

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0096b.3
(223 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
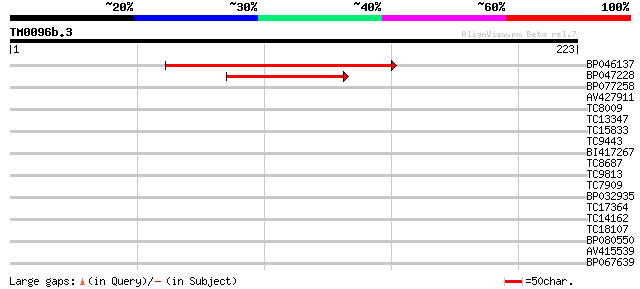
Score E
Sequences producing significant alignments: (bits) Value
BP046137 167 2e-42
BP047228 45 8e-06
BP077258 29 0.74
AV427911 28 0.97
TC8009 homologue to UP|Q9LKJ2 (Q9LKJ2) Cytosolic phosphoglycerat... 27 2.8
TC13347 similar to UP|Q8LGB4 (Q8LGB4) Protein kinase-like protei... 27 2.8
TC15833 similar to UP|Q9FJ12 (Q9FJ12) Genomic DNA, chromosome 5,... 27 3.7
TC9443 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting p... 27 3.7
BI417267 26 4.8
TC8687 similar to UP|Q9SXL2 (Q9SXL2) Serine decarboxylase, parti... 26 6.3
TC9813 similar to UP|Q9C550 (Q9C550) 2-isopropylmalate synthase ... 26 6.3
TC7909 similar to UP|CB4C_ARATH (Q9S7W1) Chlorophyll a-b binding... 26 6.3
BP032935 26 6.3
TC17364 homologue to UP|Q9VVG2 (Q9VVG2) CG13731-PA, partial (3%) 26 6.3
TC14162 similar to UP|OLP1_LYCES (Q41350) Osmotin-like protein p... 26 6.3
TC18107 weakly similar to UP|Q8LFG7 (Q8LFG7) NAM-like protein, p... 26 6.3
BP080550 25 8.2
AV415539 25 8.2
BP067639 25 8.2
>BP046137
Length = 497
Score = 167 bits (422), Expect = 2e-42
Identities = 86/91 (94%), Positives = 87/91 (95%)
Frame = -3
Query: 62 MQDVGALVELDPGTPVPEVLALDSSLILPDALKFVSVMTKAIRDLIIAKQVGVVFDGTDE 121
MQDVGALVELDPGTPVPEVLALDSSLILPDALKFVSVMTKAIR LIIAKQVGVVFDGTDE
Sbjct: 453 MQDVGALVELDPGTPVPEVLALDSSLILPDALKFVSVMTKAIRVLIIAKQVGVVFDGTDE 274
Query: 122 EAIESYAIHLVANCHGGVYLVFCIGFIIKTF 152
EAIESYAIHLVANCHGGVYLVFCIG +I F
Sbjct: 273 EAIESYAIHLVANCHGGVYLVFCIG*LIIWF 181
>BP047228
Length = 467
Score = 45.4 bits (106), Expect = 8e-06
Identities = 25/48 (52%), Positives = 35/48 (72%)
Frame = -1
Query: 86 SLILPDALKFVSVMTKAIRDLIIAKQVGVVFDGTDEEAIESYAIHLVA 133
SLILP+A V +T+A R +IA+ VGVV G++E++I+S A HLVA
Sbjct: 428 SLILPEARGSVRALTRAQRIFLIAQPVGVVPIGSEEDSIQSLADHLVA 285
>BP077258
Length = 442
Score = 28.9 bits (63), Expect = 0.74
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = +3
Query: 154 QPPPPNDYSCKEPINPHPIHILEFLGS 180
+PP N +CK P P PI L+ LG+
Sbjct: 159 KPPKANRQTCKNPNEPKPIKTLKELGT 239
>AV427911
Length = 348
Score = 28.5 bits (62), Expect = 0.97
Identities = 10/24 (41%), Positives = 14/24 (57%)
Frame = +1
Query: 151 TFYQPPPPNDYSCKEPINPHPIHI 174
T++ PPPP K P P P+H+
Sbjct: 268 TYHSPPPPPKKPYKYPSPPPPVHV 339
Score = 25.4 bits (54), Expect = 8.2
Identities = 11/26 (42%), Positives = 15/26 (57%), Gaps = 1/26 (3%)
Frame = +1
Query: 150 KTFYQ-PPPPNDYSCKEPINPHPIHI 174
K +Y+ PPPP K P P P+H+
Sbjct: 178 KYYYKSPPPPPKKPYKYPSPPPPVHV 255
>TC8009 homologue to UP|Q9LKJ2 (Q9LKJ2) Cytosolic phosphoglycerate kinase ,
complete
Length = 1697
Score = 26.9 bits (58), Expect = 2.8
Identities = 16/49 (32%), Positives = 27/49 (54%)
Frame = +2
Query: 38 LLSGNTNGLSIQTRGMLSPHIHHRMQDVGALVELDPGTPVPEVLALDSS 86
++ G + +++ G L+ + H GA +EL G P+P VLALD +
Sbjct: 1238 IIGGGDSVAAVEKVG-LADKMSHISTGGGASLELLEGKPLPGVLALDDA 1381
>TC13347 similar to UP|Q8LGB4 (Q8LGB4) Protein kinase-like protein, partial
(6%)
Length = 531
Score = 26.9 bits (58), Expect = 2.8
Identities = 12/38 (31%), Positives = 21/38 (54%)
Frame = -1
Query: 183 IHLLMCYSLLLSNSSHIVMFSKLTSLEKGKSEVIGESQ 220
+HL + L +SN SH++ + + + K K + G SQ
Sbjct: 390 LHLSFYFYLQVSNPSHLIFSTLVRIMSKNKKVISGFSQ 277
>TC15833 similar to UP|Q9FJ12 (Q9FJ12) Genomic DNA, chromosome 5, TAC
clone:K21P3 (Extensin like protein), partial (25%)
Length = 542
Score = 26.6 bits (57), Expect = 3.7
Identities = 9/18 (50%), Positives = 11/18 (61%)
Frame = +1
Query: 152 FYQPPPPNDYSCKEPINP 169
+Y PPPP+ YS P P
Sbjct: 469 YYSPPPPSQYSYSSPPPP 522
>TC9443 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), partial (41%)
Length = 1025
Score = 26.6 bits (57), Expect = 3.7
Identities = 9/25 (36%), Positives = 12/25 (48%)
Frame = -2
Query: 155 PPPPNDYSCKEPINPHPIHILEFLG 179
P P + C+ P+ P H L F G
Sbjct: 127 PEPLPQFCCRHPVEQPPFHFLHFQG 53
>BI417267
Length = 628
Score = 26.2 bits (56), Expect = 4.8
Identities = 12/36 (33%), Positives = 21/36 (58%), Gaps = 1/36 (2%)
Frame = -2
Query: 8 GHEEPILEVL-VRMARVVSLVVVDDVVYHLQLLSGN 42
GH +L +R+ R++S+ + D+ +HL SGN
Sbjct: 357 GHTTRLLTACDIRVIRILSICIRHDICHHLYWSSGN 250
>TC8687 similar to UP|Q9SXL2 (Q9SXL2) Serine decarboxylase, partial (80%)
Length = 1644
Score = 25.8 bits (55), Expect = 6.3
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = -2
Query: 144 CIGFIIKTFYQPPPPNDYSCK 164
CI +IKT P N+YSCK
Sbjct: 1610 CINILIKTHCIRTP*NNYSCK 1548
>TC9813 similar to UP|Q9C550 (Q9C550) 2-isopropylmalate synthase
(At1g74040/F2P9_9) , partial (12%)
Length = 697
Score = 25.8 bits (55), Expect = 6.3
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = +2
Query: 165 EPINPHPIHILEFLGSIIIHLLM 187
+P+ P P+ L F G IIHLLM
Sbjct: 35 KPLMPLPLQEL*FAGKQIIHLLM 103
>TC7909 similar to UP|CB4C_ARATH (Q9S7W1) Chlorophyll a-b binding protein
CP29.3, chloroplast precursor (LHCII protein 4.3)
(LHCB4.3), partial (80%)
Length = 1155
Score = 25.8 bits (55), Expect = 6.3
Identities = 12/35 (34%), Positives = 19/35 (54%), Gaps = 6/35 (17%)
Frame = -2
Query: 151 TFYQPPPPNDYSCKEPINPHPIHI------LEFLG 179
T + PP ND + EP+NP + + L+F+G
Sbjct: 575 TKHGPPSMNDLAFTEPLNPENLRVRLKRRRLDFIG 471
>BP032935
Length = 534
Score = 25.8 bits (55), Expect = 6.3
Identities = 9/16 (56%), Positives = 14/16 (87%)
Frame = -1
Query: 173 HILEFLGSIIIHLLMC 188
+I+EFL S+I+H+L C
Sbjct: 258 NIVEFLSSLIVHILEC 211
>TC17364 homologue to UP|Q9VVG2 (Q9VVG2) CG13731-PA, partial (3%)
Length = 446
Score = 25.8 bits (55), Expect = 6.3
Identities = 13/35 (37%), Positives = 22/35 (62%)
Frame = -3
Query: 124 IESYAIHLVANCHGGVYLVFCIGFIIKTFYQPPPP 158
+++ A+ VA G +YLV G++ KTF++ P P
Sbjct: 288 VKAAAMWGVAAGAGALYLVQPWGWLRKTFFEKPEP 184
>TC14162 similar to UP|OLP1_LYCES (Q41350) Osmotin-like protein precursor,
partial (87%)
Length = 1232
Score = 25.8 bits (55), Expect = 6.3
Identities = 11/21 (52%), Positives = 12/21 (56%)
Frame = +3
Query: 40 SGNTNGLSIQTRGMLSPHIHH 60
S N NG S T +L PH HH
Sbjct: 18 SNNANGSSFFTSHILLPHSHH 80
>TC18107 weakly similar to UP|Q8LFG7 (Q8LFG7) NAM-like protein, partial
(10%)
Length = 505
Score = 25.8 bits (55), Expect = 6.3
Identities = 9/29 (31%), Positives = 15/29 (51%)
Frame = +1
Query: 144 CIGFIIKTFYQPPPPNDYSCKEPINPHPI 172
CIGF+ ++ PP N+ P+ P+
Sbjct: 37 CIGFVCESVNYPPEGNENQSSSPVEDIPM 123
>BP080550
Length = 379
Score = 25.4 bits (54), Expect = 8.2
Identities = 9/18 (50%), Positives = 10/18 (55%)
Frame = -1
Query: 154 QPPPPNDYSCKEPINPHP 171
QPPPPN+Y P P
Sbjct: 244 QPPPPNNYMQSPPATKQP 191
>AV415539
Length = 419
Score = 25.4 bits (54), Expect = 8.2
Identities = 9/20 (45%), Positives = 12/20 (60%)
Frame = +3
Query: 155 PPPPNDYSCKEPINPHPIHI 174
PPPP + P NP P+H+
Sbjct: 108 PPPPPHPTRPPPGNPQPLHV 167
>BP067639
Length = 448
Score = 25.4 bits (54), Expect = 8.2
Identities = 9/14 (64%), Positives = 14/14 (99%)
Frame = -3
Query: 172 IHILEFLGSIIIHL 185
I++L+FLGS++IHL
Sbjct: 146 IYLLKFLGSVLIHL 105
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.142 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,411,405
Number of Sequences: 28460
Number of extensions: 72266
Number of successful extensions: 494
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 478
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 494
length of query: 223
length of database: 4,897,600
effective HSP length: 87
effective length of query: 136
effective length of database: 2,421,580
effective search space: 329334880
effective search space used: 329334880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0096b.3


