

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.12
(405 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
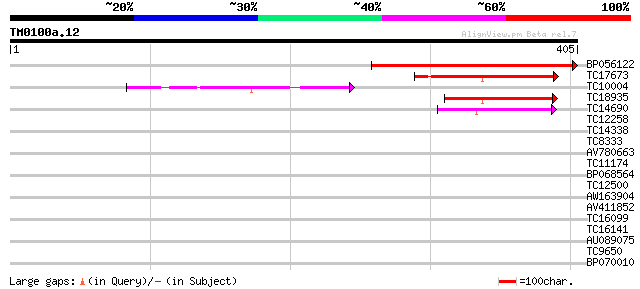
Score E
Sequences producing significant alignments: (bits) Value
BP056122 303 3e-83
TC17673 homologue to PIR|S29560|S29560 fructose-bisphosphatase ... 121 2e-28
TC10004 similar to UP|F16Q_SOLTU (P46276) Fructose-1,6-bisphosph... 98 3e-21
TC18935 homologue to UP|Q9XGG5 (Q9XGG5) Fructose-1,6-bisphosphat... 91 3e-19
TC14690 similar to UP|Q84JG8 (Q84JG8) Sedoheptulose-1,7-bisphosp... 47 7e-06
TC12258 weakly similar to UP|O50458 (O50458) PGRS_FAMILY protein... 30 0.72
TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%) 29 1.2
TC8333 weakly similar to UP|Q96GG9 (Q96GG9) RP42 homolog (Leucin... 29 1.2
AV780663 29 1.6
TC11174 weakly similar to UP|P93837 (P93837) Amp-binding protein... 28 2.1
BP068564 28 2.1
TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein... 28 2.1
AW163904 28 2.1
AV411852 28 2.7
TC16099 similar to UP|Q9LT41 (Q9LT41) RCD1, partial (91%) 28 2.7
TC16141 weakly similar to GB|AAC77862.2|20197419|AC005623 Argona... 28 2.7
AU089075 28 2.7
TC9650 similar to UP|O49485 (O49485) Phosphoglycerate dehydrogen... 28 3.6
BP070010 27 4.7
>BP056122
Length = 537
Score = 303 bits (776), Expect = 3e-83
Identities = 147/147 (100%), Positives = 147/147 (100%)
Frame = -3
Query: 259 DFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYICSLVA 318
DFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYICSLVA
Sbjct: 535 DFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYICSLVA 356
Query: 319 DLHRTLMYGGVAMNPRDHLRLVYEANPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPL 378
DLHRTLMYGGVAMNPRDHLRLVYEANPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPL
Sbjct: 355 DLHRTLMYGGVAMNPRDHLRLVYEANPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPL 176
Query: 379 FLGSLEDMEELESFGDVQQKVNPGYEV 405
FLGSLEDMEELESFGDVQQKVNPGYEV
Sbjct: 175 FLGSLEDMEELESFGDVQQKVNPGYEV 95
>TC17673 homologue to PIR|S29560|S29560 fructose-bisphosphatase - garden
pea (fragment) {Pisum sativum;} ,
partial (29%)
Length = 573
Score = 121 bits (303), Expect = 2e-28
Identities = 59/109 (54%), Positives = 82/109 (75%), Gaps = 6/109 (5%)
Frame = +3
Query: 290 LRQYIDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNPRDH------LRLVYEA 343
L++YID +++ G K YSARYI SLV D HRT++YGG+ PRD LRL+YE
Sbjct: 6 LKKYIDNLKE-PGPSGKPYSARYIGSLVGDFHRTILYGGIYGYPRDKKSKNGKLRLLYEC 182
Query: 344 NPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEELESF 392
P+SF+VEQAGG+GSDG RIL +QP ++HQR+PL++GS+E++E++E F
Sbjct: 183 APMSFIVEQAGGKGSDGHQRILDIQPTEIHQRVPLYIGSVEEVEKVEKF 329
>TC10004 similar to UP|F16Q_SOLTU (P46276) Fructose-1,6-bisphosphatase,
cytosolic (D-fructose-1,6-bisphosphate
1-phosphohydrolase) (FBPase) (CY-F1) , partial (53%)
Length = 553
Score = 97.8 bits (242), Expect = 3e-21
Identities = 59/165 (35%), Positives = 88/165 (52%), Gaps = 2/165 (1%)
Frame = +1
Query: 84 KDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEI 143
+ D +LL HI CK + + V+ G +G G G + K LD++SNE+
Sbjct: 97 RGDFSILLSHIVLGCKFVCSAVSKA-----GLAKLIGLAGETNVQGEEQKK-LDVLSNEV 258
Query: 144 ILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIFGIYN 201
+ +L SG+ +++ SEE++ T++ G Y VV DPLDGS NID + GTIFGIY
Sbjct: 259 FIKALVSSGRTSILVSEEDEEATFVEASKRGKYCVVFDPLDGSSNIDCGVSIGTIFGIY- 435
Query: 202 RLEELDNLPTEEKAQLNSLQSGSKLIAAGYVLYSSATILCVSFGS 246
+ +E + LQ G ++AAGY +Y S+ L +S GS
Sbjct: 436 ------MVKDQEPTIGDVLQPGKNMLAAGYCMYGSSCTLVISTGS 552
>TC18935 homologue to UP|Q9XGG5 (Q9XGG5) Fructose-1,6-bisphosphatase ,
partial (29%)
Length = 488
Score = 91.3 bits (225), Expect = 3e-19
Identities = 45/87 (51%), Positives = 62/87 (70%), Gaps = 6/87 (6%)
Frame = +1
Query: 311 RYICSLVADLHRTLMYGGVAMNPRDH------LRLVYEANPLSFLVEQAGGRGSDGKNRI 364
RYI S+VAD+HRTL+YGG + P D LR++YE P+SFL+EQAGG+ G+ R
Sbjct: 1 RYIGSMVADVHRTLIYGGSFLYPADKKSPNGKLRVLYEVFPMSFLMEQAGGQAFTGQQRA 180
Query: 365 LSLQPVKLHQRLPLFLGSLEDMEELES 391
L P KLH+R P+FLGS +D+EE+++
Sbjct: 181 RDLVPTKLHERSPIFLGSYDDIEEIKA 261
>TC14690 similar to UP|Q84JG8 (Q84JG8) Sedoheptulose-1,7-bisphosphatase
precursor, partial (33%)
Length = 597
Score = 46.6 bits (109), Expect = 7e-06
Identities = 28/91 (30%), Positives = 48/91 (51%), Gaps = 6/91 (6%)
Frame = +1
Query: 306 KKYSARYICSLVADLHRTLMYG-GVAMN-----PRDHLRLVYEANPLSFLVEQAGGRGSD 359
+KY+ RY +V D+++ ++ G+ N + LRL++E PL FL+E+AGG SD
Sbjct: 61 EKYTLRYTGGMVPDVNQIIVKEKGIFTNVSSPSAKAKLRLLFEVAPLGFLIEKAGGYSSD 240
Query: 360 GKNRILSLQPVKLHQRLPLFLGSLEDMEELE 390
G +L + + R + GS ++ E
Sbjct: 241 GHKSVLDIVIDNIDDRTQVAYGSKNEIIRFE 333
>TC12258 weakly similar to UP|O50458 (O50458) PGRS_FAMILY protein (PE_PGRS
family protein), partial (6%)
Length = 471
Score = 30.0 bits (66), Expect = 0.72
Identities = 21/52 (40%), Positives = 27/52 (51%), Gaps = 4/52 (7%)
Frame = +3
Query: 21 TFQSKTPLTTP----YCTPRLASISGSAKMRPLRALGGSSSSAGGDGDDGDG 68
TF S +PL++P +C PR +S SA A GG+ S GDG G G
Sbjct: 246 TFTS-SPLSSPSLRTFCAPRCC-VSSSAASFASSAGGGNGGSGVGDGGGGGG 395
Score = 26.9 bits (58), Expect = 6.1
Identities = 12/26 (46%), Positives = 13/26 (49%)
Frame = +3
Query: 114 GKQTGVGAGGGGGSSGRDAPKPLDIV 139
G G G GGGGGS G L +V
Sbjct: 360 GSGVGDGGGGGGGSGGESGDANLKLV 437
>TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%)
Length = 1751
Score = 29.3 bits (64), Expect = 1.2
Identities = 14/28 (50%), Positives = 16/28 (57%)
Frame = -2
Query: 53 GGSSSSAGGDGDDGDGVVTLIEYLGKEG 80
GG +AGGDG G+G V E G EG
Sbjct: 211 GGEEVAAGGDGYGGEGTVAAEEREGGEG 128
>TC8333 weakly similar to UP|Q96GG9 (Q96GG9) RP42 homolog (Leucine zipper
protein), partial (36%)
Length = 1390
Score = 29.3 bits (64), Expect = 1.2
Identities = 13/46 (28%), Positives = 27/46 (58%)
Frame = -1
Query: 184 SRNIDASIPTGTIFGIYNRLEELDNLPTEEKAQLNSLQSGSKLIAA 229
SRN+ +S + ++GI++R +++P + K +NS S ++ A
Sbjct: 652 SRNLYSSFSSERMYGIFSRNFSNESIPRDCKPSMNSFLENSHMVPA 515
>AV780663
Length = 568
Score = 28.9 bits (63), Expect = 1.6
Identities = 24/83 (28%), Positives = 36/83 (42%), Gaps = 8/83 (9%)
Frame = -2
Query: 58 SAGGDGDDGDGVVTLIEYLGKEGIDLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQT 117
++G G DGD + E +G G + D L + + + A++ C + K
Sbjct: 330 NSGNLGSDGDTALAF-EGIGIHGALVGDVGAALAEDAVH--EGGLAVIDMSDHCHVAKAV 160
Query: 118 GV--------GAGGGGGSSGRDA 132
GV G GGGGGS G+ A
Sbjct: 159 GVEGSSGGGGGRGGGGGSGGKGA 91
>TC11174 weakly similar to UP|P93837 (P93837) Amp-binding protein, partial
(15%)
Length = 689
Score = 28.5 bits (62), Expect = 2.1
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +3
Query: 111 CSIGKQTGVGAGGGGGSS 128
CS+ GVG GGGGG++
Sbjct: 18 CSVSPSQGVGGGGGGGAA 71
>BP068564
Length = 551
Score = 28.5 bits (62), Expect = 2.1
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = -3
Query: 111 CSIGKQTGVGAGGGGGSS 128
CS+ GVG GGGGG++
Sbjct: 489 CSVSPSEGVGGGGGGGAA 436
>TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein {Danio
rerio;} , partial (5%)
Length = 678
Score = 28.5 bits (62), Expect = 2.1
Identities = 26/93 (27%), Positives = 36/93 (37%), Gaps = 1/93 (1%)
Frame = -2
Query: 53 GGSSSSAGGDGDDGDGVVTLIEYLGKEGIDLKDDLVVLLDHIQYACKRIAAIVASPFSCS 112
GG GGDG + +G + D++ L AI +
Sbjct: 335 GGGGGGCGGDGRE-------------QGKEASPDIIAL-----------PAITEAGGGVV 228
Query: 113 IGKQTGVGAGGGG-GSSGRDAPKPLDIVSNEII 144
G+ G G GGGG G GR+ P D+ + EII
Sbjct: 227 GGRDGGGGCGGGGEGEQGRET-SPTDLETFEII 132
>AW163904
Length = 416
Score = 28.5 bits (62), Expect = 2.1
Identities = 13/38 (34%), Positives = 22/38 (57%)
Frame = +2
Query: 120 GAGGGGGSSGRDAPKPLDIVSNEIILSSLKKSGKVAVM 157
G GGGGG S ++AP P + + + I +K +A++
Sbjct: 62 GGGGGGGDSSKEAPPPPPLSAADRIEHQRRKVKLLAML 175
>AV411852
Length = 430
Score = 28.1 bits (61), Expect = 2.7
Identities = 15/45 (33%), Positives = 24/45 (53%), Gaps = 2/45 (4%)
Frame = -2
Query: 16 HPKFQTFQSKTPLTT--PYCTPRLASISGSAKMRPLRALGGSSSS 58
HP +F S T ++ P+C P +++S S+ P A SSS+
Sbjct: 423 HPPSSSFSSATSPSSNSPFCKPSASTMSSSSPQSPSTASYSSSSA 289
>TC16099 similar to UP|Q9LT41 (Q9LT41) RCD1, partial (91%)
Length = 1182
Score = 28.1 bits (61), Expect = 2.7
Identities = 17/51 (33%), Positives = 26/51 (50%), Gaps = 4/51 (7%)
Frame = -1
Query: 89 VLLDHIQYACKRIAAIVASPFSCS----IGKQTGVGAGGGGGSSGRDAPKP 135
+ L+ + A R + +++S SCS + + G GGGGG G AP P
Sbjct: 240 LFLESSRRAFSRRSGLLSSRTSCSAPAILLDEEGPP*GGGGGGGGAPAPPP 88
>TC16141 weakly similar to GB|AAC77862.2|20197419|AC005623 Argonaute
(AGO1)-like protein {Arabidopsis thaliana;} , partial
(14%)
Length = 598
Score = 28.1 bits (61), Expect = 2.7
Identities = 14/34 (41%), Positives = 19/34 (55%)
Frame = -1
Query: 117 TGVGAGGGGGSSGRDAPKPLDIVSNEIILSSLKK 150
T G GGGG +G+ + PL SNE I +K+
Sbjct: 136 TSEGRTGGGGGNGKLSSFPLPSGSNESIFLEMKR 35
>AU089075
Length = 362
Score = 28.1 bits (61), Expect = 2.7
Identities = 17/47 (36%), Positives = 22/47 (46%)
Frame = -3
Query: 1 MHSAAASPFYQLVPFHPKFQTFQSKTPLTTPYCTPRLASISGSAKMR 47
+H AASP + FHPK + + T P C L S S +A R
Sbjct: 132 IHHQAASP----IGFHPKRRLYVGGTGFQPPSCAAMLLSFSYAAWCR 4
>TC9650 similar to UP|O49485 (O49485) Phosphoglycerate dehydrogenase-like
protein , partial (47%)
Length = 1041
Score = 27.7 bits (60), Expect = 3.6
Identities = 18/51 (35%), Positives = 25/51 (48%)
Frame = -1
Query: 180 PLDGSRNIDASIPTGTIFGIYNRLEELDNLPTEEKAQLNSLQSGSKLIAAG 230
P G NI AS+P F +NR L N+P + N++ SGS + G
Sbjct: 609 PNHGHSNILASLP----FPSFNRGISLSNVPGHGGEEGNTVLSGSNSVGGG 469
>BP070010
Length = 519
Score = 27.3 bits (59), Expect = 4.7
Identities = 24/77 (31%), Positives = 35/77 (45%), Gaps = 6/77 (7%)
Frame = -1
Query: 118 GVGAGGGGGSSG----RDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWIIDDGP 173
G AGGGGG G D+ L IV N+ ++++ G V+ ++ D +DG
Sbjct: 501 GGTAGGGGGGEGDDWDSDSEDDLQIVLNDDNRMAMERGG---VVDDDDED------EDGG 349
Query: 174 YVVVT--DPLDGSRNID 188
V+V DP G D
Sbjct: 348 LVIVAGGDPSQGLEEQD 298
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.137 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,363,101
Number of Sequences: 28460
Number of extensions: 84519
Number of successful extensions: 748
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 707
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 736
length of query: 405
length of database: 4,897,600
effective HSP length: 92
effective length of query: 313
effective length of database: 2,279,280
effective search space: 713414640
effective search space used: 713414640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0100a.12


