

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0095c.6
(384 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
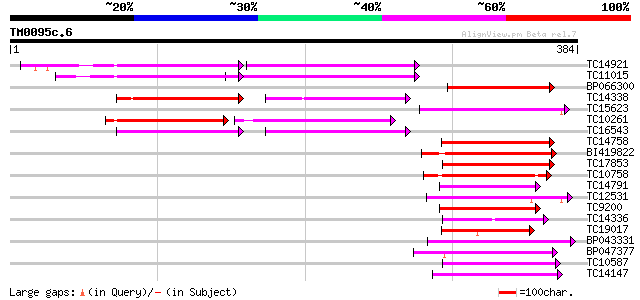
Score E
Sequences producing significant alignments: (bits) Value
TC14921 similar to UP|Q8L949 (Q8L949) Plastid protein, partial (... 94 3e-20
TC11015 similar to PIR|T52623|T52623 DAG protein homolog [import... 87 5e-18
BP066300 77 6e-15
TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%) 72 1e-13
TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding p... 72 2e-13
TC10261 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chloropl... 68 3e-12
TC16543 68 3e-12
TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding p... 64 3e-11
BI419822 63 7e-11
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 62 2e-10
TC10758 57 5e-09
TC14791 similar to UP|Q9LJ04 (Q9LJ04) ESTs AU082210(C53655), par... 55 2e-08
TC12531 similar to UP|Q9ASQ8 (Q9ASQ8) At1g22910/F19G10_13, parti... 55 3e-08
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 54 3e-08
TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, p... 52 1e-07
TC19017 similar to UP|Q8S2X6 (Q8S2X6) Glycine-rich RNA binding p... 52 2e-07
BP043331 51 4e-07
BP047377 51 4e-07
TC10587 weakly similar to GB|AAC50895.1|1518802|HSU63289 CUG-BP/... 50 5e-07
TC14147 similar to UP|ROC1_NICSY (Q08935) 29 kDa ribonucleoprote... 50 5e-07
>TC14921 similar to UP|Q8L949 (Q8L949) Plastid protein, partial (73%)
Length = 1030
Score = 94.4 bits (233), Expect = 3e-20
Identities = 50/117 (42%), Positives = 67/117 (56%)
Frame = +3
Query: 161 SSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEK 220
SSG S T LFP + HWL+ MDKPG E TK ++D Y Q L KV+G+E+
Sbjct: 288 SSGSNSFSDRPPTEMAPLFPGCDYNHWLIVMDKPGGEGATKQDMIDCYVQTLAKVLGSEE 467
Query: 221 DAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
+A+ IY+VS + FGF CE+DE+ + G+ GV V PD + E K+Y L V
Sbjct: 468 EAKKKIYNVSCERYFGFGCEIDEETSNKFEGMNGVLFVLPDSYVDPEYKDYGAELFV 638
Score = 82.8 bits (203), Expect = 9e-17
Identities = 59/164 (35%), Positives = 82/164 (49%), Gaps = 13/164 (7%)
Frame = +3
Query: 8 ISSVNPNRS-----QVHQLPTN--------ITLPNKSYHSNLPSSSSSSISCNRFFTIIA 54
I S +P RS QV PT + S +S L SS S+S S
Sbjct: 159 ILSQSPRRSVYALSQVFFTPTTRFGGIRCRVNRAGDSAYSPLNSSGSNSFSDR------- 317
Query: 55 AAAATTYPSFTPSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLY 114
T P +N HW+++MDKP +K +ID YV+TL V+GSE+EA+ +Y
Sbjct: 318 --PPTEMAPLFPGCDYN-HWLIVMDKPGGEGATKQDMIDCYVQTLAKVLGSEEEAKKKIY 488
Query: 115 DASWDTHFGFCCDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
+ S + +FGF C+IDEE S++ + GVL V PD + KDY
Sbjct: 489 NVSCERYFGFGCEIDEETSNKFEGMNGVLFVLPDSYVDPEYKDY 620
>TC11015 similar to PIR|T52623|T52623 DAG protein homolog [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(65%)
Length = 924
Score = 87.0 bits (214), Expect = 5e-18
Identities = 47/132 (35%), Positives = 73/132 (54%), Gaps = 1/132 (0%)
Frame = +1
Query: 147 PDPDFNSLKKDYSLSSGQ-AGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIV 205
P F ++ + L++G + +S LFP + HWL+ MDKPG E TK Q++
Sbjct: 163 PASRFTDVRFNSELTTGSPSSISEKPAAEMPPLFPGCDYNHWLIIMDKPGGEGATKKQMI 342
Query: 206 DYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFE 265
D Y + L KV+G+E++A+ IY +S + GF CE+DE+ + L G G+ V PD +
Sbjct: 343 DCYIKTLAKVVGSEEEAKKRIYSISCEKYLGFGCEIDEETSDKLDGKAGIMFVLPDSYVD 522
Query: 266 SENKNYEDSLSV 277
E ++Y L V
Sbjct: 523 PEYQDYGGELFV 558
Score = 83.6 bits (205), Expect = 5e-17
Identities = 48/127 (37%), Positives = 70/127 (54%)
Frame = +1
Query: 32 YHSNLPSSSSSSISCNRFFTIIAAAAATTYPSFTPSTTHNRHWMVLMDKPPLGVISKSQV 91
++S L + S SSIS A P P +N HW+++MDKP +K Q+
Sbjct: 190 FNSELTTGSPSSIS---------EKPAAEMPPLFPGCDYN-HWLIIMDKPGGEGATKKQM 339
Query: 92 IDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEISSQLASLPGVLSVRPDPDF 151
ID Y+KTL V+GSE+EA+ +Y S + + GF C+IDEE S +L G++ V PD
Sbjct: 340 IDCYIKTLAKVVGSEEEAKKRIYSISCEKYLGFGCEIDEETSDKLDGKAGIMFVLPDSYV 519
Query: 152 NSLKKDY 158
+ +DY
Sbjct: 520 DPEYQDY 540
>BP066300
Length = 420
Score = 76.6 bits (187), Expect = 6e-15
Identities = 34/73 (46%), Positives = 56/73 (76%)
Frame = -3
Query: 297 TGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEMNGK 356
+GLSF T+E++LR F+ FG+LV V +++DK++ R +G+AF+ Y TEE + A++ M+GK
Sbjct: 418 SGLSFRTTEESLRNCFKNFGQLVDVXLVMDKLANRPRGFAFLRYATEEESQKAIEGMHGK 239
Query: 357 IINGWMIVVDVAK 369
++G +I V+VAK
Sbjct: 238 FLDGRVIFVEVAK 200
>TC14338 similar to UP|Q9MAH5 (Q9MAH5) F12M16.16, partial (33%)
Length = 1751
Score = 72.4 bits (176), Expect = 1e-13
Identities = 38/86 (44%), Positives = 54/86 (62%)
Frame = +2
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+V+M+KP G ++ +ID Y+KTL V+GSE+EA+M +Y S +F F + EE+
Sbjct: 323 HWLVVMEKPE-GDPTRDDIIDGYIKTLAQVVGSEQEARMKIYSVSTKHYFAFGALVSEEL 499
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S +L LP V V PD N +KDY
Sbjct: 500 SYKLKELPKVRWVLPDSYLNVREKDY 577
Score = 58.9 bits (141), Expect = 1e-09
Identities = 34/98 (34%), Positives = 54/98 (54%)
Frame = +2
Query: 174 RTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKT 233
+ +L + +HWLV M+KP + T+ I+D Y + L +V+G+E++A+M IY VS K
Sbjct: 287 KETILLDGCDFEHWLVVMEKPEGDP-TRDDIIDGYIKTLAQVVGSEQEARMKIYSVSTKH 463
Query: 234 NFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
F F + E+ + L +P V V PD K+Y
Sbjct: 464 YFAFGALVSEELSYKLKELPKVRWVLPDSYLNVREKDY 577
>TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (88%)
Length = 683
Score = 72.0 bits (175), Expect = 2e-13
Identities = 37/104 (35%), Positives = 62/104 (59%), Gaps = 2/104 (1%)
Frame = +1
Query: 278 PNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAF 337
P +S + + + KLF+ GLS+ +K+L AF FG + + +VI+D+ + RS+G+ F
Sbjct: 142 PVASMLNYIRCMSSSKLFIGGLSYGVDDKSLEDAFSSFGTVAEARVIVDRDTGRSRGFGF 321
Query: 338 IEYTTEEAAGAALKEMNGKIINGWMIVVDVAKTNP--PRFNRGH 379
+ + +EE+A +AL M+G+ +NG I V A P PR G+
Sbjct: 322 VSFDSEESASSALSSMDGQDLNGRNIRVSYANDRPAGPRGGGGY 453
>TC10261 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chloroplast
precursor, partial (41%)
Length = 510
Score = 67.8 bits (164), Expect = 3e-12
Identities = 35/83 (42%), Positives = 51/83 (61%)
Frame = +2
Query: 66 PSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFC 125
P +N HW+++M+ P S+ Q+I+ Y+ TL TV+GS +EA+ +Y S T+ GF
Sbjct: 257 PGCDYN-HWLIVMEFPKDPAPSREQMIETYLFTLSTVLGSMEEAKKNMYAFSTTTYTGFQ 433
Query: 126 CDIDEEISSQLASLPGVLSVRPD 148
C +DE S + LPGVL V PD
Sbjct: 434 CTVDEATSEKFKGLPGVLWVLPD 502
Score = 62.0 bits (149), Expect = 2e-10
Identities = 35/109 (32%), Positives = 59/109 (54%)
Frame = +2
Query: 153 SLKKDYSLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQIL 212
+L +DYS A SS + R ++ P + HWL+ M+ P ++ Q+++ Y L
Sbjct: 191 ALDEDYS-----AKRSSSSEQRETIMLPGCDYNHWLIVMEFPKDPAPSREQMIETYLFTL 355
Query: 213 TKVMGNEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPD 261
+ V+G+ ++A+ +Y S T GF C +DE ++ G+PGV V PD
Sbjct: 356 STVLGSMEEAKKNMYAFSTTTYTGFQCTVDEATSEKFKGLPGVLWVLPD 502
>TC16543
Length = 575
Score = 67.8 bits (164), Expect = 3e-12
Identities = 34/86 (39%), Positives = 52/86 (59%)
Frame = +1
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+++M+ P S+ ++++ YVKTL ++GSE+EA +Y S T+ GF I EE+
Sbjct: 313 HWLIVMEFPENPKPSEQEMVNAYVKTLTQIVGSEEEAMKKIYSVSTHTYTGFGALISEEL 492
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S ++ PGVL V PD + KDY
Sbjct: 493 SYKVKEXPGVLWVLPDSYLDVPNKDY 570
Score = 57.8 bits (138), Expect = 3e-09
Identities = 30/98 (30%), Positives = 54/98 (54%)
Frame = +1
Query: 174 RTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKT 233
+ +L + +HWL+ M+ P ++ ++V+ Y + LT+++G+E++A IY VS T
Sbjct: 277 KETILLDGCDYEHWLIVMEFPENPKPSEQEMVNAYVKTLTQIVGSEEEAMKKIYSVSTHT 456
Query: 234 NFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
GF + E+ + + PGV V PD + NK+Y
Sbjct: 457 YTGFGALISEELSYKVKEXPGVLWVLPDSYLDVPNKDY 570
>TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding protein 2,
partial (93%)
Length = 725
Score = 64.3 bits (155), Expect = 3e-11
Identities = 30/77 (38%), Positives = 51/77 (65%)
Frame = +1
Query: 293 KLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKE 352
+ FV GL++ T TL AF +GE++ KV+ D+ + RS+G+ F+ +T+EEA +A++
Sbjct: 85 RCFVGGLAWTTDSHTLEQAFSSYGEIIDSKVVNDRETGRSRGFGFVTFTSEEAMRSAIEG 264
Query: 353 MNGKIINGWMIVVDVAK 369
MNG ++G I V+ A+
Sbjct: 265 MNGNELDGRNITVNEAQ 315
>BI419822
Length = 380
Score = 63.2 bits (152), Expect = 7e-11
Identities = 35/91 (38%), Positives = 57/91 (62%)
Frame = +2
Query: 280 SSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIE 339
SS AS E R FV GL++ T + L AF +GE+V+ KVI D+ + RS+G+ F+
Sbjct: 50 SSMASAEIEFRC---FVGGLAWATDNEALEKAFSPYGEIVESKVINDRETGRSRGFGFVT 220
Query: 340 YTTEEAAGAALKEMNGKIINGWMIVVDVAKT 370
+ +E+A A++ MNG+ ++G I V+ A++
Sbjct: 221 FASEQAMKDAIEAMNGQNLDGRNITVNEAQS 313
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 62.0 bits (149), Expect = 2e-10
Identities = 30/76 (39%), Positives = 49/76 (64%)
Frame = +2
Query: 294 LFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEM 353
LF ++ TS++ L+ AFE FG+L + KV++DK S RS+G+ F+ + ++A A+ M
Sbjct: 62 LFYWWPAWSTSDRKLKDAFEKFGKLTEAKVVVDKFSGRSRGFGFVTFDDKKAMDEAIDAM 241
Query: 354 NGKIINGWMIVVDVAK 369
NG ++G I VD A+
Sbjct: 242 NGMDLDGRTITVDKAQ 289
>TC10758
Length = 532
Score = 57.0 bits (136), Expect = 5e-09
Identities = 35/87 (40%), Positives = 54/87 (61%)
Frame = +1
Query: 281 SEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEY 340
S A ++ ++R KLFV GL+ T+ +TLR+ F +G+L + VI DK + RSKGY F+ +
Sbjct: 280 SVADRDTAVR--KLFVRGLACETTTETLRSVFSAYGDLDEAIVIFDKSTGRSKGYGFVVF 453
Query: 341 TTEEAAGAALKEMNGKIINGWMIVVDV 367
+ A ALK+ + K I+G M V +
Sbjct: 454 KHIDGAILALKDPSKK-IDGRMTVTQL 531
>TC14791 similar to UP|Q9LJ04 (Q9LJ04) ESTs AU082210(C53655), partial (37%)
Length = 1779
Score = 55.1 bits (131), Expect = 2e-08
Identities = 27/68 (39%), Positives = 41/68 (59%)
Frame = +2
Query: 292 KKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALK 351
+K+FV GL + T+ +TL + F +GE+ K + DK++ +SKGYAFI + + ALK
Sbjct: 608 RKIFVHGLGWDTTAETLTSVFSKYGEIEDCKAVTDKVTGKSKGYAFILFKHRDGCRRALK 787
Query: 352 EMNGKIIN 359
KI N
Sbjct: 788 NPQKKIGN 811
Score = 38.9 bits (89), Expect = 0.001
Identities = 25/77 (32%), Positives = 40/77 (51%)
Frame = +2
Query: 278 PNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAF 337
PN+ S+ +K+FV+ +S L F+ FGE+ + +DK + R KG+A
Sbjct: 863 PNAPPVSEYTQ---RKIFVSNVSSDIEPLKLLEFFKQFGEVEDGPLGLDKNTGRPKGFAL 1033
Query: 338 IEYTTEEAAGAALKEMN 354
Y + E+A AL+E N
Sbjct: 1034FVYRSVESAKKALEEPN 1084
>TC12531 similar to UP|Q9ASQ8 (Q9ASQ8) At1g22910/F19G10_13, partial (39%)
Length = 688
Score = 54.7 bits (130), Expect = 3e-08
Identities = 34/105 (32%), Positives = 58/105 (54%), Gaps = 6/105 (5%)
Frame = -1
Query: 283 ASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTT 342
A Q K+FV GL++ T ++T++ FE FG++++ VI DK + RSKGY F+ +
Sbjct: 388 AGQFGDTTYTKVFVGGLAWETQKETMKKYFEQFGDILEAVVITDKATGRSKGYGFVTFRE 209
Query: 343 EEAAGAALKE----MNGKIINGWMIVVDVAKTNP--PRFNRGHAR 381
EAA A + ++G+ N + + V ++ P P+ + G R
Sbjct: 208 PEAAMRACVDAAPVIDGRRANCNLASLGVQRSKPSTPKQHGGAGR 74
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 54.3 bits (129), Expect = 3e-08
Identities = 28/68 (41%), Positives = 41/68 (60%)
Frame = +1
Query: 292 KKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALK 351
+K+FV GL + T+ TL AF+ +G + K + DK++ +SKGY FI + + A ALK
Sbjct: 466 RKIFVHGLGWDTTATTLVYAFQQYGAIEDCKAVTDKVTGKSKGYGFILFKSRRGARNALK 645
Query: 352 EMNGKIIN 359
E KI N
Sbjct: 646 EPQKKIGN 669
>TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(85%)
Length = 1917
Score = 52.4 bits (124), Expect = 1e-07
Identities = 26/72 (36%), Positives = 41/72 (56%)
Frame = +2
Query: 294 LFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEM 353
+F+ L K L F FG ++ KV D S +SKGY F+++ TEEAA A++++
Sbjct: 98 IFIKNLDKAIDHKALHDTFSTFGNILSCKVATDS-SGQSKGYGFVQFETEEAAQNAIEKL 274
Query: 354 NGKIINGWMIVV 365
NG ++N + V
Sbjct: 275 NGMLLNDKQVYV 310
>TC19017 similar to UP|Q8S2X6 (Q8S2X6) Glycine-rich RNA binding protein
(Fragment), partial (26%)
Length = 320
Score = 51.6 bits (122), Expect = 2e-07
Identities = 25/67 (37%), Positives = 46/67 (68%), Gaps = 4/67 (5%)
Frame = +2
Query: 293 KLFVTGLSFYTSEKTLRAAFEGF----GELVQVKVIIDKISKRSKGYAFIEYTTEEAAGA 348
+LFV LS+ T+++++R AFE G ++ KVI+D+ + RS+G+ F+ +++ + A A
Sbjct: 107 RLFVGNLSWNTTDESMRMAFEDGAGVPGSVLDSKVILDRETGRSRGFGFVTFSSPDLASA 286
Query: 349 ALKEMNG 355
A+ +MNG
Sbjct: 287 AIAKMNG 307
>BP043331
Length = 420
Score = 50.8 bits (120), Expect = 4e-07
Identities = 28/100 (28%), Positives = 52/100 (52%)
Frame = +3
Query: 284 SQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTE 343
++ AS T LFV G+S T+++ L AF G++ V K+ + + + + +TT
Sbjct: 99 NKAASAPTSMLFVGGISPTTTDRHLDDAFSIHGDVYLATVKHTKVDGKPRQFGLVAFTTV 278
Query: 344 EAAGAALKEMNGKIINGWMIVVDVAKTNPPRFNRGHARPP 383
E A AL+ +NG+++ G + ++ K + + H PP
Sbjct: 279 ECATRALQALNGQVLQGRRLRINYTKNSHSYMDGHHYDPP 398
>BP047377
Length = 564
Score = 50.8 bits (120), Expect = 4e-07
Identities = 31/100 (31%), Positives = 55/100 (55%), Gaps = 2/100 (2%)
Frame = +3
Query: 274 SLSVPNSSEASQEASLRTKK--LFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKR 331
S + P+SS AS +++ +FV + + +E+ L + G +V +++ID+ + +
Sbjct: 69 SRASPSSSAVDFMASSQSQHRCVFVGNIPYDATEEQLIEICQEVGPVVSFRLVIDRETGK 248
Query: 332 SKGYAFIEYTTEEAAGAALKEMNGKIINGWMIVVDVAKTN 371
KGY F EY EE A +A + + G ING + VD A+ +
Sbjct: 249 PKGYGFCEYKDEETALSARRNLQGYEINGRQLRVDFAEND 368
>TC10587 weakly similar to GB|AAC50895.1|1518802|HSU63289 CUG-BP/hNab50
{Homo sapiens;} , partial (19%)
Length = 482
Score = 50.4 bits (119), Expect = 5e-07
Identities = 26/80 (32%), Positives = 43/80 (53%)
Frame = +1
Query: 294 LFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEM 353
LF+ + ++ L AF+ FG ++ KV +DK + SK + F+ Y + EAA +A+ M
Sbjct: 19 LFIYHIPQEFGDQDLANAFQPFGRVISAKVFVDKATGVSKCFGFVSYDSPEAAQSAISMM 198
Query: 354 NGKIINGWMIVVDVAKTNPP 373
NG + G + V + N P
Sbjct: 199 NGYQLGGKKLKVQHKRDNKP 258
>TC14147 similar to UP|ROC1_NICSY (Q08935) 29 kDa ribonucleoprotein A,
chloroplast precursor (CP29A), partial (63%)
Length = 1602
Score = 50.4 bits (119), Expect = 5e-07
Identities = 25/88 (28%), Positives = 47/88 (53%)
Frame = +3
Query: 287 ASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAA 346
+S ++ V L++ L + F G +V KVI D+ S RS+G+ F+ +++ +
Sbjct: 723 SSYSENRVHVGNLAWGVDNAALESLFREQGRVVDAKVIYDRESGRSRGFGFVTFSSPDEV 902
Query: 347 GAALKEMNGKIINGWMIVVDVAKTNPPR 374
+A++ ++G +NG I V A + P R
Sbjct: 903 NSAIRSLDGADLNGRAIKVSQADSKPKR 986
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.130 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,022,557
Number of Sequences: 28460
Number of extensions: 102703
Number of successful extensions: 854
Number of sequences better than 10.0: 105
Number of HSP's better than 10.0 without gapping: 818
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 849
length of query: 384
length of database: 4,897,600
effective HSP length: 92
effective length of query: 292
effective length of database: 2,279,280
effective search space: 665549760
effective search space used: 665549760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0095c.6


