

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0094a.6
(990 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
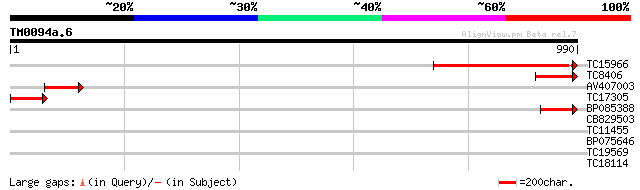
TC15966 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-tudor protein... 466 e-131
TC8406 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-tudor protein,... 151 6e-37
AV407003 138 5e-33
TC17305 weakly similar to UP|Q9FLT0 (Q9FLT0) Transcription facto... 135 3e-32
BP085388 130 1e-30
CB829503 30 2.5
TC11455 similar to UP|Q9ZUS8 (Q9ZUS8) At2g37380 protein, partial... 29 3.2
BP075646 28 7.1
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 28 7.1
TC18114 similar to GB|AAM16218.1|20334714|AY093957 At2g41900/T6D... 28 9.3
>TC15966 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-tudor protein, partial
(25%)
Length = 891
Score = 466 bits (1198), Expect = e-131
Identities = 234/250 (93%), Positives = 241/250 (95%)
Frame = +1
Query: 741 EVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRA 800
EVLGG KFYVQTVGDQKIASIQQQLA+LNLKEAPVLGAFSPKKGD VLCYFH DKSWYRA
Sbjct: 1 EVLGGDKFYVQTVGDQKIASIQQQLASLNLKEAPVLGAFSPKKGDTVLCYFHGDKSWYRA 180
Query: 801 MVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQLCSLAYIKSP 860
MVVNTPRGPVESP+DIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQLCSLAYIKSP
Sbjct: 181 MVVNTPRGPVESPQDIFEVFYIDYGNQEQVAYSQLRPLDQSVSAAPGLAQLCSLAYIKSP 360
Query: 861 SLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAVTLVAVDAEIS 920
SLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGK KGQGTGTILAVTLVAVDAEIS
Sbjct: 361 SLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKAKGQGTGTILAVTLVAVDAEIS 540
Query: 921 VNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESDDEDTAP 980
VNAAMLQEGLARMEKRNRWDRKERK GLDSL+KFQD+AR +RRGMWQYGDVESD+ED P
Sbjct: 541 VNAAMLQEGLARMEKRNRWDRKERKAGLDSLEKFQDEARTKRRGMWQYGDVESDEED-GP 717
Query: 981 PARKAGAGRR 990
PARKAG GR+
Sbjct: 718 PARKAGTGRK 747
>TC8406 similar to UP|Q8L5C2 (Q8L5C2) 110 kDa 4SNc-tudor protein, partial
(7%)
Length = 562
Score = 151 bits (381), Expect = 6e-37
Identities = 73/73 (100%), Positives = 73/73 (100%)
Frame = +1
Query: 918 EISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESDDED 977
EISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESDDED
Sbjct: 1 EISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESDDED 180
Query: 978 TAPPARKAGAGRR 990
TAPPARKAGAGRR
Sbjct: 181 TAPPARKAGAGRR 219
>AV407003
Length = 222
Score = 138 bits (347), Expect = 5e-33
Identities = 66/68 (97%), Positives = 68/68 (99%)
Frame = +1
Query: 61 EPFAWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGEKNVGVLVVSQGWAKVREQ 120
EPFAWESREYLRKLCIGKEVTFRVDY+VASINRDFGTVFLG+KNVGVLVVSQGWAKVREQ
Sbjct: 19 EPFAWESREYLRKLCIGKEVTFRVDYSVASINRDFGTVFLGDKNVGVLVVSQGWAKVREQ 198
Query: 121 GQQKGEVS 128
GQQKGEVS
Sbjct: 199 GQQKGEVS 222
Score = 29.3 bits (64), Expect = 3.2
Identities = 17/54 (31%), Positives = 28/54 (51%)
Frame = +1
Query: 432 PYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMDFGSVFL 485
P+A E++E+LR +G++V+ ++YS S DFG+VFL
Sbjct: 22 PFAWESREYLRKLCIGKEVTFRVDYS---------------VASINRDFGTVFL 138
>TC17305 weakly similar to UP|Q9FLT0 (Q9FLT0) Transcription factor-like
protein (100 kDa coactivator-like protein), partial (5%)
Length = 616
Score = 135 bits (340), Expect = 3e-32
Identities = 66/66 (100%), Positives = 66/66 (100%)
Frame = -3
Query: 1 MASAATGATGWYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVD 60
MASAATGATGWYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVD
Sbjct: 200 MASAATGATGWYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVD 21
Query: 61 EPFAWE 66
EPFAWE
Sbjct: 20 EPFAWE 3
Score = 34.3 bits (77), Expect = 0.100
Identities = 18/57 (31%), Positives = 31/57 (53%)
Frame = -3
Query: 380 FTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKVGNPRRDEKPAPYARE 436
+ G+V V SGDC+++ + PL E+ + L+S+ P++ RR P+A E
Sbjct: 167 YRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLA--RRGGVDEPFAWE 3
>BP085388
Length = 465
Score = 130 bits (327), Expect = 1e-30
Identities = 61/63 (96%), Positives = 62/63 (97%)
Frame = -3
Query: 928 EGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESDDEDTAPPARKAGA 987
EGLARMEKRNRWDRKERKVGLDSL+KFQDDARKERRGMWQYGDVESDDED APPARKAGA
Sbjct: 463 EGLARMEKRNRWDRKERKVGLDSLEKFQDDARKERRGMWQYGDVESDDEDDAPPARKAGA 284
Query: 988 GRR 990
GRR
Sbjct: 283 GRR 275
>CB829503
Length = 575
Score = 29.6 bits (65), Expect = 2.5
Identities = 35/128 (27%), Positives = 51/128 (39%), Gaps = 3/128 (2%)
Frame = +1
Query: 842 VSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKG 901
VSAAPG + A S S D E + SEL + + + R GK KG
Sbjct: 19 VSAAPG-SDTNIAAMASSRSTCPDLEDEDVDLKSELLRYIKRAHKPDKQRRFIPYGKGKG 195
Query: 902 QGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKE 961
+G GT L + + V AA+ + ++ + D+ E V D L + A K
Sbjct: 196 KGKGTQSQNCLCHLTTDDMVRAAVEAKTPLGIKAKEAMDKGE-PVSYDLLIRIMTQALKN 372
Query: 962 RR---GMW 966
+ G W
Sbjct: 373 QSCLIGYW 396
>TC11455 similar to UP|Q9ZUS8 (Q9ZUS8) At2g37380 protein, partial (10%)
Length = 575
Score = 29.3 bits (64), Expect = 3.2
Identities = 14/39 (35%), Positives = 25/39 (63%)
Frame = -1
Query: 324 VENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMW 362
VE+ A + +A+ +EEE +RR++T E+ ++ R R W
Sbjct: 350 VESRGAVVEDEAADEVEEEERRRVRTMEMRGERWRGRSW 234
>BP075646
Length = 329
Score = 28.1 bits (61), Expect = 7.1
Identities = 11/15 (73%), Positives = 12/15 (79%)
Frame = -3
Query: 975 DEDTAPPARKAGAGR 989
DE+ PPARKAG GR
Sbjct: 324 DEEDGPPARKAGTGR 280
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 28.1 bits (61), Expect = 7.1
Identities = 22/50 (44%), Positives = 24/50 (48%), Gaps = 4/50 (8%)
Frame = +1
Query: 462 PTDGSAVPSPAADSR----VMDFGSVFLLSATKADSDDTPSSIPSAGSQP 507
PT A PSP+ S M S LSATK S TP + PSA S P
Sbjct: 280 PTSPPASPSPSPPSTKPPTAMASPSTSPLSATK--SHPTPPAAPSASSTP 423
>TC18114 similar to GB|AAM16218.1|20334714|AY093957 At2g41900/T6D20.20
{Arabidopsis thaliana;}, partial (4%)
Length = 576
Score = 27.7 bits (60), Expect = 9.3
Identities = 19/62 (30%), Positives = 31/62 (49%), Gaps = 7/62 (11%)
Frame = +2
Query: 453 EMEYSRKIVPTDGSAVPSP---AADSRVMDFGSV----FLLSATKADSDDTPSSIPSAGS 505
++E R + + GSA+PSP +A++ ++ GSV F ++ TP PS S
Sbjct: 23 KLEELRPLYVSTGSAIPSPRSYSANNSALEMGSVSPIAFGSPSSMMPPVSTPPLSPSGAS 202
Query: 506 QP 507
P
Sbjct: 203 SP 208
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.133 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,974,386
Number of Sequences: 28460
Number of extensions: 167340
Number of successful extensions: 673
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 666
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 672
length of query: 990
length of database: 4,897,600
effective HSP length: 99
effective length of query: 891
effective length of database: 2,080,060
effective search space: 1853333460
effective search space used: 1853333460
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0094a.6


