

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0101.12
(310 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
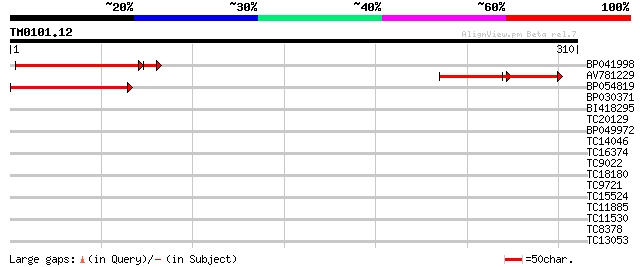
Score E
Sequences producing significant alignments: (bits) Value
BP041998 104 2e-24
AV781229 65 5e-23
BP054819 100 3e-22
BP030371 32 0.14
BI418295 32 0.18
TC20129 homologue to GB|AAF04891.1|6175165|ATAC011437 Mutator-li... 30 0.40
BP049972 30 0.40
TC14046 homologue to UP|P93769 (P93769) Elongation factor-1 alph... 29 0.89
TC16374 similar to UP|Q7LZA6 (Q7LZA6) Protamine II (Fragment), p... 29 1.2
TC9022 similar to PIR|T46225|T46225 alpha NAC-like protein - Ara... 28 2.6
TC18180 weakly similar to GB|AAF04891.1|6175165|ATAC011437 Mutat... 27 3.4
TC9721 similar to UP|Q9LTA1 (Q9LTA1) Genomic DNA, chromosome 5, ... 27 3.4
TC15524 similar to UP|Q8RWF0 (Q8RWF0) 26S proteasome subunit-lik... 27 5.8
TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 27 5.8
TC11530 26 7.5
TC8378 26 7.5
TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, parti... 26 9.8
>BP041998
Length = 485
Score = 104 bits (259), Expect(2) = 2e-24
Identities = 49/70 (70%), Positives = 58/70 (82%)
Frame = -3
Query: 4 FLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNAIFN 63
FLKSID KTWKA+V GWKHP K+Q EG+++T EKPE+E TS E+EA+L NSKA+NAIF
Sbjct: 240 FLKSIDNKTWKAIVNGWKHPTKSQVEGTSTTVVEKPEEEWTSVEDEAALGNSKALNAIFY 61
Query: 64 AVDKNMFRLI 73
VD NMFRLI
Sbjct: 60 GVDXNMFRLI 31
Score = 23.9 bits (50), Expect(2) = 2e-24
Identities = 8/10 (80%), Positives = 10/10 (100%)
Frame = -2
Query: 74 NTCTVAKDAW 83
++CTVAKDAW
Sbjct: 31 HSCTVAKDAW 2
>AV781229
Length = 208
Score = 64.7 bits (156), Expect(2) = 5e-23
Identities = 29/33 (87%), Positives = 31/33 (93%)
Frame = +3
Query: 270 QNKGVQCHECEGYGHIRSECATYLKKQKKGMVV 302
++KGV CHE EGYGHIRSECATYLKKQKKGMVV
Sbjct: 108 KSKGV*CHEWEGYGHIRSECATYLKKQKKGMVV 206
Score = 59.3 bits (142), Expect(2) = 5e-23
Identities = 30/39 (76%), Positives = 30/39 (76%)
Frame = +1
Query: 236 KIDRRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGV 274
KID RTR QDIKSDN KSYN QRKVKEEDKSGQ V
Sbjct: 4 KIDSRTRSKCQDIKSDNFKSYNFQRKVKEEDKSGQRARV 120
>BP054819
Length = 537
Score = 100 bits (249), Expect = 3e-22
Identities = 46/67 (68%), Positives = 56/67 (82%)
Frame = -3
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M AFLKSID KTWKA+VKGWKHP K + EG+++T EKPE+E TS E+EA+L NSKA+NA
Sbjct: 235 MTAFLKSIDNKTWKAIVKGWKHPTKVEVEGTSTTVVEKPEEEWTSVEDEAALGNSKALNA 56
Query: 61 IFNAVDK 67
IFN VD+
Sbjct: 55 IFNGVDR 35
>BP030371
Length = 516
Score = 32.0 bits (71), Expect = 0.14
Identities = 32/171 (18%), Positives = 69/171 (39%)
Frame = -2
Query: 119 ISEYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDEL 178
++EY M ++ +++ ++G+P+S ++ IL +K+ V I + +L +DE+
Sbjct: 509 VAEYLM*IKSITDPPRSIGDPVSAKEQAEIILEGFPTKYESLVNLIHMSDKFQALSLDEI 330
Query: 179 IGSLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKID 238
L+ + ++ + T+ ++ T E + S + +
Sbjct: 329 ESLLRDQEARIECYRKFLTEI*VNLT*TLENQGSSSANTYAPAPV--------------- 195
Query: 239 RRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSEC 289
+ + D ++N +R + G+ GVQC C GH S C
Sbjct: 194 -----HSAEASHDGYSNFNGKRCGGK--G*GRGYGVQCQVCLPTGHTASVC 63
>BI418295
Length = 523
Score = 31.6 bits (70), Expect = 0.18
Identities = 27/137 (19%), Positives = 54/137 (38%), Gaps = 14/137 (10%)
Frame = +2
Query: 19 GWKHPVKAQAEGSTS----TPEEKPEDELTSAEEEASLEN----------SKAMNAIFNA 64
GW+ V+ + E P+ LT + A+ EN S +
Sbjct: 11 GWRQQVEGAIQSHQLHDFLVDPEIPQRFLTEEDRVAATENPAYTTWIKQDSMVFTWLLTT 190
Query: 65 VDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYHM 124
+ + + C + W + E R + ++L+ + +N+T + I+EY
Sbjct: 191 LSPFVLPTVINCERSSQVWREILDFFENQCRAQSAKLR---SELKNITKGS-KSITEYLR 358
Query: 125 RVRDLSNASFALGEPMS 141
R++ + N ++GEP+S
Sbjct: 359 RIKTIVNTLVSIGEPVS 409
>TC20129 homologue to GB|AAF04891.1|6175165|ATAC011437 Mutator-like
transposase {Arabidopsis thaliana;} , partial (12%)
Length = 588
Score = 30.4 bits (67), Expect = 0.40
Identities = 11/32 (34%), Positives = 18/32 (55%)
Frame = +3
Query: 259 QRKVKEEDKSGQNKGVQCHECEGYGHIRSECA 290
+++V+ ED+ + V C C GH R+ CA
Sbjct: 174 KKRVRAEDRGRVKRVVHCSRCNQTGHFRTTCA 269
>BP049972
Length = 496
Score = 30.4 bits (67), Expect = 0.40
Identities = 11/32 (34%), Positives = 18/32 (55%)
Frame = -1
Query: 259 QRKVKEEDKSGQNKGVQCHECEGYGHIRSECA 290
+++V+ ED+ + V C C GH R+ CA
Sbjct: 286 KKRVRAEDRGRVKRVVHCSRCNQTGHFRTTCA 191
>TC14046 homologue to UP|P93769 (P93769) Elongation factor-1 alpha, partial
(84%)
Length = 1506
Score = 29.3 bits (64), Expect = 0.89
Identities = 24/83 (28%), Positives = 38/83 (44%), Gaps = 5/83 (6%)
Frame = +1
Query: 171 SSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVS-NTTEGDDADSESENL----SEALAL 225
S + DE++ + +Y K+G P+K I FV + EGD+ S NL L
Sbjct: 568 SKARYDEILKEVSSYLKKVGYNPDK----IPFVPISGFEGDNMIERSTNLDWYKGPTLLE 735
Query: 226 LARKFNRALRKIDRRTRPNVQDI 248
+ N+ R D+ R +QD+
Sbjct: 736 ALDQINKPKRPTDKPLRLPLQDV 804
>TC16374 similar to UP|Q7LZA6 (Q7LZA6) Protamine II (Fragment), partial
(63%)
Length = 1130
Score = 28.9 bits (63), Expect = 1.2
Identities = 31/124 (25%), Positives = 53/124 (42%), Gaps = 4/124 (3%)
Frame = +3
Query: 170 ISSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFVSNTTE-GDDADSESENLSEALALLAR 228
I L D L LQ+ + EK K+ + A ++ E +D DSE+E +E +
Sbjct: 486 IICLSEDTLRDHLQSKRHARSEKLYKQGRLKAMLNINGEIENDEDSETEIQTEDTEENVQ 665
Query: 229 KFNRALRKIDRRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQC---HECEGYGHI 285
K + K +++ + + KSD S + + ++ KS + V+C CEG
Sbjct: 666 KKKKKNTKGEKQNKKRLGKKKSDKSNTRKMRSTKRQAKKSRKE*EVRCLFSTICEGVFFF 845
Query: 286 RSEC 289
C
Sbjct: 846 SGYC 857
>TC9022 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (75%)
Length = 1083
Score = 27.7 bits (60), Expect = 2.6
Identities = 22/91 (24%), Positives = 37/91 (40%)
Frame = +3
Query: 211 DADSESENLSEALALLARKFNRALRKIDRRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQ 270
DA SE+E L + L+ + + P V+D+K D+ + ++D+
Sbjct: 120 DASSEAEALEQVLSAEETTLKKKPQP-QEDDAPIVEDVKDDDKDEADDDEDEDDDDEDDD 296
Query: 271 NKGVQCHECEGYGHIRSECATYLKKQKKGMV 301
+ EG RSE KK +K M+
Sbjct: 297 KEDGALGGSEGSKQSRSE-----KKSRKAML 374
>TC18180 weakly similar to GB|AAF04891.1|6175165|ATAC011437 Mutator-like
transposase {Arabidopsis thaliana;} , partial (4%)
Length = 513
Score = 27.3 bits (59), Expect = 3.4
Identities = 14/37 (37%), Positives = 18/37 (47%)
Frame = +3
Query: 253 SKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSEC 289
+K Y SQ VK E + C C+G GH +S C
Sbjct: 171 TKRYVSQNIVKRE--------LHCSRCKGLGHNKSTC 257
>TC9721 similar to UP|Q9LTA1 (Q9LTA1) Genomic DNA, chromosome 5, TAC
clone:K2I5, partial (12%)
Length = 520
Score = 27.3 bits (59), Expect = 3.4
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = -3
Query: 18 KGWKHPVKAQAEGSTSTPEEKPEDELTSAEE 48
+G V AQ EGS+S+PE P D + ++
Sbjct: 236 QGLSGTVPAQPEGSSSSPEPGPPDRILQQQQ 144
>TC15524 similar to UP|Q8RWF0 (Q8RWF0) 26S proteasome subunit-like protein,
partial (47%)
Length = 583
Score = 26.6 bits (57), Expect = 5.8
Identities = 18/66 (27%), Positives = 32/66 (48%)
Frame = -1
Query: 198 KSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRRTRPNVQDIKSDNSKSYN 257
+SIA + N+ + ESE + E L + RK + K+ R +Q ++S + S
Sbjct: 202 QSIASLENSMSEKLLELESELMPELLLVQIRKRIVPIVKLRVRAAKRIQVLQSSHGDSTT 23
Query: 258 SQRKVK 263
++K K
Sbjct: 22 EEKKKK 5
>TC11885 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (26%)
Length = 555
Score = 26.6 bits (57), Expect = 5.8
Identities = 10/16 (62%), Positives = 10/16 (62%)
Frame = +2
Query: 276 CHECEGYGHIRSECAT 291
CH C GHI SEC T
Sbjct: 401 CHNCGLPGHIASECTT 448
>TC11530
Length = 647
Score = 26.2 bits (56), Expect = 7.5
Identities = 8/26 (30%), Positives = 20/26 (76%)
Frame = -3
Query: 215 ESENLSEALALLARKFNRALRKIDRR 240
+ EN+ E + ++++K+NR L +++R+
Sbjct: 642 KKENICECIKVVSQKYNRMLHRLNRQ 565
>TC8378
Length = 694
Score = 26.2 bits (56), Expect = 7.5
Identities = 13/40 (32%), Positives = 23/40 (57%), Gaps = 6/40 (15%)
Frame = -2
Query: 269 GQNKGVQCHE------CEGYGHIRSECATYLKKQKKGMVV 302
GQ+ G + H C+G+ H RS ++ LKK++ +V+
Sbjct: 423 GQDAGKKHHMDALKPLCQGFMHKRSNSSSLLKKERYNLVM 304
>TC13053 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (9%)
Length = 450
Score = 25.8 bits (55), Expect = 9.8
Identities = 8/14 (57%), Positives = 9/14 (64%)
Frame = +3
Query: 276 CHECEGYGHIRSEC 289
CH C G GH+ EC
Sbjct: 69 CHNCGGRGHLAYEC 110
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.310 0.126 0.341
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,835,915
Number of Sequences: 28460
Number of extensions: 35171
Number of successful extensions: 199
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 196
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 199
length of query: 310
length of database: 4,897,600
effective HSP length: 90
effective length of query: 220
effective length of database: 2,336,200
effective search space: 513964000
effective search space used: 513964000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0101.12


