

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.3
(244 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
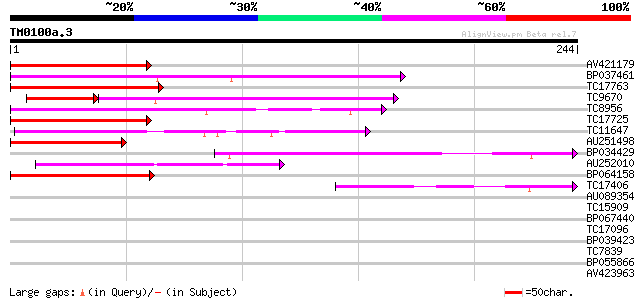
Score E
Sequences producing significant alignments: (bits) Value
AV421179 118 1e-27
BP037461 105 5e-24
TC17763 similar to UP|Q7Y1U9 (Q7Y1U9) SVP-like floral repressor,... 97 3e-21
TC9670 homologue to UP|O64958 (O64958) CUM1, partial (65%) 68 9e-20
TC8956 similar to UP|Q9ZS28 (Q9ZS28) MADs-box protein, GDEF1, pa... 87 3e-18
TC17725 homologue to UP|Q8LLR2 (Q8LLR2) MADS-box protein 2, part... 80 4e-16
TC11647 similar to UP|Q7X9H7 (Q7X9H7) MADS-box protein AGL62, pa... 79 5e-16
AU251498 69 6e-13
BP034429 68 2e-12
AU252010 67 4e-12
BP064158 65 1e-11
TC17406 similar to UP|Q7Y1U9 (Q7Y1U9) SVP-like floral repressor,... 43 6e-05
AU089354 36 0.005
TC15909 similar to UP|GCH2_ARATH (P47924) Riboflavin biosynthesi... 28 1.1
BP067440 27 2.5
TC17096 similar to PIR|C53394|C53394 ribosomal protein L12.C pre... 27 2.5
BP039423 27 2.5
TC7839 homologue to UP|Q93XW1 (Q93XW1) 14-3-3 protein, complete 27 3.2
BP055866 27 4.2
AV423963 27 4.2
>AV421179
Length = 207
Score = 118 bits (295), Expect = 1e-27
Identities = 61/61 (100%), Positives = 61/61 (100%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS
Sbjct: 6 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 185
Query: 61 S 61
S
Sbjct: 186S 188
>BP037461
Length = 551
Score = 105 bits (263), Expect = 5e-24
Identities = 66/173 (38%), Positives = 94/173 (54%), Gaps = 3/173 (1%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+++K+I+N +RQVTF+KRR GL KKA ELS LCDA++ALI+FS KL+E+ SS
Sbjct: 8 MGRGRVELKRIENKINRQVTFAKRRNGLLKKAYELSVLCDAEVALIIFSNRGKLYEFCSS 187
Query: 61 SR--GGLLPLPDANSAT*DKTTPPSPTMQFESDSND-TLRKKVEEKTHELRQLNGEDLQG 117
S L N + ++ S L+ + E R L GEDL
Sbjct: 188 SSMLKTLERYQKCNYGAPEANVSTREALELSSQQEYLKLKARYEALQRSQRNLMGEDLGP 367
Query: 118 LTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVL 170
L +L+ LE L SL + + + + ++S L+RKE L E N+ L+Q L
Sbjct: 368 LNSKELESLERQLDSSLKQIRSTRTQFMLDQLSDLQRKEHMLSEANRSLRQRL 526
>TC17763 similar to UP|Q7Y1U9 (Q7Y1U9) SVP-like floral repressor, partial
(35%)
Length = 737
Score = 96.7 bits (239), Expect = 3e-21
Identities = 46/66 (69%), Positives = 58/66 (87%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQIKKIDN ++RQVTFSKRR+GLFKKA+ELS LCDAD+AL+VFS+T KLFEY++
Sbjct: 501 MAREKIQIKKIDNATARQVTFSKRRRGLFKKAEELSVLCDADVALVVFSSTGKLFEYSNL 680
Query: 61 SRGGLL 66
S +L
Sbjct: 681 SMKEIL 698
>TC9670 homologue to UP|O64958 (O64958) CUM1, partial (65%)
Length = 1060
Score = 68.2 bits (165), Expect(2) = 9e-20
Identities = 39/130 (30%), Positives = 70/130 (53%), Gaps = 1/130 (0%)
Frame = +3
Query: 39 CDADIALIVFSATNKLFEYASSS-RGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLR 97
CDA++ALIVFS+ +L+EYA++S + + A S + + QF D LR
Sbjct: 93 CDAEVALIVFSSRGRLYEYANNSVKATIDRYKKACSDSSGAGSASEANAQFYQQEADKLR 272
Query: 98 KKVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEV 157
++ + RQ+ GE L + +L+ LE L++ ++ + K+E EI ++++E+
Sbjct: 273 VQISNLQNNNRQMMGESLGSMNAKELKNLETKLEKGISRIRSKKNELLFAEIEYMQKREI 452
Query: 158 ELIEENQKLK 167
+L NQ L+
Sbjct: 453 DLHNNNQLLR 482
Score = 44.3 bits (103), Expect(2) = 9e-20
Identities = 21/31 (67%), Positives = 25/31 (79%)
Frame = +1
Query: 8 IKKIDNISSRQVTFSKRRKGLFKKAQELSTL 38
IK+I+N ++RQVTF KRR GL KKA ELS L
Sbjct: 1 IKRIENTTNRQVTFCKRRNGLLKKAYELSVL 93
>TC8956 similar to UP|Q9ZS28 (Q9ZS28) MADs-box protein, GDEF1, partial
(58%)
Length = 1100
Score = 86.7 bits (213), Expect = 3e-18
Identities = 58/170 (34%), Positives = 95/170 (55%), Gaps = 8/170 (4%)
Frame = +2
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +I+IK I+N ++RQVT+SKRR G+FKKA ELS LCDA ++LI+FS NK+ EY S
Sbjct: 137 MGRGKIEIKLIENPTNRQVTYSKRRNGIFKKAHELSVLCDAKVSLIMFSKNNKMHEYISP 316
Query: 61 SRGGLLPLPDANSAT*DKTTPPS------PTMQFESDSNDTLRKKVEEKTHELRQLNGED 114
+ D S ++ + N+ LR+++ + E G D
Sbjct: 317 GLTTKRIIDQYQKTLGDIDLWRSHYEKMLENLKKLKEINNKLRRQIRHRLGE-----GLD 481
Query: 115 LQGLTLHQLQKLEEVLKRSLASVSRVKDEKF--MQEISTLKRKEVELIEE 162
+ L+ QL+KLEE + ++S+ ++++ KF ++ + RK+V +E+
Sbjct: 482 MDDLSFQQLRKLEEDM---VSSIGKIRERKFHVIKTRTDTCRKKVRSLEQ 622
>TC17725 homologue to UP|Q8LLR2 (Q8LLR2) MADS-box protein 2, partial (45%)
Length = 970
Score = 79.7 bits (195), Expect = 4e-16
Identities = 38/61 (62%), Positives = 49/61 (80%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R R+++K+I+N +RQVTF+KRR GL KKA ELS LCDA++ALIVFS KL+E+ SS
Sbjct: 276 MGRGRVELKRIENKINRQVTFAKRRNGLLKKAYELSVLCDAEVALIVFSTRGKLYEFCSS 455
Query: 61 S 61
S
Sbjct: 456 S 458
>TC11647 similar to UP|Q7X9H7 (Q7X9H7) MADS-box protein AGL62, partial (25%)
Length = 843
Score = 79.3 bits (194), Expect = 5e-16
Identities = 61/165 (36%), Positives = 86/165 (51%), Gaps = 12/165 (7%)
Frame = +1
Query: 3 RKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSSR 62
R++I++KK+ N S+ QVTFSKRR GLFKKA EL LCD +IAL+VFS K+F +
Sbjct: 91 RQKIEMKKMTNESNLQVTFSKRRSGLFKKASELCILCDVEIALVVFSPGEKVFSFGH--- 261
Query: 63 GGLLPLPDANSAT*DKTTPP--SPTMQF-ESDSNDTLRKKVEEKTHELRQLN-------- 111
P +A PP S TMQ+ E+ N KV E EL Q+N
Sbjct: 262 ----PSVEAVIKRYFSQAPPQTSDTMQYIEAQRN----SKVHELNVELSQINNLLDTVRN 417
Query: 112 -GEDLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRK 155
GE+L LH+ + + + + R K E+F ++ LK++
Sbjct: 418 HGEELN--RLHKAAQAQFWWACQIEEMDRPKLEQFRMALNELKKQ 546
>AU251498
Length = 342
Score = 69.3 bits (168), Expect = 6e-13
Identities = 33/50 (66%), Positives = 42/50 (84%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSA 50
M R +I+IK+I+N ++RQVTF KRR GL KKA ELS LCDA++ALIVFS+
Sbjct: 45 MGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSS 194
>BP034429
Length = 567
Score = 67.8 bits (164), Expect = 2e-12
Identities = 50/158 (31%), Positives = 81/158 (50%), Gaps = 2/158 (1%)
Frame = -1
Query: 89 ESDSN-DTLRKKVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSLASVSRVKDEKFMQ 147
E +SN L K+V +T +LR++ GED +GL L +LE+ L L V +K+++ M
Sbjct: 564 EDNSNLAELHKEVANRTEQLRRMTGEDFEGLEFDDLLELEKTLPSGLKRVIELKEKRIMD 385
Query: 148 EISTLKRKEVELIEENQKLKQVLYSLFKLLFNEYLCSIILSYGGIDSQHILMKYM*F*VP 207
EI+ +++K ++L EEN+ LKQ + L K + L S I
Sbjct: 384 EITAVQKKGIQLEEENKLLKQKMAMLCKGKSHFLLDSNI--------------------- 268
Query: 208 SLTQYGQRQSLESTIS-SSSYLLEEDGSDTSLKLGLVY 244
++ + S+ + S +S L++D S TSLKLGL +
Sbjct: 267 TMQEVASSDSMNNVCSCNSGPSLDDDSSVTSLKLGLPF 154
>AU252010
Length = 328
Score = 66.6 bits (161), Expect = 4e-12
Identities = 38/107 (35%), Positives = 64/107 (59%)
Frame = +1
Query: 12 DNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASSSRGGLLPLPDA 71
+N S+RQVT+SKR+ G+ KKA+E++ LCDA ++LI+F+A+ K+ +Y S S L+ + +
Sbjct: 1 ENSSNRQVTYSKRKNGILKKAKEITVLCDAQVSLIIFAASGKMHDYISPST-TLVDMLER 177
Query: 72 NSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGL 118
T K + + + L+K+ + ELR L G+D+ L
Sbjct: 178 YHKTSGKRLWDAKHENLNGEI-ERLKKENDGMQIELRHLKGDDINSL 315
>BP064158
Length = 394
Score = 65.1 bits (157), Expect = 1e-11
Identities = 31/62 (50%), Positives = 45/62 (72%)
Frame = +3
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R ++Q +K++N ++RQVTFSKRR GL KKA ELS LC D+ALI+FS + + ++ +
Sbjct: 165 MGRVKLQNQKVENTTNRQVTFSKRRNGLIKKAYELSVLCXFDVALIMFSPSGRACLFSGN 344
Query: 61 SR 62
R
Sbjct: 345 RR 350
>TC17406 similar to UP|Q7Y1U9 (Q7Y1U9) SVP-like floral repressor, partial
(33%)
Length = 523
Score = 42.7 bits (99), Expect = 6e-05
Identities = 36/108 (33%), Positives = 54/108 (49%), Gaps = 4/108 (3%)
Frame = +2
Query: 141 KDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFNEYLCSIILSYGGIDSQHILMK 200
K EK M EI+ L+ K L+EEN++LK+ + + S L +G +S+ ++M+
Sbjct: 8 KGEKIMNEINGLQIKGKPLMEENERLKRHVAGMI---------STGLMHGDTESELLVME 160
Query: 201 YM*F*VPSLTQYGQRQSLESTI----SSSSYLLEEDGSDTSLKLGLVY 244
+ S ES S++ LE+D SDTSLKLGL Y
Sbjct: 161 -------------EGHSSESVTNVCNSTTGPPLEDDSSDTSLKLGLPY 265
>AU089354
Length = 178
Score = 36.2 bits (82), Expect = 0.005
Identities = 15/35 (42%), Positives = 24/35 (67%)
Frame = +3
Query: 27 GLFKKAQELSTLCDADIALIVFSATNKLFEYASSS 61
G+ KKA E++ LC A ++LI F+A+ K+ +Y S
Sbjct: 3 GILKKAIEITVLCXAQVSLIXFAASGKMHDYIRPS 107
>TC15909 similar to UP|GCH2_ARATH (P47924) Riboflavin biosynthesis protein
ribA [Includes: GTP cyclohydrolase II ;
3,4-dihydroxy-2-butanone 4-phosphate synthase (DHBP
synthase)] , partial (21%)
Length = 825
Score = 28.5 bits (62), Expect = 1.1
Identities = 14/37 (37%), Positives = 19/37 (50%)
Frame = +2
Query: 60 SSRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTL 96
S+ GGL+ PD N+ K P ++ FE N TL
Sbjct: 320 SAGGGLVSHPDTNNVVIAKNLPGDESVGFEDQPNGTL 430
>BP067440
Length = 413
Score = 27.3 bits (59), Expect = 2.5
Identities = 13/45 (28%), Positives = 22/45 (48%)
Frame = +1
Query: 97 RKKVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSLASVSRVK 141
RKKV +H+L+ + +G HQ Q V + + +S+ K
Sbjct: 247 RKKVSNPSHKLQSPYQREAEGRDQHQYQTFNRVSEEQIHIISQCK 381
>TC17096 similar to PIR|C53394|C53394 ribosomal protein L12.C precursor,
chloroplast - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (74%)
Length = 677
Score = 27.3 bits (59), Expect = 2.5
Identities = 15/53 (28%), Positives = 25/53 (46%)
Frame = +2
Query: 82 PSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSL 134
P T+ F + T+R + E + G+D+ LTL Q + L + L+ L
Sbjct: 71 PKTTLHFPTPLRRTIRLRPISAVSEKVEKLGDDISSLTLEQAKTLVDYLQDKL 229
>BP039423
Length = 566
Score = 27.3 bits (59), Expect = 2.5
Identities = 18/76 (23%), Positives = 39/76 (50%), Gaps = 7/76 (9%)
Frame = +1
Query: 117 GLTLHQLQKLEEVLKRS------LASVSRVKDEKFMQEISTLKRK-EVELIEENQKLKQV 169
GLTL +++ L S + ++ + E+ ++ + RK E+ELI++ K++
Sbjct: 274 GLTLKNSAEIKGTLGESDFLIDYIGLLAELPSERQYEKSGNITRKIELELIDDKGKVRCA 453
Query: 170 LYSLFKLLFNEYLCSI 185
L+ + + EYL ++
Sbjct: 454 LFGNYVDIVKEYLANV 501
>TC7839 homologue to UP|Q93XW1 (Q93XW1) 14-3-3 protein, complete
Length = 1284
Score = 26.9 bits (58), Expect = 3.2
Identities = 13/38 (34%), Positives = 21/38 (55%)
Frame = -2
Query: 163 NQKLKQVLYSLFKLLFNEYLCSIILSYGGIDSQHILMK 200
+ K+ +L+ F F ++CSIIL GG + + I K
Sbjct: 902 SSKISSLLFIGFCWCFFNFICSIILHVGGPEGEVITKK 789
>BP055866
Length = 541
Score = 26.6 bits (57), Expect = 4.2
Identities = 23/86 (26%), Positives = 36/86 (41%)
Frame = +3
Query: 82 PSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTLHQLQKLEEVLKRSLASVSRVK 141
PS F + + T V+ E + + ++ + L L+KL +R L +
Sbjct: 48 PSSNNGFTNGTTTTNTNTVDLSLLEAIEKSQHTIEAVDLRTLKKLVLAFERRLKDNIEAR 227
Query: 142 DEKFMQEISTLKRKEVELIEENQKLK 167
K+ + EVEL EE QKLK
Sbjct: 228 -LKYPNQPDRFADSEVELHEELQKLK 302
>AV423963
Length = 381
Score = 26.6 bits (57), Expect = 4.2
Identities = 25/101 (24%), Positives = 38/101 (36%), Gaps = 9/101 (8%)
Frame = +1
Query: 67 PLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGE---------DLQG 117
P P A + PPSP+ + S K H + L G L G
Sbjct: 40 PSPSAPATPPIALKPPSPSPRRNDSSPSPQTNSTSSKNHHVPILAGSAGVAAFLLISLTG 219
Query: 118 LTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVE 158
+ L + K+ V K + +S + F+ + LKR E+E
Sbjct: 220 IYLCKTNKVTTV-KPWVTGLSGQLQKAFVTGVPKLKRSELE 339
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.136 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,191,321
Number of Sequences: 28460
Number of extensions: 36352
Number of successful extensions: 246
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 241
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 244
length of query: 244
length of database: 4,897,600
effective HSP length: 88
effective length of query: 156
effective length of database: 2,393,120
effective search space: 373326720
effective search space used: 373326720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0100a.3


