

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0105.17
(488 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
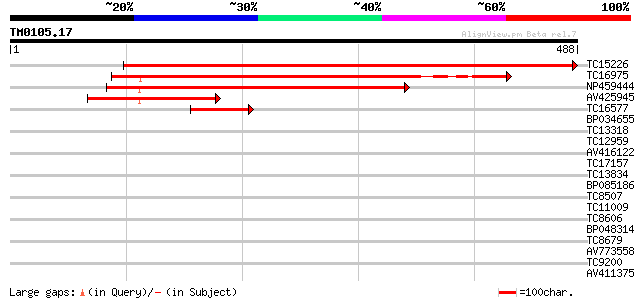
Score E
Sequences producing significant alignments: (bits) Value
TC15226 UP|O82459 (O82459) Rac GTPase activating protein 2 (Frag... 790 0.0
TC16975 UP|O82458 (O82458) Rac GTPase activating protein 1, part... 382 e-107
NP459444 rac GTPase activating protein 3 [Lotus japonicus] 373 e-104
AV425945 143 7e-35
TC16577 similar to UP|O82458 (O82458) Rac GTPase activating prot... 82 2e-16
BP034655 40 0.001
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 40 0.001
TC12959 similar to UP|AAS52042 (AAS52042) ADR122Cp, partial (6%) 38 0.003
AV416122 37 0.006
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 37 0.009
TC13834 homologue to UP|Q948R9 (Q948R9) RPS6-like protein (AT3g1... 36 0.016
BP085186 35 0.021
TC8507 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetyla... 35 0.028
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 35 0.028
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 34 0.047
BP048314 34 0.047
TC8679 homologue to UP|Q43466 (Q43466) Protein kinase 3 , partia... 34 0.061
AV773558 33 0.080
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 33 0.080
AV411375 33 0.10
>TC15226 UP|O82459 (O82459) Rac GTPase activating protein 2 (Fragment),
partial (92%)
Length = 1436
Score = 790 bits (2040), Expect = 0.0
Identities = 390/390 (100%), Positives = 390/390 (100%)
Frame = +3
Query: 99 KSLVTCSVDREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPTELQPEVPQKVPSASAKV 158
KSLVTCSVDREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPTELQPEVPQKVPSASAKV
Sbjct: 3 KSLVTCSVDREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPTELQPEVPQKVPSASAKV 182
Query: 159 FGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVRCQLNR 218
FGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVRCQLNR
Sbjct: 183 FGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEEFVRCQLNR 362
Query: 219 GLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTNLVKLLPSTEAALL 278
GLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTNLVKLLPSTEAALL
Sbjct: 363 GLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEEDCTNLVKLLPSTEAALL 542
Query: 279 DWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKTLILKTL 338
DWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKTLILKTL
Sbjct: 543 DWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALIHAVQVMNFLKTLILKTL 722
Query: 339 RERDESMAKARQLSSLLNSPSCKGDSHPFKDNREESSAQPVDTCATMPPDKSEFSRMEWC 398
RERDESMAKARQLSSLLNSPSCKGDSHPFKDNREESSAQPVDTCATMPPDKSEFSRMEWC
Sbjct: 723 RERDESMAKARQLSSLLNSPSCKGDSHPFKDNREESSAQPVDTCATMPPDKSEFSRMEWC 902
Query: 399 VDEKVWSSEEKGTGGGALESVSGGSSPSRYESGPLESRYRGIYDSEHWLRLRKGVRRLCQ 458
VDEKVWSSEEKGTGGGALESVSGGSSPSRYESGPLESRYRGIYDSEHWLRLRKGVRRLCQ
Sbjct: 903 VDEKVWSSEEKGTGGGALESVSGGSSPSRYESGPLESRYRGIYDSEHWLRLRKGVRRLCQ 1082
Query: 459 HPVFQLSKSTKKRADLGIVNTREGGGEAWA 488
HPVFQLSKSTKKRADLGIVNTREGGGEAWA
Sbjct: 1083HPVFQLSKSTKKRADLGIVNTREGGGEAWA 1172
>TC16975 UP|O82458 (O82458) Rac GTPase activating protein 1, partial (88%)
Length = 1686
Score = 382 bits (981), Expect = e-107
Identities = 205/352 (58%), Positives = 253/352 (71%), Gaps = 7/352 (1%)
Frame = +1
Query: 88 AILDILVAALKKSLV-TCSVDREDV-----SSLDISWPTEVRHVSHVTFDRFNGFLGLPT 141
++L +L+A +KSL+ +CS D SS++I WP+ VRHV+HVTFDRF+GFLGLP
Sbjct: 4 SLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGFLGLPV 183
Query: 142 ELQPEVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFR 201
E +PEVP++ PSAS VFGVS +SMQ S+D RGNSVPTILL+MQ LY+ GGL+AEGIFR
Sbjct: 184 EFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFR 363
Query: 202 INADNSQEEFVRCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCNSEE 261
INA+NSQEE VR QLNRG+VP GV+VHCL+GLIKAWFRELPTG+LD L+PE+VM SEE
Sbjct: 364 INAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEE 543
Query: 262 DCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLTALI 321
+C LV+LLP TEAALLDWAINLMADV + E FNKMNARNIAMVFAPNMT M DPLTAL+
Sbjct: 544 ECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALM 723
Query: 322 HAVQVMNFLKTLILKTLRERDESMAKARQLSSLLNSPSCKGDSHPFKDNREESSAQPVDT 381
+AVQVMNFLKTL++KTLR R+ES+ K+ + +L + F D+ +S +Q
Sbjct: 724 YAVQVMNFLKTLVVKTLRVREESIVKSNPVPNL----------NSFDDDGHQSDSQ---- 861
Query: 382 CATMPPDKSEFSRMEWCVDE-KVWSSEEKGTGGGALESVSGGSSPSRYESGP 432
+P D SE C DE V+ S E + G + S E+ P
Sbjct: 862 --VLPKDGSENGND--CSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSP 1005
>NP459444 rac GTPase activating protein 3 [Lotus japonicus]
Length = 1300
Score = 373 bits (958), Expect = e-104
Identities = 187/266 (70%), Positives = 218/266 (81%), Gaps = 5/266 (1%)
Frame = +2
Query: 84 QNQFAILDILVAALKKSLVTCSVDRED-----VSSLDISWPTEVRHVSHVTFDRFNGFLG 138
QNQ + L+AALKKS+V CSVD D V ++I WPT V+HV+HVTFDRFNGFLG
Sbjct: 14 QNQGSPAAFLLAALKKSMVACSVDSPDDVISAVHPMEIGWPTNVKHVTHVTFDRFNGFLG 193
Query: 139 LPTELQPEVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEG 198
LP EL+ VP VPSAS VFGVSA+SM CSYD +GNSVPTILL+MQ RLYS+GGL AEG
Sbjct: 194 LPLELEVHVPAPVPSASVSVFGVSAESMHCSYDSKGNSVPTILLLMQERLYSQGGLMAEG 373
Query: 199 IFRINADNSQEEFVRCQLNRGLVPRGVEVHCLSGLIKAWFRELPTGVLDSLTPEQVMHCN 258
IFRIN +N QEE +R QLNRG+VP ++VHCL+GLIKAWFRELP+GVLD L+PEQV+ CN
Sbjct: 374 IFRINPENGQEEHLRDQLNRGVVPDNIDVHCLAGLIKAWFRELPSGVLDGLSPEQVLECN 553
Query: 259 SEEDCTNLVKLLPSTEAALLDWAINLMADVVEHEQFNKMNARNIAMVFAPNMTQMVDPLT 318
+EE+ LVK L TE ALL+WA++LMADVVE E+ NKM+ARNIAMVFAPNMTQM DPLT
Sbjct: 554 TEEEFVQLVKQLKPTELALLNWALDLMADVVEEEEHNKMDARNIAMVFAPNMTQMSDPLT 733
Query: 319 ALIHAVQVMNFLKTLILKTLRERDES 344
AL+HAVQVMN LKTLILKTL ER+E+
Sbjct: 734 ALMHAVQVMNLLKTLILKTLSEREEA 811
>AV425945
Length = 415
Score = 143 bits (360), Expect = 7e-35
Identities = 66/119 (55%), Positives = 88/119 (73%), Gaps = 5/119 (4%)
Frame = +1
Query: 68 GSKDKGNWRGQSHNQNQNQFAILDILVAALKKSLVTCSVDRED-----VSSLDISWPTEV 122
G + + + + QNQ +++ +L+AAL+KS+V C VDR D V ++I WPT+V
Sbjct: 58 GKRTRASRTAEEEQDRQNQLSLVALLLAALRKSMVACRVDRPDEAISTVHQMEIGWPTDV 237
Query: 123 RHVSHVTFDRFNGFLGLPTELQPEVPQKVPSASAKVFGVSAKSMQCSYDERGNSVPTIL 181
+H++HVTFDRFNGFLGLP E + E+P +VPSAS VFGVSA+SMQCSYD +GNSVPTIL
Sbjct: 238 QHITHVTFDRFNGFLGLPVEFEVEIPGRVPSASVSVFGVSAESMQCSYDSKGNSVPTIL 414
>TC16577 similar to UP|O82458 (O82458) Rac GTPase activating protein 1,
partial (16%)
Length = 726
Score = 82.0 bits (201), Expect = 2e-16
Identities = 39/55 (70%), Positives = 45/55 (80%)
Frame = +1
Query: 156 AKVFGVSAKSMQCSYDERGNSVPTILLMMQNRLYSEGGLKAEGIFRINADNSQEE 210
A +FGVS +SMQ SYD RGNSVPTILL MQ LY++GGL+AEGIFRIN DN+ E
Sbjct: 166 ATMFGVSTESMQLSYDTRGNSVPTILLQMQRHLYAQGGLQAEGIFRINTDNTHTE 330
>BP034655
Length = 517
Score = 39.7 bits (91), Expect = 0.001
Identities = 16/26 (61%), Positives = 23/26 (87%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E++EEEEEEEEE D+EDD
Sbjct: 157 EEEEEEEEEEEEEEEEEEEEEDEEDD 234
Score = 39.3 bits (90), Expect = 0.001
Identities = 16/26 (61%), Positives = 23/26 (87%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E++EEEEEEEEE DDE+D
Sbjct: 166 EEEEEEEEEEEEEEEEEEDEEDDEED 243
Score = 38.1 bits (87), Expect = 0.003
Identities = 15/26 (57%), Positives = 23/26 (87%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E++EEEEEEEEE +DE+D
Sbjct: 154 EEEEEEEEEEEEEEEEEEEEEEDEED 231
Score = 37.4 bits (85), Expect = 0.006
Identities = 14/26 (53%), Positives = 24/26 (91%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E++EEEEEEEE+ +D+ED+
Sbjct: 169 EEEEEEEEEEEEEEEEEDEEDDEEDE 246
Score = 37.0 bits (84), Expect = 0.007
Identities = 13/27 (48%), Positives = 24/27 (88%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDDY 54
++EE++E++EEEEE+EE+ D+++DY
Sbjct: 178 EEEEEEEEEEEEEEDEEDDEEDEDEDY 258
Score = 36.6 bits (83), Expect = 0.009
Identities = 14/26 (53%), Positives = 23/26 (87%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E++EEEEEEEEE ++ED+
Sbjct: 148 EEEEEEEEEEEEEEEEEEEEEEEEDE 225
Score = 36.6 bits (83), Expect = 0.009
Identities = 14/26 (53%), Positives = 23/26 (87%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E++EEEEEEEEE ++E+D
Sbjct: 145 EEEEEEEEEEEEEEEEEEEEEEEEED 222
Score = 36.2 bits (82), Expect = 0.012
Identities = 14/25 (56%), Positives = 22/25 (88%)
Frame = +1
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DEE++E++EEEEEEEEE ++E++
Sbjct: 142 DEEEEEEEEEEEEEEEEEEEEEEEE 216
Score = 36.2 bits (82), Expect = 0.012
Identities = 14/26 (53%), Positives = 23/26 (87%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E++EEEEEE+EE +DED+
Sbjct: 175 EEEEEEEEEEEEEEEDEEDDEEDEDE 252
Score = 35.0 bits (79), Expect = 0.028
Identities = 13/25 (52%), Positives = 22/25 (88%)
Frame = +1
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
D+E++E++EEEEEEEEE ++E++
Sbjct: 139 DDEEEEEEEEEEEEEEEEEEEEEEE 213
Score = 34.3 bits (77), Expect = 0.047
Identities = 12/27 (44%), Positives = 23/27 (84%)
Frame = +1
Query: 27 VKDEEDDEDDEEEEEEEEELYNDDEDD 53
+ D++++E++EEEEEEEEE ++E++
Sbjct: 130 LSDDDEEEEEEEEEEEEEEEEEEEEEE 210
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 39.7 bits (91), Expect = 0.001
Identities = 14/35 (40%), Positives = 28/35 (80%)
Frame = +3
Query: 19 PPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
PP P + ++EEDD+++E++E+E+++ +DD+DD
Sbjct: 225 PPKPVQTDNEEEEDDDEEEDDEDEDDDDDDDDDDD 329
Score = 35.8 bits (81), Expect = 0.016
Identities = 14/32 (43%), Positives = 25/32 (77%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDDYWSNPV 59
+D++DD+DD+++ EEEE+ +DDE++ S V
Sbjct: 294 EDDDDDDDDDDDGEEEEDDDDDDEEEEGSEEV 389
Score = 35.4 bits (80), Expect = 0.021
Identities = 12/26 (46%), Positives = 23/26 (88%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+DE+DD+DD+++++ EEE +DD+D+
Sbjct: 288 EDEDDDDDDDDDDDGEEEEDDDDDDE 365
Score = 32.7 bits (73), Expect = 0.14
Identities = 10/25 (40%), Positives = 21/25 (84%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE++D+DD+++++++ E DD+DD
Sbjct: 285 DEDEDDDDDDDDDDDGEEEEDDDDD 359
Score = 32.7 bits (73), Expect = 0.14
Identities = 13/25 (52%), Positives = 20/25 (80%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDED 52
++EEDD+DD+EEEE EE+ + +D
Sbjct: 333 EEEEDDDDDDEEEEGSEEVEVEGKD 407
Score = 31.2 bits (69), Expect = 0.40
Identities = 11/28 (39%), Positives = 24/28 (85%), Gaps = 2/28 (7%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEE--EELYNDDEDD 53
+D+ED++DD+++++++ EE +DD+DD
Sbjct: 279 EDDEDEDDDDDDDDDDDGEEEEDDDDDD 362
>TC12959 similar to UP|AAS52042 (AAS52042) ADR122Cp, partial (6%)
Length = 517
Score = 38.1 bits (87), Expect = 0.003
Identities = 15/39 (38%), Positives = 27/39 (68%), Gaps = 2/39 (5%)
Frame = +1
Query: 19 PPSPFFHNVKD--EEDDEDDEEEEEEEEELYNDDEDDYW 55
P S F+ ++ D DD+DD+++ E+E++ N+D+DD W
Sbjct: 85 PSSRFWDSLSDGSSSDDDDDDDDSEDEDDDDNEDDDDEW 201
>AV416122
Length = 414
Score = 37.4 bits (85), Expect = 0.006
Identities = 17/26 (65%), Positives = 22/26 (84%)
Frame = +3
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EEDDE+D+EEE+EEE DDEDD
Sbjct: 303 EEEEDDEEDKEEEDEEE----DDEDD 368
Score = 35.0 bits (79), Expect = 0.028
Identities = 13/23 (56%), Positives = 21/23 (90%)
Frame = +3
Query: 31 EDDEDDEEEEEEEEELYNDDEDD 53
E++EDDEE++EEE+E +D++DD
Sbjct: 303 EEEEDDEEDKEEEDEEEDDEDDD 371
Score = 28.5 bits (62), Expect = 2.6
Identities = 10/24 (41%), Positives = 19/24 (78%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDED 52
DEED E+++EEE++E++ N + +
Sbjct: 318 DEEDKEEEDEEEDDEDDDSNGEAE 389
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 36.6 bits (83), Expect = 0.009
Identities = 15/25 (60%), Positives = 23/25 (92%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE++DEDD+++EEEE++ +DDEDD
Sbjct: 794 DEDEDEDDDDDEEEEDD-DDDDEDD 865
Score = 35.4 bits (80), Expect = 0.021
Identities = 12/25 (48%), Positives = 22/25 (88%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE+ DED++E+++++EE +DD+DD
Sbjct: 782 DEDGDEDEDEDDDDDEEEEDDDDDD 856
Score = 34.3 bits (77), Expect = 0.047
Identities = 11/26 (42%), Positives = 22/26 (84%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+D+ED ++DE+E+++++E DD+DD
Sbjct: 776 EDDEDGDEDEDEDDDDDEEEEDDDDD 853
Score = 33.1 bits (74), Expect = 0.10
Identities = 11/26 (42%), Positives = 23/26 (88%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+D ++DED++++++EEEE +DD++D
Sbjct: 785 EDGDEDEDEDDDDDEEEEDDDDDDED 862
Score = 29.3 bits (64), Expect = 1.5
Identities = 11/24 (45%), Positives = 20/24 (82%)
Frame = +2
Query: 30 EEDDEDDEEEEEEEEELYNDDEDD 53
E+ +EDDE+ +E+E+E +DDE++
Sbjct: 764 EDIEEDDEDGDEDEDEDDDDDEEE 835
Score = 28.9 bits (63), Expect = 2.0
Identities = 10/17 (58%), Positives = 16/17 (93%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEE 44
++EEDD+DD+E++EE E
Sbjct: 827 EEEEDDDDDDEDDEEGE 877
Score = 28.1 bits (61), Expect = 3.4
Identities = 8/17 (47%), Positives = 17/17 (99%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEE 45
D+E++EDD++++E++EE
Sbjct: 821 DDEEEEDDDDDDEDDEE 871
Score = 27.7 bits (60), Expect = 4.4
Identities = 9/17 (52%), Positives = 16/17 (93%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEE 45
+EE+D+DD++E++EE E
Sbjct: 827 EEEEDDDDDDEDDEEGE 877
Score = 27.7 bits (60), Expect = 4.4
Identities = 12/25 (48%), Positives = 19/25 (76%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
D ED E+D+E+ +E+E+ DD+DD
Sbjct: 758 DFEDIEEDDEDGDEDED--EDDDDD 826
>TC13834 homologue to UP|Q948R9 (Q948R9) RPS6-like protein
(AT3g17170/K14A17_29), partial (13%)
Length = 517
Score = 35.8 bits (81), Expect = 0.016
Identities = 14/34 (41%), Positives = 23/34 (67%)
Frame = +1
Query: 19 PPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDED 52
PP P FH + DD+DDEE +E+ ++ ++ +ED
Sbjct: 112 PPPPEFHTLGASVDDDDDEELDEDYDDDWDGEED 213
>BP085186
Length = 388
Score = 35.4 bits (80), Expect = 0.021
Identities = 16/36 (44%), Positives = 25/36 (69%)
Frame = -1
Query: 26 NVKDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVST 61
N ++EE++E++EEEEEEEEE +E+ V+T
Sbjct: 262 NEEEEEEEEEEEEEEEEEEEEAEPEEEKNEEENVNT 155
Score = 33.5 bits (75), Expect = 0.080
Identities = 15/36 (41%), Positives = 25/36 (68%)
Frame = -1
Query: 18 SPPSPFFHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
SP + ++EE++E++EEEEEEEEE + E++
Sbjct: 289 SPDNADGEGNEEEEEEEEEEEEEEEEEEEEAEPEEE 182
Score = 28.5 bits (62), Expect = 2.6
Identities = 14/31 (45%), Positives = 21/31 (67%), Gaps = 1/31 (3%)
Frame = -1
Query: 29 DEEDDE-DDEEEEEEEEELYNDDEDDYWSNP 58
D D E ++EEEEEEEEE ++E++ + P
Sbjct: 283 DNADGEGNEEEEEEEEEEEEEEEEEEEEAEP 191
Score = 27.3 bits (59), Expect = 5.7
Identities = 11/26 (42%), Positives = 18/26 (68%)
Frame = -1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++EE++E + EEE+ EEE N E +
Sbjct: 220 EEEEEEEAEPEEEKNEEENVNTQEGE 143
>TC8507 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase
HD2-P39, partial (40%)
Length = 620
Score = 35.0 bits (79), Expect = 0.028
Identities = 19/50 (38%), Positives = 28/50 (56%), Gaps = 2/50 (4%)
Frame = +3
Query: 29 DEEDDEDDEEEEEE--EEELYNDDEDDYWSNPVSTPLINPAGSKDKGNWR 76
DEEDD DDE++E++ +EE+ + D D S+ T P D+G R
Sbjct: 3 DEEDDSDDEDDEDDDSDEEMDDADSDSDESDSDDTDEETPVKKADQGKKR 152
Score = 34.3 bits (77), Expect = 0.047
Identities = 16/46 (34%), Positives = 24/46 (51%)
Frame = +3
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSKDKGN 74
D+EDDEDD+ +EE ++ + DE D TP+ K + N
Sbjct: 21 DDEDDEDDDSDEEMDDADSDSDESDSDDTDEETPVKKADQGKKRAN 158
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 35.0 bits (79), Expect = 0.028
Identities = 13/18 (72%), Positives = 18/18 (99%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEEL 46
D++DD+DD+EEEEEEE+L
Sbjct: 182 DDDDDDDDDEEEEEEEDL 235
Score = 33.5 bits (75), Expect = 0.080
Identities = 12/17 (70%), Positives = 17/17 (99%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEE 45
D++DD+DD++EEEEEEE
Sbjct: 179 DDDDDDDDDDEEEEEEE 229
Score = 32.3 bits (72), Expect = 0.18
Identities = 11/17 (64%), Positives = 17/17 (99%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEE 45
D++DD+DD+++EEEEEE
Sbjct: 176 DDDDDDDDDDDEEEEEE 226
Score = 30.0 bits (66), Expect = 0.89
Identities = 11/26 (42%), Positives = 19/26 (72%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDDY 54
D +DD+DD+++++EEEE D +Y
Sbjct: 170 DPDDDDDDDDDDDEEEEEEEDLGTEY 247
Score = 27.7 bits (60), Expect = 4.4
Identities = 13/36 (36%), Positives = 22/36 (61%), Gaps = 11/36 (30%)
Frame = +2
Query: 29 DEEDDEDDEEEE-----------EEEEELYNDDEDD 53
D++DDE+DEEEE + +++ +DD+DD
Sbjct: 101 DDDDDEEDEEEEGGVEGGRGGGGDPDDDDDDDDDDD 208
Score = 27.7 bits (60), Expect = 4.4
Identities = 14/30 (46%), Positives = 18/30 (59%), Gaps = 8/30 (26%)
Frame = +2
Query: 32 DDEDDEEEEEEEEELYN--------DDEDD 53
DD+DDEE+EEEE + DD+DD
Sbjct: 101 DDDDDEEDEEEEGGVEGGRGGGGDPDDDDD 190
Score = 26.9 bits (58), Expect = 7.5
Identities = 10/27 (37%), Positives = 20/27 (74%), Gaps = 2/27 (7%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEEL--YNDDEDD 53
D++D EDD+ +E+++++ DD+DD
Sbjct: 35 DDDDGEDDDGDEDDDDDAPGGGDDDDD 115
Score = 26.9 bits (58), Expect = 7.5
Identities = 11/25 (44%), Positives = 18/25 (72%)
Frame = +2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DE+DD+D E+++ +E DD+DD
Sbjct: 26 DEDDDDDGEDDDGDE-----DDDDD 85
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 34.3 bits (77), Expect = 0.047
Identities = 11/26 (42%), Positives = 23/26 (88%)
Frame = -2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+D+EDD+DD+E+++EE++ D+E++
Sbjct: 407 EDQEDDDDDDEDDDEEDDGGEDEEEE 330
Score = 32.3 bits (72), Expect = 0.18
Identities = 14/25 (56%), Positives = 19/25 (76%)
Frame = -2
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
DEEDD ++EEEE EE N+DE++
Sbjct: 368 DEEDDGGEDEEEEGVEEEDNEDEEE 294
Score = 32.3 bits (72), Expect = 0.18
Identities = 16/49 (32%), Positives = 26/49 (52%), Gaps = 5/49 (10%)
Frame = -2
Query: 29 DEEDDEDDEEE-----EEEEEELYNDDEDDYWSNPVSTPLINPAGSKDK 72
D++DDEDD+EE +EEEE + +D +D + + P + K
Sbjct: 392 DDDDDEDDDEEDDGGEDEEEEGVEEEDNEDEEEDEEDEEALQPPKKRKK 246
Score = 32.3 bits (72), Expect = 0.18
Identities = 10/28 (35%), Positives = 24/28 (85%)
Frame = -2
Query: 26 NVKDEEDDEDDEEEEEEEEELYNDDEDD 53
N +DEED++ +++E++++++ +D+EDD
Sbjct: 437 NGEDEEDEDGEDQEDDDDDDEDDDEEDD 354
Score = 29.6 bits (65), Expect = 1.2
Identities = 11/24 (45%), Positives = 20/24 (82%)
Frame = -2
Query: 30 EEDDEDDEEEEEEEEELYNDDEDD 53
EE+ ED+E+E+ E++E +DD++D
Sbjct: 443 EENGEDEEDEDGEDQEDDDDDDED 372
Score = 28.9 bits (63), Expect = 2.0
Identities = 10/26 (38%), Positives = 20/26 (76%)
Frame = -2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
+D ED EDD++++E+++E + ED+
Sbjct: 416 EDGEDQEDDDDDDEDDDEEDDGGEDE 339
Score = 28.1 bits (61), Expect = 3.4
Identities = 9/26 (34%), Positives = 21/26 (80%)
Frame = -2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++ +DE+DE+ E++E++ +D++DD
Sbjct: 443 EENGEDEEDEDGEDQEDDDDDDEDDD 366
Score = 27.3 bits (59), Expect = 5.7
Identities = 11/24 (45%), Positives = 18/24 (74%)
Frame = -2
Query: 28 KDEEDDEDDEEEEEEEEELYNDDE 51
+DEE++ +EE+ E+EEE D+E
Sbjct: 347 EDEEEEGVEEEDNEDEEEDEEDEE 276
>BP048314
Length = 394
Score = 34.3 bits (77), Expect = 0.047
Identities = 13/25 (52%), Positives = 20/25 (80%)
Frame = +1
Query: 25 HNVKDEEDDEDDEEEEEEEEELYND 49
H ++EE++E++EEEEEEEE + D
Sbjct: 181 HEEEEEEEEEEEEEEEEEEEAMCKD 255
Score = 32.3 bits (72), Expect = 0.18
Identities = 13/18 (72%), Positives = 17/18 (94%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEE 45
K EE++E++EEEEEEEEE
Sbjct: 178 KHEEEEEEEEEEEEEEEE 231
Score = 29.6 bits (65), Expect = 1.2
Identities = 11/17 (64%), Positives = 16/17 (93%)
Frame = +1
Query: 29 DEEDDEDDEEEEEEEEE 45
D+ ++E++EEEEEEEEE
Sbjct: 175 DKHEEEEEEEEEEEEEE 225
Score = 28.5 bits (62), Expect = 2.6
Identities = 11/22 (50%), Positives = 17/22 (77%)
Frame = +1
Query: 32 DDEDDEEEEEEEEELYNDDEDD 53
D ++EEEEEEEEE ++E++
Sbjct: 175 DKHEEEEEEEEEEEEEEEEEEE 240
Score = 28.5 bits (62), Expect = 2.6
Identities = 11/22 (50%), Positives = 18/22 (81%)
Frame = +1
Query: 30 EEDDEDDEEEEEEEEELYNDDE 51
++ +E++EEEEEEEEE ++E
Sbjct: 175 DKHEEEEEEEEEEEEEEEEEEE 240
>TC8679 homologue to UP|Q43466 (Q43466) Protein kinase 3 , partial (70%)
Length = 972
Score = 33.9 bits (76), Expect = 0.061
Identities = 16/25 (64%), Positives = 20/25 (80%)
Frame = +2
Query: 30 EEDDEDDEEEEEEEEELYNDDEDDY 54
EE+DE+D EEE EEEE D+ED+Y
Sbjct: 641 EEEDEEDVEEEVEEEE---DEEDEY 706
Score = 30.0 bits (66), Expect = 0.89
Identities = 12/21 (57%), Positives = 18/21 (85%)
Frame = +2
Query: 28 KDEEDDEDDEEEEEEEEELYN 48
+DEED E++ EEEE+EE+ Y+
Sbjct: 647 EDEEDVEEEVEEEEDEEDEYD 709
>AV773558
Length = 469
Score = 33.5 bits (75), Expect = 0.080
Identities = 14/44 (31%), Positives = 26/44 (58%), Gaps = 10/44 (22%)
Frame = -2
Query: 20 PSPFFHNVKDEE----------DDEDDEEEEEEEEELYNDDEDD 53
P F N DEE +D+DD+++++++++ Y DDED+
Sbjct: 432 PDGLFFNEGDEELDDDFVLVYSEDDDDDDDDDDDDDDYTDDEDE 301
Score = 27.3 bits (59), Expect = 5.7
Identities = 9/17 (52%), Positives = 15/17 (87%)
Frame = -2
Query: 29 DEEDDEDDEEEEEEEEE 45
D++DD+DD ++E+EEE
Sbjct: 345 DDDDDDDDYTDDEDEEE 295
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 33.5 bits (75), Expect = 0.080
Identities = 17/43 (39%), Positives = 29/43 (66%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDDYWSNPVSTPLINPAGSK 70
++EE+ E++EEEEEEEEE+ + E++ P+ L+ P G +
Sbjct: 253 EEEEEGEEEEEEEEEEEEVEAEAEEED-DEPIQ-KLLEPMGKE 375
Score = 33.1 bits (74), Expect = 0.10
Identities = 13/30 (43%), Positives = 23/30 (76%)
Frame = +1
Query: 24 FHNVKDEEDDEDDEEEEEEEEELYNDDEDD 53
+ V++E ++E +EEEEEEEEE ++E++
Sbjct: 196 YEEVEEEVEEEVEEEEEEEEEEEEGEEEEE 285
Score = 33.1 bits (74), Expect = 0.10
Identities = 13/26 (50%), Positives = 22/26 (84%)
Frame = +1
Query: 27 VKDEEDDEDDEEEEEEEEELYNDDED 52
V++EE++E++EEE EEEEE ++E+
Sbjct: 229 VEEEEEEEEEEEEGEEEEEEEEEEEE 306
Score = 32.0 bits (71), Expect = 0.23
Identities = 12/25 (48%), Positives = 21/25 (84%)
Frame = +1
Query: 29 DEEDDEDDEEEEEEEEELYNDDEDD 53
+EE++E++EEEE EEEE ++E++
Sbjct: 232 EEEEEEEEEEEEGEEEEEEEEEEEE 306
Score = 31.6 bits (70), Expect = 0.30
Identities = 13/24 (54%), Positives = 20/24 (83%)
Frame = +1
Query: 30 EEDDEDDEEEEEEEEELYNDDEDD 53
EE+ E++EEEEEEEEE ++E++
Sbjct: 220 EEEVEEEEEEEEEEEEGEEEEEEE 291
Score = 31.2 bits (69), Expect = 0.40
Identities = 12/27 (44%), Positives = 22/27 (81%)
Frame = +1
Query: 27 VKDEEDDEDDEEEEEEEEELYNDDEDD 53
V++E ++E++EEEEEEE E ++E++
Sbjct: 217 VEEEVEEEEEEEEEEEEGEEEEEEEEE 297
Score = 29.6 bits (65), Expect = 1.2
Identities = 11/26 (42%), Positives = 21/26 (80%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDD 53
++ E++E++EEEEEE EE ++E++
Sbjct: 223 EEVEEEEEEEEEEEEGEEEEEEEEEE 300
Score = 26.6 bits (57), Expect = 9.8
Identities = 11/27 (40%), Positives = 20/27 (73%)
Frame = +1
Query: 27 VKDEEDDEDDEEEEEEEEELYNDDEDD 53
V++ E+ E++ EEE EEEE ++E++
Sbjct: 187 VQEYEEVEEEVEEEVEEEEEEEEEEEE 267
>AV411375
Length = 418
Score = 33.1 bits (74), Expect = 0.10
Identities = 13/29 (44%), Positives = 21/29 (71%)
Frame = +1
Query: 28 KDEEDDEDDEEEEEEEEELYNDDEDDYWS 56
+ +E++ED+E E EEEE Y+DD+ + S
Sbjct: 205 QQQEEEEDEEVEVEEEEASYSDDDTSFLS 291
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.132 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,687,319
Number of Sequences: 28460
Number of extensions: 125251
Number of successful extensions: 2619
Number of sequences better than 10.0: 168
Number of HSP's better than 10.0 without gapping: 1046
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1787
length of query: 488
length of database: 4,897,600
effective HSP length: 94
effective length of query: 394
effective length of database: 2,222,360
effective search space: 875609840
effective search space used: 875609840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0105.17


