = 5e-05
Identities = 25/85 (29%), Positives = 39/85 (45%), Gaps = 7/85 (8%)
Frame = +1
Query: 52 ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPP-------LLSAALQFNVL 104
+ +L + + L+ +L+ +P +F V A + N T+ LL L N
Sbjct: 190 LLAIILFLALVFLIFYLIVQPRAFKFHVTDAKLTQFNYTTTNNNNSNNHLLRYNLVLNFT 369
Query: 105 IRNPNKRVSIYYDRFSAFVSYRNQP 129
RNPNK+++IYYD A Y P
Sbjct: 370 ARNPNKKLNIYYDEVEAVAFYDGIP 444
>TC11198 similar to UP|Q9LJT8 (Q9LJT8) Non-race specific disease resistance
protein-like, partial (17%)
Length = 882
Score = 42.7 bits (99), Expect = 5e-05
Identities = 27/83 (32%), Positives = 42/83 (50%), Gaps = 6/83 (7%)
Frame = +2
Query: 49 CTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTT--SPPLLSAALQFNVLIR 106
C+ F+ +G+ L +WL R +P+ + + ALN T SP ++ L F + +
Sbjct: 80 CSRCLGFITTMGLAALFIWLSLRVDEPKCYIDYIYIPALNKTLNSPFTTNSTLNFTLKLV 259
Query: 107 NPNKRVSIYYD----RFSAFVSY 125
NPNK I YD RF+ FV +
Sbjct: 260 NPNKDKGIKYDAVRLRFAIFVDH 328
>TC10531 similar to UP|Q96UV7 (Q96UV7) Endo-1,4-beta-xylanase , partial
(6%)
Length = 387
Score = 39.7 bits (91), Expect = 4e-04
Identities = 34/123 (27%), Positives = 50/123 (40%), Gaps = 8/123 (6%)
Frame = +2
Query: 2 EEDPNPPKGQNQHHHLPPP-NKIQMNNHKASPSPYGGGGGGSSPKRAACTFITVFLLAVG 60
+E P P ++PPP + Q +HK Y C F T ++L
Sbjct: 56 KEPPPPQLNPAAAINIPPPAEQPQPRHHKGRTCCY-------------CLFRTFWILLAT 196
Query: 61 ITLL------VLWLVYRPHKPRFTVVGAAVYALNTTSPPL-LSAALQFNVLIRNPNKRVS 113
I LL V+W++ +P F A + N S L L N +RNPNK++
Sbjct: 197 ILLLACLIILVVWILIQPRSYTFHASQAKLTEFNYNSTTATLRHNLVVNYTVRNPNKKLK 376
Query: 114 IYY 116
IY+
Sbjct: 377 IYF 385
>AV418706
Length = 374
Score = 36.2 bits (82), Expect = 0.005
Identities = 24/102 (23%), Positives = 44/102 (42%), Gaps = 9/102 (8%)
Frame = +3
Query: 19 PPNKIQMNN-HKASPSPYGGGGGGSSPKRAACTF-------ITVFLLAVGITLLVLWLVY 70
PP K Q++ ++ + P S CT + + ++ +G VL+ +Y
Sbjct: 69 PPTKSQLHGPNRPTYRPRAAPPRRRSSNSCCCTLFCWLILILFILIILLGAAGAVLYFLY 248
Query: 71 RPHKPRFTVVGAAVYALN-TTSPPLLSAALQFNVLIRNPNKR 111
P +P FTV + ++S L++ Q + NPNK+
Sbjct: 249 HPQRPSFTVTSLKLSTFKLSSSSSTLTSKFQLTLSATNPNKK 374
>TC7968 similar to UP|Q39360 (Q39360) (Clone PIS63-1), partial (15%)
Length = 1193
Score = 30.4 bits (67), Expect = 0.27
Identities = 22/46 (47%), Positives = 24/46 (51%)
Frame = -3
Query: 36 GGGGGGSSPKRAACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVG 81
GGGGG + A I VF L G L+LWLVYR F VVG
Sbjct: 297 GGGGGRVQTRLMAIKAIAVFSLW-GEDWLLLWLVYR----YFIVVG 175
>TC10007
Length = 594
Score = 30.0 bits (66), Expect = 0.35
Identities = 15/33 (45%), Positives = 18/33 (54%), Gaps = 1/33 (3%)
Frame = -3
Query: 4 DPNPPKGQNQHHHLPP-PNKIQMNNHKASPSPY 35
+PNP NQHHH P PNK +H+ P Y
Sbjct: 145 NPNPSPPSNQHHHPQP*PNK---THHQHPPPSY 56
>TC10727 weakly similar to UP|Q15287 (Q15287) RNA-binding protein
(RNA-binding protein S1, serine-rich domain), partial
(15%)
Length = 932
Score = 30.0 bits (66), Expect = 0.35
Identities = 16/46 (34%), Positives = 19/46 (40%)
Frame = +3
Query: 5 PNPPKGQNQHHHLPPPNKIQMNNHKASPSPYGGGGGGSSPKRAACT 50
P PP + HH PPP +SP P G +SP A T
Sbjct: 15 PPPPPPHHNHHRHPPPT--------SSPLPAHGASSSTSPPSPAPT 128
>TC18699 weakly similar to UP|Q9FHE4 (Q9FHE4) Genomic DNA, chromosome 5, TAC
clone:K17O22 (AT5g45100/K17O22_9), partial (28%)
Length = 537
Score = 28.9 bits (63), Expect = 0.79
Identities = 15/44 (34%), Positives = 19/44 (43%), Gaps = 5/44 (11%)
Frame = -1
Query: 5 PNPPKG-----QNQHHHLPPPNKIQMNNHKASPSPYGGGGGGSS 43
P PP G + HHLPPP + H +P+P G S
Sbjct: 261 PPPPHGGGCCRNSTQHHLPPPPPRRGGYH*RAPAPVPSRNAGKS 130
>TC13196
Length = 490
Score = 28.5 bits (62), Expect = 1.0
Identities = 13/49 (26%), Positives = 21/49 (42%), Gaps = 3/49 (6%)
Frame = -2
Query: 4 DPNPPKGQNQHHHLPPPNKIQM---NNHKASPSPYGGGGGGSSPKRAAC 49
+P+PP+ + HH PP + N + +P P G P+ C
Sbjct: 207 EPSPPQPKRAPHHSNPPPRSSSEYPRNSEQNPQPAAAPPCGPHPRSCTC 61
>TC12406 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, partial (6%)
Length = 450
Score = 28.1 bits (61), Expect = 1.3
Identities = 10/22 (45%), Positives = 15/22 (67%)
Frame = +2
Query: 22 KIQMNNHKASPSPYGGGGGGSS 43
++Q+ A P P GGGGGG++
Sbjct: 272 QVQVQPQNAMPGPNGGGGGGAA 337
>TC18155 homologue to UP|O82161 (O82161) Phi-1 protein, partial (18%)
Length = 428
Score = 28.1 bits (61), Expect = 1.3
Identities = 11/30 (36%), Positives = 15/30 (49%)
Frame = +2
Query: 5 PNPPKGQNQHHHLPPPNKIQMNNHKASPSP 34
P+ P + H PP N NH+ +PSP
Sbjct: 305 PSQPNHPSPHGGKPPRNTTTSQNHQTTPSP 394
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,148,440
Number of Sequences: 28460
Number of extensions: 90270
Number of successful extensions: 1092
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 1015
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1069
length of query: 231
length of database: 4,897,600
effective HSP length: 87
effective length of query: 144
effective length of database: 2,421,580
effective search space: 348707520
effective search space used: 348707520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0105.5


