

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.9
(742 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
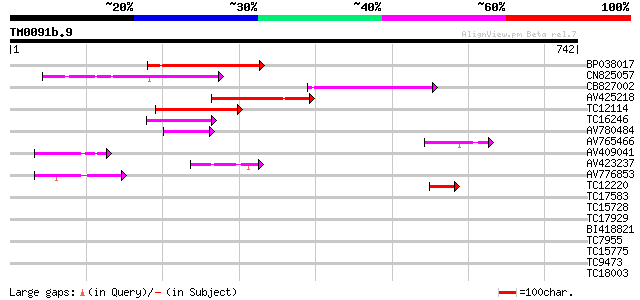
Sequences producing significant alignments: (bits) Value
BP038017 179 2e-45
CN825057 127 8e-30
CB827002 125 3e-29
AV425218 124 5e-29
TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transp... 109 1e-24
TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired re... 82 4e-16
AV780484 59 3e-09
AV765466 51 8e-07
AV409041 49 2e-06
AV423237 46 2e-05
AV776853 42 3e-04
TC12220 42 3e-04
TC17583 weakly similar to UP|Q9LIE5 (Q9LIE5) Far-red impaired re... 34 0.074
TC15728 similar to AAS09986 (AAS09986) MYB transcription factor,... 32 0.28
TC17929 31 0.63
BI418821 31 0.82
TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcrip... 31 0.82
TC15775 similar to UP|WR17_ARATH (Q9SJA8) Probable WRKY transcri... 31 0.82
TC9473 similar to AAS09986 (AAS09986) MYB transcription factor, ... 30 1.1
TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 30 1.4
>BP038017
Length = 570
Score = 179 bits (453), Expect = 2e-45
Identities = 80/153 (52%), Positives = 116/153 (75%)
Frame = +2
Query: 181 NRDCWNYIRNVRRKNLDVGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARS 240
N + Y+++ +++ L DAQ + +Y K+ Q +N F+YAIQ D+++RM+N+FW DARS
Sbjct: 113 NTTPFTYVKSSQKRTLG-RDAQNLLNYFKKMQGKNPGFYYAIQLDDENRMINVFWADARS 289
Query: 241 RLSYQLFGDVITFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKT 300
R +Y FGD + FDT Y+ N+Y +PFAPF G+N+H Q++L+GCALL DESE+SF WLF+T
Sbjct: 290 RSAYNYFGDAVIFDTMYRPNQYQVPFAPFTGVNHHGQNVLFGCALLLDESESSFTWLFRT 469
Query: 301 LLQAMGGKKPISIITDQDLAMKAALAKVFPESR 333
L AM + P+SI TDQD A++AA+A+VFPE+R
Sbjct: 470 WLSAMNDRPPVSITTDQDRAIQAAVAQVFPETR 568
>CN825057
Length = 721
Score = 127 bits (318), Expect = 8e-30
Identities = 76/242 (31%), Positives = 123/242 (50%), Gaps = 5/242 (2%)
Frame = +1
Query: 44 FYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISRTEEIQDSTNQTKRKCSTIRSG 103
FY +AK GF +++S+ + + K SR +S + R+ S ++
Sbjct: 10 FYQEYAKSMGFTTSIKNSRRSKKTKEFIDA---KFACSRYGVTPESDGGSNRRSSVKKTD 180
Query: 104 CEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHKKMSVAAKSLVEKFEEEGI 163
C+A + V R KW+I F +HNH ++ P + R H+ + +A K+ ++
Sbjct: 181 CKACMHVKR-KPDGKWIIHEFIKEHNHELL-PALAYHFRIHRNVKLAEKNNMDILHAVSE 354
Query: 164 PTGKVAAIFNDGDSTFTN-----RDCWNYIRNVRRKNLDVGDAQAVFDYCKRKQVENSNF 218
T K+ + N D + + + +D GDAQ + +Y K Q EN NF
Sbjct: 355 RTRKMYVEMSRQSGGCLNIESLVGDLNDQFKKGQYLAMDEGDAQVMLEYFKHIQKENPNF 534
Query: 219 FYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNKYSMPFAPFVGLNNHSQS 278
FY+I +E+ R+ N+FW+DA+S Y F DV++FDT+Y + +PFAPFVG+N+H Q
Sbjct: 535 FYSIDLNEEQRLRNIFWIDAKSINDYLSFNDVVSFDTSYIKSNEKLPFAPFVGVNHHCQP 714
Query: 279 IL 280
IL
Sbjct: 715 IL 720
>CB827002
Length = 534
Score = 125 bits (313), Expect = 3e-29
Identities = 65/170 (38%), Positives = 101/170 (59%)
Frame = +1
Query: 390 KENEWLQGLYKIRESWIPIYNRSTFFAGMNTTQRSESINAFFDSFVDASTTLQEFVLKFE 449
+++EWL LY P++ R TFFA M+ TQRS+S+N++FD +V+AST L +F +E
Sbjct: 28 QDHEWLS-LYSSCRQGAPVHLRDTFFAEMSITQRSDSMNSYFDGYVNASTNLNQFFKLYE 204
Query: 450 KAVESRLEAERKENYESRHKSRILSTGSKLEEHAASIYTRNIFAKFKGELEKINRFTRHK 509
KA+ESR E E + +Y++ + +L T S +E+ A+ +YTR IF +F+ EL F K
Sbjct: 205 KALESRNEKEVRADYDTMNTLPVLRTPSPMEKQASELYTRKIFMRFQEELVGTLTFMASK 384
Query: 510 IRRDGPKYVYQVSNCYDARDTFTVDIDLDSQIAKCGYELYEFMGILCRHI 559
DG Y V+ + + V ++ A C +++EF G+LCRHI
Sbjct: 385 ADDDGEVITYHVAKFGEDHKAYYVKFNVLEMKASCSCQMFEFSGLLCRHI 534
>AV425218
Length = 419
Score = 124 bits (311), Expect = 5e-29
Identities = 60/135 (44%), Positives = 84/135 (61%)
Frame = +2
Query: 265 PFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQAMGGKKPISIITDQDLAMKAA 324
P A VG+N+H QS+L+GCALL E SFVWLF++LL M G P IITD AMK A
Sbjct: 5 PLATLVGVNHHGQSVLFGCALLSSEDSESFVWLFQSLLHCMSGVPPQGIITDHSEAMKKA 184
Query: 325 LAKVFPESRHRLCLWHIIKKFPEKLAHIYHKQSIFKRDMKRCIRGSHSIQSFEEEWMRIM 384
+ V P +RHR CL +I+KK P+KL +SI + ++ + + I FE W +I+
Sbjct: 185 IETVLPSTRHRWCLSYIMKKLPQKLLGYAQYESI-RHHLQNVVYDAVVIDEFERNWKKIV 361
Query: 385 VEYNLKENEWLQGLY 399
++ L++NEWL L+
Sbjct: 362 EDFGLEDNEWLNELF 406
>TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transposase-like
{Oryza sativa (japonica cultivar-group);}, partial (8%)
Length = 488
Score = 109 bits (273), Expect = 1e-24
Identities = 48/113 (42%), Positives = 79/113 (69%)
Frame = +1
Query: 192 RRKNLDVGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVI 251
R+++L G+A + Y +RK VENS F++A Q D++ ++ N+FWVDAR + Y FGD++
Sbjct: 19 RQRSLLHGEAGYILQYFQRKLVENSPFYHAYQLDDEDQITNVFWVDARMLIDYGYFGDMV 198
Query: 252 TFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVWLFKTLLQA 304
+ D+TY T+ + P A F G N+H +++++G ALL DE+ S+ WLF++ L+A
Sbjct: 199 SLDSTYCTHSSNRPLAVFSGFNHHRKAVIFGAALLYDETTESY*WLFESFLEA 357
>TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired response
protein-like, partial (4%)
Length = 660
Score = 81.6 bits (200), Expect = 4e-16
Identities = 39/92 (42%), Positives = 51/92 (55%)
Frame = +2
Query: 179 FTNRDCWNYIRNVRRKNLDVGDAQAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDA 238
FT D N+I +R+ + GDA A Y + K + FFY D + NLFW D
Sbjct: 86 FTKTDLSNHIAKPKRERIQNGDAAAALSYLEGKADNDPMFFYKFTKTGDESLENLFWCDG 265
Query: 239 RSRLSYQLFGDVITFDTTYKTNKYSMPFAPFV 270
SR+ Y +FGDVI FD+TYK NKY+ P F+
Sbjct: 266 VSRMDYNVFGDVIAFDSTYKKNKYNKPLVVFL 361
>AV780484
Length = 529
Score = 58.9 bits (141), Expect = 3e-09
Identities = 27/67 (40%), Positives = 39/67 (57%)
Frame = -1
Query: 202 QAVFDYCKRKQVENSNFFYAIQCDEDSRMVNLFWVDARSRLSYQLFGDVITFDTTYKTNK 261
+A+ +Y Q EN NFFYAI D + + ++FWVD + RL Y+ ++ T Y NK
Sbjct: 205 KAMTEYFVSTQGENPNFFYAIDLDLNRHLTSVFWVDIKGRLDYETSMMLVLIHTHYLKNK 26
Query: 262 YSMPFAP 268
Y +PF P
Sbjct: 25 YKIPFFP 5
>AV765466
Length = 599
Score = 50.8 bits (120), Expect = 8e-07
Identities = 29/94 (30%), Positives = 49/94 (51%), Gaps = 3/94 (3%)
Frame = -2
Query: 543 KCGYELYEFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDANRG---LEDTYNDNDLGE 599
KC L+EF GILCRH L I + V ++P ++++RW K+ R ++ +Y L
Sbjct: 595 KCDCRLFEFRGILCRHSLAILSQERVKEVPDRYVLDRWRKNMRRKYVYVKTSYCVQHLKP 416
Query: 600 KSDTLKILRRVHVQREASFLADLAEESEEAYNFI 633
+ + L++L + +A A E EE +F+
Sbjct: 415 QMERLELL-----CNQFKSIAVSAAEFEETSSFV 329
>AV409041
Length = 349
Score = 49.3 bits (116), Expect = 2e-06
Identities = 31/101 (30%), Positives = 48/101 (46%)
Frame = +3
Query: 33 MEFESIEKVREFYNSFAKKNGFGVRVRSSKPKRAVLVCCNEGQHKVKISRTEEIQDSTNQ 92
+EFES E FY +AK GFG SS+ RA + ++ ++ D+ N
Sbjct: 45 IEFESHEAAYAFYKEYAKSAGFGTAKLSSRRSRASKEFIDAKFSCIRYGNKQQSDDAINP 224
Query: 93 TKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMV 133
+ + GC+AS+ V R KW + SF +HNH ++
Sbjct: 225 R----PSPKIGCKASMHVKR-RQDGKWYVYSFVKEHNHELL 332
>AV423237
Length = 310
Score = 46.2 bits (108), Expect = 2e-05
Identities = 35/100 (35%), Positives = 53/100 (53%), Gaps = 4/100 (4%)
Frame = +2
Query: 237 DARSRLSYQLFGDVITFDTTYKTNKYSMPFAPFVGLNNHSQSILYGCALLQDESEASFVW 296
DA SR++Y FGD + FDTT + S+ G ++ + +++ AL+ +ESE+SFVW
Sbjct: 17 DATSRMNYSYFGDAVIFDTTIDNHIESI--CSSWG*SSWATCVIWLVALIANESESSFVW 190
Query: 297 LFKTLLQAMGGKKPI----SIITDQDLAMKAALAKVFPES 332
L +G + SI TD D + K L VFP +
Sbjct: 191 -----LSGLGSMPCLDATCSITTDLDHSYK-LLWLVFPSN 292
>AV776853
Length = 625
Score = 42.4 bits (98), Expect = 3e-04
Identities = 34/128 (26%), Positives = 55/128 (42%), Gaps = 7/128 (5%)
Frame = +1
Query: 33 MEFESIEKVREFYNSFAKKNGFGVRVRS------SKPKRAVLVCCNEG-QHKVKISRTEE 85
++F S E+ +FY ++AK GF VR VC +G + K +RT+
Sbjct: 238 LDFGSEEEAYQFYQAYAKYQGFIVRKDDIGRDYHGNVNMRQFVCNRQGLRSKKHYNRTDR 417
Query: 86 IQDSTNQTKRKCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVMVSPKSVSYMRCHK 145
+D T + C A L V KW + SF HNH + + V ++ ++
Sbjct: 418 KRDHKPVT-------HTNCLAKLRVHLDYKIGKWKVVSFEECHNHELTPARFVHFIPPYR 576
Query: 146 KMSVAAKS 153
M+ A K+
Sbjct: 577 VMNDADKA 600
>TC12220
Length = 473
Score = 42.4 bits (98), Expect = 3e-04
Identities = 16/39 (41%), Positives = 27/39 (69%)
Frame = +1
Query: 550 EFMGILCRHILVIFQAKGVVQIPSHFIMERWTKDANRGL 588
E+ G+LCRH L +F + ++PS +I+ RWT++A G+
Sbjct: 1 EYEGVLCRHPLRVFXILELREVPSRYILHRWTRNAEDGV 117
>TC17583 weakly similar to UP|Q9LIE5 (Q9LIE5) Far-red impaired response
protein, mutator-like transposase-like protein,
phytochrome A signaling protein-like, partial (3%)
Length = 666
Score = 34.3 bits (77), Expect = 0.074
Identities = 14/29 (48%), Positives = 18/29 (61%)
Frame = +3
Query: 239 RSRLSYQLFGDVITFDTTYKTNKYSMPFA 267
R + Y+ FGDV++ DTTY TN P A
Sbjct: 3 RMIMDYEYFGDVVSLDTTYSTNNAYRPLA 89
>TC15728 similar to AAS09986 (AAS09986) MYB transcription factor, partial
(34%)
Length = 1207
Score = 32.3 bits (72), Expect = 0.28
Identities = 10/21 (47%), Positives = 16/21 (75%)
Frame = +1
Query: 715 SVVKRKACHECGEHGHNSRTC 735
+++ + C +CG +GHNSRTC
Sbjct: 178 AMMMSRTCSQCGNNGHNSRTC 240
>TC17929
Length = 791
Score = 31.2 bits (69), Expect = 0.63
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = +2
Query: 719 RKACHECGEHGHNSRTCKKRNQN 741
R+ C+ CGE GH R C K + +
Sbjct: 53 RQTCYRCGESGHKMRNCPKEHSS 121
>BI418821
Length = 614
Score = 30.8 bits (68), Expect = 0.82
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = +2
Query: 721 ACHECGEHGHNSRTCKKRNQN 741
AC+ CG+ GH +R C + N N
Sbjct: 473 ACYNCGDAGHLARDCNRSNNN 535
Score = 28.9 bits (63), Expect = 3.1
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = +2
Query: 722 CHECGEHGHNSRTCKKRNQN 741
C+ CG+ GH +R C + N N
Sbjct: 392 CYNCGDTGHLARDCHRSNNN 451
>TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcription
factor 11 (WRKY DNA-binding protein 11), partial (27%)
Length = 579
Score = 30.8 bits (68), Expect = 0.82
Identities = 14/37 (37%), Positives = 20/37 (53%)
Frame = +3
Query: 96 KCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNHVM 132
KCST+R GC A V R T +I ++ +H H +
Sbjct: 162 KCSTVR-GCPARKHVERATDDPTMLIVTYEGEHRHAL 269
>TC15775 similar to UP|WR17_ARATH (Q9SJA8) Probable WRKY transcription
factor 17 (WRKY DNA-binding protein 17), partial (53%)
Length = 1284
Score = 30.8 bits (68), Expect = 0.82
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = +3
Query: 96 KCSTIRSGCEASLIVSRGTTKSKWMIKSFNNDHNH 130
KCST+R GC A V R T +I ++ ++H+H
Sbjct: 810 KCSTVR-GCPARKHVERATDDPAMLIVTYEDEHDH 911
>TC9473 similar to AAS09986 (AAS09986) MYB transcription factor, partial
(50%)
Length = 887
Score = 30.4 bits (67), Expect = 1.1
Identities = 14/39 (35%), Positives = 20/39 (50%)
Frame = +3
Query: 700 QNSRFKSGLEVSLNKSVVKRKACHECGEHGHNSRTCKKR 738
Q+S F + N ++ +R C C GHN+RTC R
Sbjct: 135 QHSPFLPSFDPLRNAAMTRR--CSHCSHGGHNARTCPNR 245
>TC18003 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (42%)
Length = 587
Score = 30.0 bits (66), Expect = 1.4
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +1
Query: 722 CHECGEHGHNSRTCKKR 738
C+ECGE GH +R C+ R
Sbjct: 166 CYECGEPGHFARECRSR 216
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.132 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,226,360
Number of Sequences: 28460
Number of extensions: 158258
Number of successful extensions: 816
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 800
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 812
length of query: 742
length of database: 4,897,600
effective HSP length: 97
effective length of query: 645
effective length of database: 2,136,980
effective search space: 1378352100
effective search space used: 1378352100
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0091b.9


