

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.11
(245 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
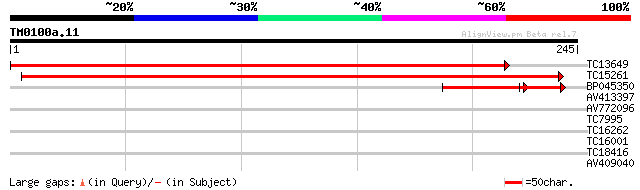
TC13649 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, ... 443 e-125
TC15261 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, ... 322 3e-89
BP045350 36 1e-04
AV413397 27 3.2
AV772096 27 4.2
TC7995 similar to PIR|S63686|S63686 sterol 24-C-methyltransferas... 26 7.2
TC16262 similar to UP|Q9FZ10 (Q9FZ10) Serine threonine kinase ho... 26 7.2
TC16001 25 9.4
TC18416 25 9.4
AV409040 25 9.4
>TC13649 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, partial
(68%)
Length = 690
Score = 443 bits (1139), Expect = e-125
Identities = 216/216 (100%), Positives = 216/216 (100%)
Frame = +2
Query: 1 MVSQIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMI 60
MVSQIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMI
Sbjct: 41 MVSQIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMI 220
Query: 61 NESARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVT 120
NESARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVT
Sbjct: 221 NESARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVT 400
Query: 121 LRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGF 180
LRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGF
Sbjct: 401 LRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGF 580
Query: 181 LKPLENVVVYSHGCATFDVPLEVARNTKGALAHPQE 216
LKPLENVVVYSHGCATFDVPLEVARNTKGALAHPQE
Sbjct: 581 LKPLENVVVYSHGCATFDVPLEVARNTKGALAHPQE 688
>TC15261 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, partial
(75%)
Length = 1031
Score = 322 bits (826), Expect = 3e-89
Identities = 145/234 (61%), Positives = 188/234 (79%)
Frame = +1
Query: 6 IELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINESAR 65
++L+K EIP++Q+ + L+ D GLVLVD++NGFCTVGAGN+AP E N +I++M+ ES
Sbjct: 49 LQLVKEEIPVKQQPLRLSGDIKTGLVLVDVVNGFCTVGAGNMAPTEPNEKITKMVEESVS 228
Query: 66 LARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLRRKE 125
L++ F EKN P+ AFLDSHHP+ PE PYP HC+ G+DE LVP L WLENE N TLRRKE
Sbjct: 229 LSKKFAEKNWPIFAFLDSHHPDIPEPPYPSHCLIGSDEGKLVPELLWLENEPNATLRRKE 408
Query: 126 CFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKPLE 185
C DG++GS E+DGSNVF DWVKKN+I+ +LV GICTD+CVLDFVCS +SA+NR FL PLE
Sbjct: 409 CIDGFLGSYEKDGSNVFADWVKKNQIKQILVAGICTDVCVLDFVCSVLSARNRQFLPPLE 588
Query: 186 NVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVLF 239
NV+V + C+T+DVPL VA+ K ++HPQE +HH+GLY+A RGA+IA+EV+F
Sbjct: 589 NVIVSTEACSTYDVPLHVAKTNKDWVSHPQELMHHIGLYIACGRGAEIASEVIF 750
>BP045350
Length = 380
Score = 35.8 bits (81), Expect(2) = 1e-04
Identities = 14/37 (37%), Positives = 23/37 (61%)
Frame = -1
Query: 188 VVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLY 224
+V C+T+DVPL VA+ + ++ P +HH+G Y
Sbjct: 380 IVSPEACSTYDVPLHVAQTYQDWVSPPPGLMHHLGPY 270
Score = 25.0 bits (53), Expect(2) = 1e-04
Identities = 11/20 (55%), Positives = 16/20 (80%)
Frame = -2
Query: 221 MGLYMAKERGAKIANEVLFG 240
+ L +A RGA+IA+EV+FG
Sbjct: 280 LALTLACGRGAQIASEVIFG 221
>AV413397
Length = 382
Score = 26.9 bits (58), Expect = 3.2
Identities = 11/32 (34%), Positives = 16/32 (49%)
Frame = -1
Query: 81 LDSHHPNKPEDPYPPHCIAGTDESNLVPALRW 112
L+ P P+ YP HC G +E++ RW
Sbjct: 187 LEGQSPESPKHTYPVHC--GPEENHSDQGFRW 98
>AV772096
Length = 431
Score = 26.6 bits (57), Expect = 4.2
Identities = 14/40 (35%), Positives = 22/40 (55%), Gaps = 1/40 (2%)
Frame = -3
Query: 159 ICTDICVLDFV-CSTMSAKNRGFLKPLENVVVYSHGCATF 197
+C+ + DF C T S K+ G+L P E+ ++Y C F
Sbjct: 168 LCSFFILSDFTPCHTHSYKSFGYLGPNEDPLLYILTCRMF 49
>TC7995 similar to PIR|S63686|S63686 sterol 24-C-methyltransferase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (98%)
Length = 1524
Score = 25.8 bits (55), Expect = 7.2
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = -2
Query: 82 DSHHPNKPEDPYPPHCIAGTDES 104
+SH P + E+P PP C A +S
Sbjct: 689 NSHTPPRRENPNPPSCCAAWPDS 621
>TC16262 similar to UP|Q9FZ10 (Q9FZ10) Serine threonine kinase homolog
COK-4, partial (6%)
Length = 623
Score = 25.8 bits (55), Expect = 7.2
Identities = 15/48 (31%), Positives = 21/48 (43%)
Frame = +3
Query: 174 SAKNRGFLKPLENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHM 221
+A RGF LE VV SHGC + A + F++H+
Sbjct: 333 TATTRGFNFSLEYVVFMSHGCMPCRIYFFGASQLATSTTA*SSFMYHL 476
>TC16001
Length = 706
Score = 25.4 bits (54), Expect = 9.4
Identities = 10/19 (52%), Positives = 13/19 (67%)
Frame = +1
Query: 74 NLPVMAFLDSHHPNKPEDP 92
NL V+AFLD+H N + P
Sbjct: 541 NLAVLAFLDTHRENYSDPP 597
>TC18416
Length = 426
Score = 25.4 bits (54), Expect = 9.4
Identities = 11/26 (42%), Positives = 13/26 (49%)
Frame = +2
Query: 87 NKPEDPYPPHCIAGTDESNLVPALRW 112
N P P IAG + S L AL+W
Sbjct: 182 NNPSSPLTSIAIAGRNPSKLTQALQW 259
>AV409040
Length = 348
Score = 25.4 bits (54), Expect = 9.4
Identities = 10/22 (45%), Positives = 11/22 (49%)
Frame = +2
Query: 75 LPVMAFLDSHHPNKPEDPYPPH 96
LP SH P+ P P PPH
Sbjct: 59 LPPKCSAHSHSPSLPPPPLPPH 124
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,557,383
Number of Sequences: 28460
Number of extensions: 65913
Number of successful extensions: 354
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 353
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 354
length of query: 245
length of database: 4,897,600
effective HSP length: 88
effective length of query: 157
effective length of database: 2,393,120
effective search space: 375719840
effective search space used: 375719840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0100a.11


