

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
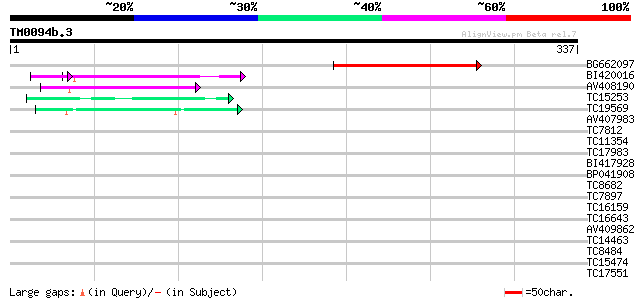
Query= TM0094b.3
(337 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG662097 96 9e-21
BI420016 39 3e-05
AV408190 42 1e-04
TC15253 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-pro... 42 1e-04
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 40 7e-04
AV407983 39 0.001
TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic me... 38 0.003
TC11354 similar to UP|CAE54480 (CAE54480) Alpha glucosidase II ... 37 0.004
TC17983 similar to UP|Q9XEX5 (Q9XEX5) Cytidine deaminase , part... 37 0.006
BI417928 37 0.006
BP041908 36 0.008
TC8682 similar to UP|HPPD_SOLSC (Q9ARF9) 4-hydroxyphenylpyruvate... 36 0.008
TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-li... 36 0.010
TC16159 homologue to UP|Q94A23 (Q94A23) AT5g64310/MSJ1_15, parti... 36 0.010
TC16643 similar to UP|Q9FIH3 (Q9FIH3) Serine/threonine protein k... 35 0.014
AV409862 35 0.014
TC14463 similar to UP|Q9SLF1 (Q9SLF1) Nodulin-like protein (At2g... 35 0.014
TC8484 similar to UP|Q9XGP1 (Q9XGP1) ESTs AU070372(S13446), part... 35 0.014
TC15474 UP|Q7Y1B9 (Q7Y1B9) Ammonium transporter, complete 35 0.018
TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, part... 35 0.018
>BG662097
Length = 350
Score = 95.9 bits (237), Expect = 9e-21
Identities = 52/88 (59%), Positives = 64/88 (72%)
Frame = -2
Query: 193 LRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCKL 252
L +VE L +DL AKE T+ASQEK LA+K KEI+DL E L + E LAE +SDL KCKL
Sbjct: 274 LENQVEGLSKDLSAKEATIASQEKSLASKVKEISDLHERLATKCELLAEVRSDLESKCKL 95
Query: 253 LADYEAQAATLKEKIRSEAAHAAAKERV 280
LA+ +AQA L+E+IR EAA A A ER+
Sbjct: 94 LAEKDAQAIALEEQIRREAAFATANERI 11
>BI420016
Length = 559
Score = 39.3 bits (90), Expect(2) = 3e-05
Identities = 37/114 (32%), Positives = 47/114 (40%), Gaps = 5/114 (4%)
Frame = +2
Query: 32 PDPQTT--PLTG-TGGGFVAPDPVVTDSACDPLASD-HPSEVNASQTPDAPRPVTLAEGQ 87
P+PQT P T T GG +P PV + P + P S TP+AP +
Sbjct: 155 PEPQTLKKPTTQCTHGGNPSPKPVPASTPSPPSSPTLTPPPSPPSPTPNAPLSPSSPPSP 334
Query: 88 RSPSSIRP-PIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYPFS 140
+P S P P P+P G + P+SP PT S SSP FS
Sbjct: 335 PTPPSPPPSPAPNPTPSATGSTTPSSP-----------LTPTSASLSSPSSLFS 463
Score = 24.3 bits (51), Expect(2) = 3e-05
Identities = 11/25 (44%), Positives = 14/25 (56%)
Frame = +1
Query: 13 KRQSSPVSACSPKRSKVGEPDPQTT 37
+ SSP S+ S S G P+PQ T
Sbjct: 106 RNNSSPSSSNSSSMSAAGTPNPQKT 180
>AV408190
Length = 382
Score = 42.4 bits (98), Expect = 1e-04
Identities = 31/100 (31%), Positives = 44/100 (44%), Gaps = 5/100 (5%)
Frame = +3
Query: 19 VSACSPKRSKVGEPDP----QTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNA-SQ 73
V++ +P +S P QTTP T AP S+ PLA+ P+ +
Sbjct: 72 VASAAPIQSSTASGSPSLSLQTTP*TSLLSSPPAPTMARPPSSTLPLAASSPTNTSGPPS 251
Query: 74 TPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSP 113
TP P T A + + SS PP PS + PS P++P
Sbjct: 252 TPSPPLSPTWASERATSSSSFPPTPSTSPSSASPSCPSAP 371
>TC15253 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-protein
(ATAGP4) (AT5G10430/F12B17_220), partial (42%)
Length = 575
Score = 42.0 bits (97), Expect = 1e-04
Identities = 34/123 (27%), Positives = 42/123 (33%)
Frame = +3
Query: 11 GGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVN 70
G +P + P + P P T P T P P T A P S
Sbjct: 156 GAAPTQAPTTTPPPPPAAAPAPPPATPPPAAT------PAPTTTPPAATPAPS------- 296
Query: 71 ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGP 130
+P AP P G +PS+ PP P P GP PG SA P P
Sbjct: 297 --ASPPAPTPTASPTGAPTPSASSPPAPIPSGPASGPGPAAGPGPN------SADTPPPP 452
Query: 131 SSS 133
S++
Sbjct: 453 SAA 461
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 39.7 bits (91), Expect = 7e-04
Identities = 41/129 (31%), Positives = 50/129 (37%), Gaps = 6/129 (4%)
Frame = +1
Query: 16 SSPVSACSPKRSKVGEP--DPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ 73
SS S+ SP +P P T+P T T +P V + P S +AS+
Sbjct: 49 SSSSSSYSPPPPPSQQPTHSPSTSP-TSTTTPPPSPTKVTENPPTAPSTSTKSPTSSASE 225
Query: 74 TPDAPRPVTLAEGQRSPSSIRPPI---PSPVKVTLGPSAPNSPGVGGEGENLS-AAQPTG 129
P P P T Q S PP PSP T P+A SP S P
Sbjct: 226 EPSPPNPSTSTTQQPKHSPTSPPASPSPSPPS-TKPPTAMASPSTSPLSATKSHPTPPAA 402
Query: 130 PSSSSPPYP 138
PS+SS P P
Sbjct: 403 PSASSTPPP 429
Score = 37.4 bits (85), Expect = 0.004
Identities = 31/107 (28%), Positives = 42/107 (38%), Gaps = 3/107 (2%)
Frame = +1
Query: 56 SACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGV 115
S C L S ++ P P SP+S P PSP KVT P P +P
Sbjct: 19 SPCGLLLHTSSSSSSSYSPPPPPSQQPTHSPSTSPTSTTTPPPSPTKVTENP--PTAPST 192
Query: 116 GGEGENLSAAQ---PTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
+ SA++ P PS+S+ P S P +PS +P
Sbjct: 193 STKSPTSSASEEPSPPNPSTSTTQQPKHSPTSPP------ASPSPSP 315
>AV407983
Length = 417
Score = 39.3 bits (90), Expect = 0.001
Identities = 26/83 (31%), Positives = 40/83 (47%), Gaps = 7/83 (8%)
Frame = +1
Query: 61 LASDHPSEVNASQTPDAPRPVTLAEG-QRSPSS---IRPPIPSPVKVTLGPSAPNSPGVG 116
+ S HPS N+S P++ + SPSS + PP+ +P +L SAP+S
Sbjct: 133 IPSPHPSPQNSSSAS*NTSPISASSSPSHSPSSSPPLLPPLSAPTSSSLSSSAPSSTSSS 312
Query: 117 GEGENLSAAQ---PTGPSSSSPP 136
N S A PT P++++ P
Sbjct: 313 SASSNPSLADLALPTPPTTTTTP 381
>TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic membrane
protein 1, complete
Length = 1149
Score = 37.7 bits (86), Expect = 0.003
Identities = 42/160 (26%), Positives = 60/160 (37%), Gaps = 11/160 (6%)
Frame = +1
Query: 1 MSVKPRVAEVGGKRQSSPVSACSPKRSKV-----GEPDPQTTPLTGTGGGFVAPDPVVTD 55
+SV R V G+R S S P R + G+P ++P + + P+ T+
Sbjct: 34 VSVDHRFPTVAGRRCRSATSPSVPLRRQPTQTP*GQPWLSSSPPSSSSSPAPVPESPTTN 213
Query: 56 SACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNS--- 112
S P+ +P P SPS P P+ PSA +S
Sbjct: 214 S---------PTTAPPPPPASSPPPSPTHSPSSSPSPSAPTSPAATSTPPSPSASSSAVT 366
Query: 113 --PGVGGEGENLSAAQPTGPSSSSPPYPFS-FLLSHPYLE 149
+LS + P+ P SSP P S +LLS LE
Sbjct: 367 SPSSAVSSTSSLSFSDPSSPPCSSPSSPDSLYLLSVFQLE 486
>TC11354 similar to UP|CAE54480 (CAE54480) Alpha glucosidase II , partial
(6%)
Length = 583
Score = 37.4 bits (85), Expect = 0.004
Identities = 27/106 (25%), Positives = 41/106 (38%), Gaps = 2/106 (1%)
Frame = -1
Query: 66 PSEVNASQTPDAPRPVTLAEGQRSPSSIRPP--IPSPVKVTLGPSAPNSPGVGGEGENLS 123
PS S +P A P T + P+ P PSP T ++P++ + +
Sbjct: 340 PSSPPMSPSPTATSPPTSSPNTHPPTPTPNP*SSPSPSTTTASSASPSTKTLPSTRPRPA 161
Query: 124 AAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQEAVLN 169
+ PT S SSPP +S P + PS +P + N
Sbjct: 160 SESPTSSSPSSPPPSSGSSVSPPRISPPPPPPSNSPTATPPSSATN 23
Score = 36.2 bits (82), Expect = 0.008
Identities = 36/120 (30%), Positives = 45/120 (37%), Gaps = 1/120 (0%)
Frame = -1
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDP-VVTDSACDPLASDHPSEVNASQT 74
SSP + SP + P T P T T +P P T S+ P PS
Sbjct: 337 SSPPMSPSPTATSPPTSSPNTHPPTPTPNP*SSPSPSTTTASSASPSTKTLPST------ 176
Query: 75 PDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSS 134
PRP + + SPSS PP S V+ +P P + T PSSSS
Sbjct: 175 --RPRPASESPTSSSPSS--PPPSSGSSVSPPRISPPPPPPSNSPTATPPSSATNPSSSS 8
>TC17983 similar to UP|Q9XEX5 (Q9XEX5) Cytidine deaminase , partial (42%)
Length = 519
Score = 36.6 bits (83), Expect = 0.006
Identities = 36/117 (30%), Positives = 50/117 (41%), Gaps = 16/117 (13%)
Frame = +2
Query: 47 VAPDPVVTDSACDPL--ASDHPSEVNASQTPDAPRPV---------TLAEGQRSPS---- 91
V+P P + ++ P + DHPS + S D+P V +LA +PS
Sbjct: 86 VSP*PNSSQASSTPQKPSPDHPSPTSGSPPSDSPHRVGSSSESTSNSLASPSTTPSTPSS 265
Query: 92 SIRPPIPSPVKVTLGPS-APNSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPY 147
S P PS K PS +P P + ++A+P SSSP P SHPY
Sbjct: 266 SSSPTSPSTTKQISTPSPSPPPPVATAASSSKNSAEPRRFKSSSPQNPTR--NSHPY 430
>BI417928
Length = 509
Score = 36.6 bits (83), Expect = 0.006
Identities = 30/103 (29%), Positives = 41/103 (39%), Gaps = 4/103 (3%)
Frame = +2
Query: 36 TTPLTGTGGGFVAPDPVVTDSACDPL----ASDHPSEVNASQTPDAPRPVTLAEGQRSPS 91
TTPL P P S PL +S P S P P T A+ +
Sbjct: 170 TTPLIPPPTLSPPPPPPPRSSTPPPLRLSSSSPWPPFCPTSPPPKPTTPPTTAKA----T 337
Query: 92 SIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSS 134
++ P+P+P PSAP+ P G + LS PS++S
Sbjct: 338 TVASPLPTPPTTPSPPSAPSPPSTYGPSKPLSVTLTGAPSTAS 466
>BP041908
Length = 487
Score = 36.2 bits (82), Expect = 0.008
Identities = 28/97 (28%), Positives = 39/97 (39%), Gaps = 13/97 (13%)
Frame = +3
Query: 16 SSPVSACSPKRSKVGEPDPQT-TPLTGTGGGFVAPDPVVTDSACD------------PLA 62
+S +S P P P T TP T + P P T+SA PL
Sbjct: 177 ASKISLTLPHDPASSTPPPATSTPTTTSTSPPGEPPPASTNSATSKKETSS*SSSPTPLN 356
Query: 63 SDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPS 99
S PS + + P P + + ++SPS +PP PS
Sbjct: 357 SSSPSSPHLTSAPWPPPQIPSSPPRKSPSKPKPPTPS 467
>TC8682 similar to UP|HPPD_SOLSC (Q9ARF9) 4-hydroxyphenylpyruvate
dioxygenase (4HPPD) (HPD) (HPPDase) , partial (29%)
Length = 576
Score = 36.2 bits (82), Expect = 0.008
Identities = 35/134 (26%), Positives = 50/134 (37%), Gaps = 9/134 (6%)
Frame = +1
Query: 12 GKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTD---SACDPLASDHPSE 68
G S ++ P +++ G P +T T + G P P SAC +
Sbjct: 181 GSNSSGSTASSGPTQNQTGSP*KGST--TSSSGAPTPPTPHAASPGASACRSSQNQTSPP 354
Query: 69 VNASQTPDAPRPVTLAEGQ------RSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENL 122
P + P T A +SPS PP PSP+ P++P+ P
Sbjct: 355 ETTLTPPTSSAPATSASSSPPHTLPKSPSP--PPPPSPLSPP-PPASPSPPPTASPSAPS 525
Query: 123 SAAQPTGPSSSSPP 136
+ PT PS SPP
Sbjct: 526 PSKSPT-PSKPSPP 564
>TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-like protein
2 precursor (Phytocyanin-like protein), partial (31%)
Length = 1482
Score = 35.8 bits (81), Expect = 0.010
Identities = 32/125 (25%), Positives = 50/125 (39%)
Frame = +3
Query: 14 RQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ 73
R + P + PK S P P+ +P +G+ P P T + P + P
Sbjct: 516 RGTQPPPSPPPKSSP--SPSPKASPPSGSPPTAGEPSPAGTPPSVLPSPAGAPVPAGG-- 683
Query: 74 TPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSS 133
P A P PS+ P PSPV + P+A +PG + ++ P G ++
Sbjct: 684 -PSAGTP---------PSASTSPSPSPV--SAPPAASPAPGAESPSSSPTSNSPAGGPTA 827
Query: 134 SPPYP 138
+ P P
Sbjct: 828 TSPSP 842
Score = 35.4 bits (80), Expect = 0.014
Identities = 33/129 (25%), Positives = 44/129 (33%), Gaps = 1/129 (0%)
Frame = +3
Query: 11 GGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVN 70
G + SP SP S P + P G P V+ A P+ + PS
Sbjct: 519 GTQPPPSPPPKSSPSPSPKASPPSGSPPTAGEPSPAGTPPSVLPSPAGAPVPAGGPSAGT 698
Query: 71 ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGP 130
+P P SP S PP SP PS+ + G ++ P G
Sbjct: 699 PPSASTSPSP--------SPVSA-PPAASPAPGAESPSSSPTSNSPAGGPTATSPSPAGD 851
Query: 131 S-SSSPPYP 138
S + PP P
Sbjct: 852 SPAGGPPAP 878
>TC16159 homologue to UP|Q94A23 (Q94A23) AT5g64310/MSJ1_15, partial (19%)
Length = 605
Score = 35.8 bits (81), Expect = 0.010
Identities = 26/88 (29%), Positives = 39/88 (43%)
Frame = +2
Query: 34 PQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSI 93
P TP+ + ++P P V+ SA P AS P+ V + +P P T+ P+S
Sbjct: 140 PTKTPVAASPRKSLSPSPAVSPSAQTPAASP-PTPVESGPSPS---PATV---NSPPASS 298
Query: 94 RPPIPSPVKVTLGPSAPNSPGVGGEGEN 121
P +P ++ PS SP G N
Sbjct: 299 DAPAVTPSSISSPPSEAQSPSQNGAALN 382
>TC16643 similar to UP|Q9FIH3 (Q9FIH3) Serine/threonine protein kinase-like
protein, partial (28%)
Length = 585
Score = 35.4 bits (80), Expect = 0.014
Identities = 25/98 (25%), Positives = 37/98 (37%), Gaps = 14/98 (14%)
Frame = +2
Query: 63 SDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENL 122
S P +++ T + P P A + P+ P P+P + P+ P P +N
Sbjct: 80 SSSPPPSSSASTANPPNPEPAATTRSEPALSETPTPAPPSPSSTPAGPGIPNSRSPWKNS 259
Query: 123 SAAQPTGPSS--------------SSPPYPFSFLLSHP 146
A Q T P + +SPP P S S P
Sbjct: 260 PAPQKTSPRA*LSATAASA*STRLASPPAPSSPSRSSP 373
>AV409862
Length = 420
Score = 35.4 bits (80), Expect = 0.014
Identities = 28/85 (32%), Positives = 36/85 (41%)
Frame = +3
Query: 52 VVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPN 111
+ T S+ P A+ PS N S T P T SPS PP S T SA
Sbjct: 54 IPTLSSPSPFAASPPSPRNPSPTTLPPPTTT-----NSPSKPEPPATSTPSATSSTSASK 218
Query: 112 SPGVGGEGENLSAAQPTGPSSSSPP 136
+ +A+ P+ PS+SSPP
Sbjct: 219 T----------AASTPSTPSASSPP 263
>TC14463 similar to UP|Q9SLF1 (Q9SLF1) Nodulin-like protein
(At2g16660/T24I21.7), partial (40%)
Length = 1122
Score = 35.4 bits (80), Expect = 0.014
Identities = 29/88 (32%), Positives = 38/88 (42%), Gaps = 2/88 (2%)
Frame = +2
Query: 51 PVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPS-- 108
P SA P + +PS + +P P P SPSS PS +K ++ PS
Sbjct: 137 PPANGSASSPPSGSNPSPATTTPSPTTPTPSN--PSCTSPSSSSTTSPS-LKTSVKPSVS 307
Query: 109 APNSPGVGGEGENLSAAQPTGPSSSSPP 136
+P SP + P GPSSSS P
Sbjct: 308 SPASP---------PTSSPLGPSSSSAP 364
>TC8484 similar to UP|Q9XGP1 (Q9XGP1) ESTs AU070372(S13446), partial (76%)
Length = 1121
Score = 35.4 bits (80), Expect = 0.014
Identities = 26/84 (30%), Positives = 34/84 (39%), Gaps = 5/84 (5%)
Frame = +1
Query: 60 PLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSP-----VKVTLGPSAPNSPG 114
P S HP+ + +P +P T+A +PS PP PSP +L P P S
Sbjct: 142 PPLSSHPNPSSPPNSPPSPSSSTVA--TPNPSDSHPPPPSPPPSPSATNSLNPPYPTSTT 315
Query: 115 VGGEGENLSAAQPTGPSSSSPPYP 138
+ S P G SS P P
Sbjct: 316 PARSKPSPSLNSPRGRRQSSSPSP 387
Score = 26.9 bits (58), Expect = 4.9
Identities = 18/50 (36%), Positives = 21/50 (42%), Gaps = 1/50 (2%)
Frame = +1
Query: 90 PSSIRPPIPSPVKVTLGPSAP-NSPGVGGEGENLSAAQPTGPSSSSPPYP 138
P S PP P+ PS+P NSP + S PS S PP P
Sbjct: 115 PGSFLPPPSPPLSSHPNPSSPPNSP----PSPSSSTVATPNPSDSHPPPP 252
>TC15474 UP|Q7Y1B9 (Q7Y1B9) Ammonium transporter, complete
Length = 1950
Score = 35.0 bits (79), Expect = 0.018
Identities = 35/126 (27%), Positives = 52/126 (40%), Gaps = 6/126 (4%)
Frame = +3
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP 75
SSP S+ P + P TP T F+ P +P+ +AS++P
Sbjct: 234 SSPCSSVLPCSAPA--PSEPKTP*TSC*PMFLMQQPAA-----------YPTTSSASRSP 374
Query: 76 DAPRPV--TLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTG---- 129
P P + A S +++RPP + T GPS P E+ +A P G
Sbjct: 375 SEPPPTASSAATSSASNTTLRPPTTTASSSTSGPSPSPLP------ESPAAPSPRGHNSS 536
Query: 130 PSSSSP 135
P+SS+P
Sbjct: 537 PTSSTP 554
>TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, partial (22%)
Length = 505
Score = 35.0 bits (79), Expect = 0.018
Identities = 32/100 (32%), Positives = 38/100 (38%), Gaps = 1/100 (1%)
Frame = +3
Query: 57 ACDPLAS-DHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGV 115
A PL S PS T P P + + SS PP PSP + P P+S
Sbjct: 171 ASHPLPSLPSPSSAAVGPTSSTP-PTSPTSAAAATSSATPPSPSPPPTSASPPPPSST-- 341
Query: 116 GGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAP 155
G S++ T PS S P P S P L AP
Sbjct: 342 -PNGTPPSSSPATTPSPSRPSTP-----SSPTRASLPSAP 443
Score = 32.3 bits (72), Expect = 0.12
Identities = 24/72 (33%), Positives = 33/72 (45%), Gaps = 6/72 (8%)
Frame = +3
Query: 75 PDAPRPVTLAEGQRSPSSIR-PPIPSPVKVTLGP--SAPNSPGVGGEGENLSAAQPT--- 128
P P +T + SP+S P +PSP +GP S P + SA P+
Sbjct: 123 PPFPPSLTTSFSTSSPASHPLPSLPSPSSAAVGPTSSTPPTSPTSAAAATSSATPPSPSP 302
Query: 129 GPSSSSPPYPFS 140
P+S+SPP P S
Sbjct: 303 PPTSASPPPPSS 338
Score = 30.4 bits (67), Expect = 0.44
Identities = 25/78 (32%), Positives = 37/78 (47%), Gaps = 2/78 (2%)
Frame = +3
Query: 61 LASDHPS--EVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGE 118
LA+ H S V+ QT P P + R+ ++ PP P + + S+P S +
Sbjct: 21 LATSHGS*NLVSPIQTTPPPPPPPPSPNHRN--NLPPPFPPSLTTSFSTSSPASHPLPSL 194
Query: 119 GENLSAAQPTGPSSSSPP 136
SAA GP+SS+PP
Sbjct: 195 PSPSSAA--VGPTSSTPP 242
Score = 30.0 bits (66), Expect = 0.57
Identities = 33/112 (29%), Positives = 41/112 (36%)
Frame = +3
Query: 23 SPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVT 82
SP + VG P T P + T T SA P S P T +P P +
Sbjct: 198 SPSSAAVG-PTSSTPPTSPTSAA------AATSSATPPSPSPPP-------TSASPPPPS 335
Query: 83 LAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSS 134
PSS PSP + PS P+SP A+ P+ P SS
Sbjct: 336 STPNGTPPSSSPATTPSPSR----PSTPSSP--------TRASLPSAPGFSS 455
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.310 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,288,797
Number of Sequences: 28460
Number of extensions: 83509
Number of successful extensions: 927
Number of sequences better than 10.0: 197
Number of HSP's better than 10.0 without gapping: 800
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 885
length of query: 337
length of database: 4,897,600
effective HSP length: 91
effective length of query: 246
effective length of database: 2,307,740
effective search space: 567704040
effective search space used: 567704040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0094b.3


