

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0102b.4
(278 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
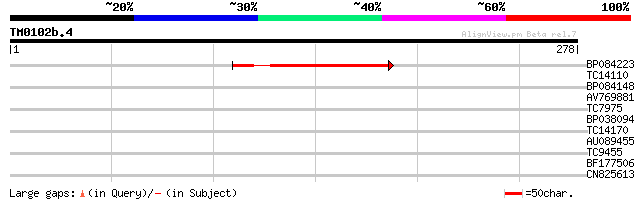
BP084223 80 5e-16
TC14110 homologue to UP|Q9AT55 (Q9AT55) S-adenosyl-L-methionine ... 32 0.16
BP084148 29 1.0
AV769881 27 5.0
TC7975 homologue to UP|Q9AT56 (Q9AT56) S-adenosyl-L-methionine s... 27 5.0
BP038094 26 6.5
TC14170 similar to UP|PSBS_LYCES (P54773) Photosystem II 22 kDa ... 26 8.5
AU089455 26 8.5
TC9455 similar to UP|Q9LKX5 (Q9LKX5) Acyl-CoA oxidase ACX3 (At1... 26 8.5
BF177506 26 8.5
CN825613 26 8.5
>BP084223
Length = 347
Score = 79.7 bits (195), Expect = 5e-16
Identities = 39/79 (49%), Positives = 56/79 (70%)
Frame = +3
Query: 110 INRVTWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFI 169
INR+ + T MDCFK+++ Q+GD+I +D T++F VLEF RL +RTT L+FI
Sbjct: 126 INRINYPGLTN-------MDCFKYMVTQLGDIIMVDPRTTDFSVLEFVRLHIRTTALDFI 284
Query: 170 TLARKILIKGTTYTISIIE 188
+LARK++I TTYTI ++E
Sbjct: 285 SLARKVMINETTYTIRVME 341
>TC14110 homologue to UP|Q9AT55 (Q9AT55) S-adenosyl-L-methionine synthetase
(S-adenosylmethionine synthetase) (Methionine
adenosyltransferase) (AdoMet synthetase) , complete
Length = 1576
Score = 31.6 bits (70), Expect = 0.16
Identities = 33/130 (25%), Positives = 57/130 (43%)
Frame = -2
Query: 89 SKGWRSEFVDNLKPWSSSFAPINRVTWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDT 148
S G S N+ PWS API+ +V+C P W + +L + T
Sbjct: 507 SSGVSSVAYPNICPWSP--APISSGRFVKCP*TP---WAISGLCCSMLTRTLQLSASRPT 343
Query: 149 SEFVVLEFSRLLVRTTTLNFITLARKILIKGTTYTISIIEEGEALLHPNCCCREESSVED 208
S F ++++ T + TLA ++ T + +++ +A L C R+ SS
Sbjct: 342 SSETKPMFLQVVL--TIFS*STLALVVISPKTITMLVLVQVSQATLLSGSCSRQASSTAS 169
Query: 209 DVFSGWSLNG 218
+++S SL+G
Sbjct: 168 EIWS-QSLSG 142
>BP084148
Length = 518
Score = 28.9 bits (63), Expect = 1.0
Identities = 18/82 (21%), Positives = 37/82 (44%)
Frame = -2
Query: 64 VRYLRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKPWSSSFAPINRVTWVRCTGIPL 123
+R L + ++ L L GS G + NL+ S+ P + +RCT + +
Sbjct: 469 IRLLPSCQRMIRRRLRMRLTRLSHGSMGTNLLRLMNLRTR*RSWRPFAIPSLLRCTKVQV 290
Query: 124 HMWTMDCFKHLLLQVGDVIKID 145
MW ++ + ++L + + +D
Sbjct: 289 LMWVVEPWMRMVLLLAVEVVLD 224
>AV769881
Length = 528
Score = 26.6 bits (57), Expect = 5.0
Identities = 14/53 (26%), Positives = 25/53 (46%)
Frame = -1
Query: 183 TISIIEEGEALLHPNCCCREESSVEDDVFSGWSLNGVTVTPELSESGGEDDDG 235
+++I+ E LH N CCR + F+ + + GV + + G + DG
Sbjct: 237 SLAILAELSGWLHSNACCRMVT------FTEFGIRGVRIREQRQVRVGLEQDG 97
>TC7975 homologue to UP|Q9AT56 (Q9AT56) S-adenosyl-L-methionine synthetase
(S-adenosylmethionine synthetase) (Methionine
adenosyltransferase) (AdoMet synthetase) , complete
Length = 1564
Score = 26.6 bits (57), Expect = 5.0
Identities = 34/135 (25%), Positives = 56/135 (41%), Gaps = 5/135 (3%)
Frame = -3
Query: 89 SKGWRSEFVDNLKPWSSSFAPINRVTWVRCT-----GIPLHMWTMDCFKHLLLQVGDVIK 143
S G S N+ PWS + R+ CT G+ M+T K L L
Sbjct: 506 SSGVSSVA*PNM*PWSPAPMSSGRLVKWPCTP*AISGLCCSMFT----KTLHLSASRPTS 339
Query: 144 IDEDTSEFVVLEFSRLLVRTTTLNFITLARKILIKGTTYTISIIEEGEALLHPNCCCREE 203
D + +V SR + +TL + ++ K T + +++ +A L C R+
Sbjct: 338 SDTNP---IVRHVSRTIFS*STLALVVISPK-----TMTMLVLVQVSQATLLSGSCSRQA 183
Query: 204 SSVEDDVFSGWSLNG 218
SS +++S SL+G
Sbjct: 182 SSTASEIWS-HSLSG 141
>BP038094
Length = 571
Score = 26.2 bits (56), Expect = 6.5
Identities = 12/29 (41%), Positives = 18/29 (61%)
Frame = +3
Query: 178 KGTTYTISIIEEGEALLHPNCCCREESSV 206
+ T+ + I +E LL P CC +EES+V
Sbjct: 405 RARTHNMRIPKELLCLLGPLCCLKEESTV 491
>TC14170 similar to UP|PSBS_LYCES (P54773) Photosystem II 22 kDa protein,
chloroplast precursor (CP22), partial (81%)
Length = 1201
Score = 25.8 bits (55), Expect = 8.5
Identities = 23/93 (24%), Positives = 38/93 (40%), Gaps = 1/93 (1%)
Frame = +3
Query: 134 LLLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFITL-ARKILIKGTTYTISIIEEGEA 192
L+ V +D + L+ RL + + L+F L + L T+T + + +A
Sbjct: 153 LMSSVSSSYSVDLKKDPLLHLQSQRLRPKFSQLSFNPLPSNSSLFSSRTFTTLALFKSKA 332
Query: 193 LLHPNCCCREESSVEDDVFSGWSLNGVTVTPEL 225
P +++ VED VF G T EL
Sbjct: 333 KAPPPKVVKQKPKVEDGVFGTSGGIGFTKQNEL 431
>AU089455
Length = 595
Score = 25.8 bits (55), Expect = 8.5
Identities = 12/23 (52%), Positives = 16/23 (69%)
Frame = +1
Query: 9 SKVDRGRMTALELWSGPEIPKEE 31
S VD+GR + EL + PE+ KEE
Sbjct: 214 SSVDKGRTDSKELITVPEMDKEE 282
>TC9455 similar to UP|Q9LKX5 (Q9LKX5) Acyl-CoA oxidase ACX3 (At1g06290) ,
partial (50%)
Length = 1361
Score = 25.8 bits (55), Expect = 8.5
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = +2
Query: 193 LLHPNCCCREESSVE 207
L++ CCCREE S E
Sbjct: 872 LVNTECCCREERSAE 916
>BF177506
Length = 447
Score = 25.8 bits (55), Expect = 8.5
Identities = 16/37 (43%), Positives = 21/37 (56%), Gaps = 4/37 (10%)
Frame = +1
Query: 224 ELSESGGEDDDGLWPVETKSQHDLQG----SPNHSLG 256
E E+GG+D +GL ++ QH LQG SP LG
Sbjct: 76 E*EEAGGDDGEGLAYLQGGLQHLLQGFQDPSPYAELG 186
>CN825613
Length = 257
Score = 25.8 bits (55), Expect = 8.5
Identities = 11/32 (34%), Positives = 19/32 (59%)
Frame = -3
Query: 234 DGLWPVETKSQHDLQGSPNHSLGDNKNPRSNT 265
D L+PV+ QH ++ PNH+ ++ R +T
Sbjct: 243 DQLYPVQMYIQHHVEIFPNHNDHQDQTTRDHT 148
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,506,702
Number of Sequences: 28460
Number of extensions: 81177
Number of successful extensions: 417
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 416
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 417
length of query: 278
length of database: 4,897,600
effective HSP length: 89
effective length of query: 189
effective length of database: 2,364,660
effective search space: 446920740
effective search space used: 446920740
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0102b.4


