

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
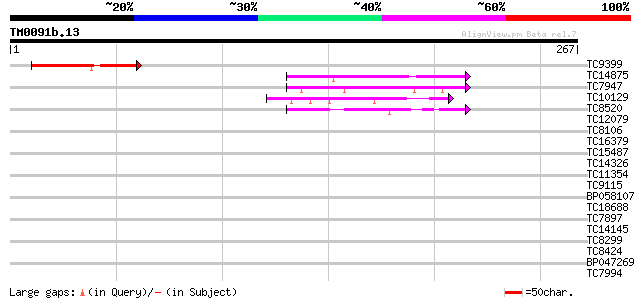
Query= TM0091b.13
(267 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC9399 similar to UP|GBF_DICDI (P36417) G-box binding factor (GB... 56 7e-09
TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%) 44 3e-05
TC7947 similar to UP|Q9ST69 (Q9ST69) Ribosomal protein S5, parti... 42 8e-05
TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%) 40 4e-04
TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and sucros... 39 7e-04
TC12079 39 0.001
TC8106 weakly similar to UP|Q9XIV1 (Q9XIV1) mRNA expressed in cu... 39 0.001
TC16379 similar to UP|O22265 (O22265) Expressed protein (At2g474... 38 0.002
TC15487 similar to UP|O23144 (O23144) Proton pump interactor (AT... 38 0.002
TC14326 weakly similar to UP|EXLP_TOBAC (Q03211) Pistil-specific... 38 0.002
TC11354 similar to UP|CAE54480 (CAE54480) Alpha glucosidase II ... 37 0.003
TC9115 similar to PIR|T00425|T00425 photolyase/blue-light recept... 37 0.003
BP058107 37 0.003
TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%) 37 0.003
TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-li... 37 0.003
TC14145 similar to UP|Q7XXS4 (Q7XXS4) Thiamine biosynthetic enzy... 37 0.003
TC8299 similar to UP|Q9XEX8 (Q9XEX8) Remorin 1, partial (67%) 37 0.005
TC8424 weakly similar to GB|BAC22265.1|24414015|AP003803 arm rep... 36 0.008
BP047269 35 0.010
TC7994 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomeras... 35 0.010
>TC9399 similar to UP|GBF_DICDI (P36417) G-box binding factor (GBF),
partial (3%)
Length = 718
Score = 55.8 bits (133), Expect = 7e-09
Identities = 27/53 (50%), Positives = 37/53 (68%), Gaps = 1/53 (1%)
Frame = +3
Query: 11 MTFTVLIHHKGFFVDKPCLHYREGCVN-VHSGIDLDTWSFFECRDLVKDLGYT 62
M+F +IHH G FV++P LHY + N VH+ + D WSFFEC D+V ++GYT
Sbjct: 504 MSFKAIIHHSGQFVEEPRLHYVDVKSNDVHA--ETDRWSFFECLDVVAEVGYT 656
>TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%)
Length = 1240
Score = 43.9 bits (102), Expect = 3e-05
Identities = 28/88 (31%), Positives = 39/88 (43%), Gaps = 1/88 (1%)
Frame = +1
Query: 131 PPFKKPSPSMPAQPKTDSPKP-PQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
PP P+ P+ PKT + P P + P P SS S AA + AP T P
Sbjct: 109 PPLPAPTI*PPSNPKTSTSTPTPASPPASTSPLTPPTTSPSSSTSTAAPSASAPLTTTP- 285
Query: 190 KKALSQHVSAPKKVTPSKRPATQPLRRS 217
+ ++P++ TPS P+T RS
Sbjct: 286 --TTATSTTSPRRQTPSSSPSTTASPRS 363
>TC7947 similar to UP|Q9ST69 (Q9ST69) Ribosomal protein S5, partial (81%)
Length = 1294
Score = 42.4 bits (98), Expect = 8e-05
Identities = 30/96 (31%), Positives = 40/96 (41%), Gaps = 9/96 (9%)
Frame = +2
Query: 131 PPFKKP-SPSMPAQPKTDSPKPPQIKG--CQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
PP P SPS P P SP+PP P P PP S P+ PP++
Sbjct: 122 PPLLSPLSPSAPPHPSPSSPQPPNSPSSLSSPDPSSPPPPPPPSPPNANPPTTLTPPSST 301
Query: 188 PM---KKALSQHVSAPKK---VTPSKRPATQPLRRS 217
P K + S H +PK + PS ++P +S
Sbjct: 302 PQTQKKTSPSTHQKSPKASSLLLPSTTAPSKPRTKS 409
>TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%)
Length = 562
Score = 40.0 bits (92), Expect = 4e-04
Identities = 32/96 (33%), Positives = 40/96 (41%), Gaps = 8/96 (8%)
Frame = +1
Query: 122 TPITMIRYNP-PFKKPSPSM-----PAQPKTDSP-KPPQIKGCQPKNVLPKADPPK-SSP 173
+PI+ Y P P K PSP+ P P T SP P PK P PP SP
Sbjct: 292 SPISQPPYTPTPPKTPSPTSQPPHTPTPPNTPSPISQPPYTPTPPKTPSPTNQPPHIPSP 471
Query: 174 SNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRP 209
N++ PPNT S PK +P+ +P
Sbjct: 472 PNSSSPTSQPPNTP----------SPPKTPSPTSQP 549
Score = 37.0 bits (84), Expect = 0.003
Identities = 28/80 (35%), Positives = 36/80 (45%), Gaps = 3/80 (3%)
Frame = +1
Query: 138 PSMPAQPKTDSPKPPQIKGCQPKN--VLPKADPPKSSPSNAAEAIRAP-PNTAPMKKALS 194
PS+P PKT SP G QP + + PK P SP N +AP PN P
Sbjct: 22 PSIPTPPKTPSP------GNQPPHTPIPPKTPSPSYSPPNVPSPPKAPSPNNHPPYTP-- 177
Query: 195 QHVSAPKKVTPSKRPATQPL 214
+ PK +P+ +P PL
Sbjct: 178 ---TPPKTPSPTSQPPYIPL 228
Score = 35.4 bits (80), Expect = 0.010
Identities = 34/95 (35%), Positives = 40/95 (41%), Gaps = 13/95 (13%)
Frame = +1
Query: 132 PFKKPSPS-----MPAQPKTDSPK--PPQI----KGCQPKNVLPKADPPKSSPSNAAEA- 179
P K PSP P PKT SP PP + K P N P P +PS ++
Sbjct: 37 PPKTPSPGNQPPHTPIPPKTPSPSYSPPNVPSPPKAPSPNNHPPYTPTPPKTPSPTSQPP 216
Query: 180 -IRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
I PP T P + HV P TPS P +QP
Sbjct: 217 YIPLPPKT-PSPISQPPHVPTPPS-TPS--PISQP 309
Score = 33.9 bits (76), Expect = 0.030
Identities = 22/58 (37%), Positives = 27/58 (45%), Gaps = 3/58 (5%)
Frame = +1
Query: 132 PFKKPSPSMPAQPKTDSP--KPPQIKGCQPKNVLPKADPPKS-SPSNAAEAIRAPPNT 186
P +P P P PKT SP +PP I P + P + PP + SP PPNT
Sbjct: 391 PISQP-PYTPTPPKTPSPTNQPPHIPS-PPNSSSPTSQPPNTPSPPKTPSPTSQPPNT 558
>TC8520 homologue to UP|Q9FZ15 (Q9FZ15) Tuber-specific and
sucrose-responsive element binding factor, partial (36%)
Length = 1778
Score = 39.3 bits (90), Expect = 7e-04
Identities = 27/88 (30%), Positives = 38/88 (42%), Gaps = 1/88 (1%)
Frame = -2
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAA-EAIRAPPNTAPM 189
PP +P+P +PAQ K++ P P P + P+ PP+ + A E R P T P
Sbjct: 844 PPSPEPAPPLPAQDKSEQPSP------APLDSSPQQSPPRGETATAK*EVRRHPHTTTPF 683
Query: 190 KKALSQHVSAPKKVTPSKRPATQPLRRS 217
QH + P R +P R S
Sbjct: 682 -----QHTAEP--APDGTRARVKPPRNS 620
>TC12079
Length = 414
Score = 38.9 bits (89), Expect = 0.001
Identities = 21/78 (26%), Positives = 33/78 (41%)
Frame = +2
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
P P+ P P + KPP G +PK+ PK+ P P A T P+ ++ Q
Sbjct: 89 PEPASPPNPSPSTTKPPSTIG-RPKSKTPKSPPSSXDPRIRPHGDLAMAPTGPIPRSSLQ 265
Query: 196 HVSAPKKVTPSKRPATQP 213
++ P + +R P
Sbjct: 266 YLXVPDXSSSXRRIHXPP 319
>TC8106 weakly similar to UP|Q9XIV1 (Q9XIV1) mRNA expressed in cucumber
hypocotyls (Arabinogalactan protein), partial (32%)
Length = 760
Score = 38.5 bits (88), Expect = 0.001
Identities = 32/114 (28%), Positives = 45/114 (39%), Gaps = 16/114 (14%)
Frame = +1
Query: 117 FGVLCTPITMIRYNPPFKK----------PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKA 166
FG TP++ +PP K P+ + PA SP P + P +P +
Sbjct: 4 FGTRVTPVSAPLASPPSKAAAAPAPVAKAPASTPPAAVPVSSPPPAPVPVASPPAPVPVS 183
Query: 167 DPPKSSPSNAAEAIRAP---PNTAPMKKALSQHVSAPKKVTPSKR---PATQPL 214
PP +P+ A A+ P AP K + KKV K+ PA PL
Sbjct: 184 SPPAPAPTTPAPAVTPTAEVPAPAPSK--------SKKKVKKGKKHGAPAPSPL 321
>TC16379 similar to UP|O22265 (O22265) Expressed protein
(At2g47450/T30B22.25), partial (16%)
Length = 449
Score = 38.1 bits (87), Expect = 0.002
Identities = 32/94 (34%), Positives = 40/94 (42%), Gaps = 15/94 (15%)
Frame = +3
Query: 137 SPSMPAQPKTDSPKPPQIKGCQ--PKNVLPKADP-PKSSPSNAAEAIRAPP--------N 185
SPS A P T P PP K + P +L K +P KS+ S+AAE PP
Sbjct: 114 SPSAAANPFTSPPSPPHSKTSRKPPMTLLKKMNPTAKSNESSAAEPFPTPPPWSISSSGK 293
Query: 186 TAPMKKA----LSQHVSAPKKVTPSKRPATQPLR 215
TA + SQ S+P P P +P R
Sbjct: 294 TATRRPGSPLISSQTTSSPSTKPPGGPPQGKPTR 395
>TC15487 similar to UP|O23144 (O23144) Proton pump interactor
(AT4g27500/F27G19_100), partial (16%)
Length = 1153
Score = 38.1 bits (87), Expect = 0.002
Identities = 32/118 (27%), Positives = 49/118 (41%), Gaps = 11/118 (9%)
Frame = +2
Query: 137 SPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQH 196
S ++P QPK + PKP P+ LP K N ++A P+ +++A
Sbjct: 236 SKAIPKQPK-EEPKP------SPQETLPTQKESK----NKVRDLKAKPDKKDLEEADEYE 382
Query: 197 VSAPKKVTPSKRPATQP-----LRRSNRL-----IWSNRKNL-EQSQVVAVQKIQANA 243
P+K TP K P P ++R + W +K L E++ A K Q A
Sbjct: 383 FEVPRKETPPKEPEIDPAKLKEMKREEEIAKAKQAWERKKKLAEKAAAKAALKAQKEA 556
>TC14326 weakly similar to UP|EXLP_TOBAC (Q03211) Pistil-specific
extensin-like protein precursor (PELP), partial (11%)
Length = 630
Score = 37.7 bits (86), Expect = 0.002
Identities = 25/85 (29%), Positives = 35/85 (40%)
Frame = +2
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
+ PP K PSP + P +P PP +K P PP +PS +++PP AP
Sbjct: 287 FPPPVKAPSPPIVKSPPVKAPTPPIVKS------PPSYPPPVKAPS--PPIVKSPPVKAP 442
Query: 189 MKKALSQHVSAPKKVTPSKRPATQP 213
V +P P + T P
Sbjct: 443 TPPI----VKSPPYYPPPVKAPTPP 505
Score = 35.4 bits (80), Expect = 0.010
Identities = 20/63 (31%), Positives = 28/63 (43%)
Frame = +2
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
Y PP K PSP + P +P PP +K P PP +P+ ++ PP P
Sbjct: 380 YPPPVKAPSPPIVKSPPVKAPTPPIVKS------PPYYPPPVKAPT--PPQVKTPPPHPP 535
Query: 189 MKK 191
+ K
Sbjct: 536 VVK 544
Score = 32.7 bits (73), Expect = 0.066
Identities = 25/93 (26%), Positives = 38/93 (39%), Gaps = 1/93 (1%)
Frame = +2
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA 182
P I +PP K P+P + P + PP +K P V PP +P+ +++
Sbjct: 308 PSPPIVKSPPVKAPTPPIVKSPPS---YPPPVKAPSPPIV---KSPPVKAPT--PPIVKS 463
Query: 183 PP-NTAPMKKALSQHVSAPKKVTPSKRPATQPL 214
PP P+K V P P +P P+
Sbjct: 464 PPYYPPPVKAPTPPQVKTPPPHPPVVKPPVAPI 562
>TC11354 similar to UP|CAE54480 (CAE54480) Alpha glucosidase II , partial
(6%)
Length = 583
Score = 37.4 bits (85), Expect = 0.003
Identities = 24/96 (25%), Positives = 36/96 (37%), Gaps = 13/96 (13%)
Frame = -1
Query: 135 KPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSP-------------SNAAEAIR 181
+P P+ P P + P P P P PP +P S+A+ + +
Sbjct: 370 EPDPAPPPPPPSSPPMSPSPTATSPPTSSPNTHPPTPTPNP*SSPSPSTTTASSASPSTK 191
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRRS 217
P+T P + S S+P PS + P R S
Sbjct: 190 TLPSTRPRPASESPTSSSPSSPPPSSGSSVSPPRIS 83
Score = 29.3 bits (64), Expect = 0.73
Identities = 14/53 (26%), Positives = 21/53 (39%)
Frame = -1
Query: 135 KPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
+P+ P SP P P + P PP +SP+ PP++A
Sbjct: 169 RPASESPTSSSPSSPPPSSGSSVSPPRISPPPPPPSNSPT------ATPPSSA 29
>TC9115 similar to PIR|T00425|T00425 photolyase/blue-light receptor (PHR2)
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (55%)
Length = 1018
Score = 37.4 bits (85), Expect = 0.003
Identities = 21/80 (26%), Positives = 34/80 (42%)
Frame = -2
Query: 142 AQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSAPK 201
A P T+ P C PK+ P PP S P ++ +P + P+ + + P
Sbjct: 867 AYPPTEKANPQHGTKCSPKSTSPPHSPPSSPPQSSPPP--SPHHAKPLCEHTQRQHQQPS 694
Query: 202 KVTPSKRPATQPLRRSNRLI 221
P++ PA Q +R L+
Sbjct: 693 PAPPTQSPACQL*QRDPTLL 634
Score = 35.0 bits (79), Expect = 0.013
Identities = 18/60 (30%), Positives = 27/60 (45%), Gaps = 1/60 (1%)
Frame = +1
Query: 128 RYNPPFKKPSPSMPAQPKTD-SPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNT 186
R NPP PSP P+ P T +P P P + P + ++ ++A + PP T
Sbjct: 316 RPNPPSNPPSPPTPSAPLTP*APTAPSTLPTPPHSAAPPSSGSATTSASATTRLSTPPTT 495
>BP058107
Length = 496
Score = 37.4 bits (85), Expect = 0.003
Identities = 22/81 (27%), Positives = 32/81 (39%)
Frame = +1
Query: 135 KPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALS 194
+P P P P+ P+ P P+ + P+A PP+ P A PP A + A
Sbjct: 112 RPCPLRPLGPRARPPRTPSPGSPSPRALPPRAPPPRGLPPRALPPRALPPAAAAPRIAEK 291
Query: 195 QHVSAPKKVTPSKRPATQPLR 215
P K KR + +R
Sbjct: 292 ATPWVPTKELRRKRSSALSVR 354
Score = 27.7 bits (60), Expect = 2.1
Identities = 23/89 (25%), Positives = 39/89 (42%)
Frame = +1
Query: 148 SPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSK 207
SP+P + + + P+A PP+ +PS + + RA P AP + L P+ + P+
Sbjct: 106 SPRPCPL-----RPLGPRARPPR-TPSPGSPSPRALPPRAPPPRGLPPRALPPRALPPA- 264
Query: 208 RPATQPLRRSNRLIWSNRKNLEQSQVVAV 236
A P W K L + + A+
Sbjct: 265 --AAAPRIAEKATPWVPTKELRRKRSSAL 345
>TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%)
Length = 514
Score = 37.0 bits (84), Expect = 0.003
Identities = 30/103 (29%), Positives = 44/103 (42%), Gaps = 11/103 (10%)
Frame = +3
Query: 122 TPITMIRYNPPFKKPS----PSMPAQPKTDSPKPPQIKGCQPKNVLP-------KADPPK 170
+P+++ + +PP PS P P + + +P PP P A PP
Sbjct: 102 SPLSLNQISPPNAPPS*LSAPPSPEERFSGTPPPPPPATGSASTATPTPPTSSRSASPPS 281
Query: 171 SSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
SP+N+ A P T+ ALS VS P +P+ P T P
Sbjct: 282 PSPANSHMASSPPSPTS----ALSVSVSTP---SPAHSPPTSP 389
>TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-like protein
2 precursor (Phytocyanin-like protein), partial (31%)
Length = 1482
Score = 37.0 bits (84), Expect = 0.003
Identities = 33/100 (33%), Positives = 42/100 (42%), Gaps = 17/100 (17%)
Frame = +3
Query: 131 PPFKKPSPSMPAQPKTDSPKP---PQIKGCQPKNVLPK-------------ADPPKSSPS 174
PP PSPS A P + SP P G P +VLP PP +S S
Sbjct: 543 PPKSSPSPSPKASPPSGSPPTAGEPSPAGTPP-SVLPSPAGAPVPAGGPSAGTPPSASTS 719
Query: 175 NAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRP-ATQP 213
+ + APP +P A S S+P +P+ P AT P
Sbjct: 720 PSPSPVSAPPAASPAPGAESPS-SSPTSNSPAGGPTATSP 836
Score = 33.1 bits (74), Expect = 0.050
Identities = 21/59 (35%), Positives = 28/59 (46%)
Frame = +3
Query: 155 KGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+G QP P + PPKSSPS + +A +PP+ +P P V PS A P
Sbjct: 516 RGTQP----PPSPPPKSSPSPSPKA--SPPSGSPPTAGEPSPAGTPPSVLPSPAGAPVP 674
>TC14145 similar to UP|Q7XXS4 (Q7XXS4) Thiamine biosynthetic enzyme, partial
(83%)
Length = 1372
Score = 37.0 bits (84), Expect = 0.003
Identities = 25/86 (29%), Positives = 36/86 (41%), Gaps = 5/86 (5%)
Frame = +3
Query: 131 PPFKKPSPSMPAQ-----PKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPN 185
PP P PS P Q P ++P PP A PP + P N + PP+
Sbjct: 132 PPQPSPPPSFPTQNPHSPPSMENPSPP-------------APPPSNPPPNKTQ----PPS 260
Query: 186 TAPMKKALSQHVSAPKKVTPSKRPAT 211
P L+ +S P + PS+ P++
Sbjct: 261 PCP*TPLLT--ISKPSNLHPSRNPSS 332
Score = 31.2 bits (69), Expect = 0.19
Identities = 28/121 (23%), Positives = 46/121 (37%), Gaps = 19/121 (15%)
Frame = +3
Query: 130 NPPFKKPSPSMPAQPKTDSPKP----PQIKGCQPKNVLPKADPPK--------------- 170
NP P PS P KT P P P + +P N+ P +P
Sbjct: 198 NPSPPAPPPSNPPPNKTQPPSPCP*TPLLTISKPSNLHPSRNPSSHVK*PAAT*LT*SLT 377
Query: 171 SSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRRSNRLIWSNRKNLEQ 230
+P++++ A+ P + AP A + ++P + S A + S+ W + L
Sbjct: 378 PTPTSSSSALDPPASHAPTSSARTLPSASPSLSSLSAPAAARGSVASSSPPWWSVSRLTS 557
Query: 231 S 231
S
Sbjct: 558 S 560
>TC8299 similar to UP|Q9XEX8 (Q9XEX8) Remorin 1, partial (67%)
Length = 1098
Score = 36.6 bits (83), Expect = 0.005
Identities = 25/116 (21%), Positives = 51/116 (43%), Gaps = 11/116 (9%)
Frame = +2
Query: 131 PPFKKPSPSMPAQPKTDSPKP-----PQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPN 185
PP P+ S PA P+P P+ + K+++P+ PP+S P + ++AI
Sbjct: 200 PPAPAPTESTPAPEAEAKPEPVVHDAPKEDVAEEKSIIPQPSPPESKPVDDSKAIVKVEK 379
Query: 186 T------APMKKALSQHVSAPKKVTPSKRPATQPLRRSNRLIWSNRKNLEQSQVVA 235
T P++ ++++ + T + + S + I N+ + + S + A
Sbjct: 380 TQEAAEEKPLEGSINRDAVLTRVATEKRLSLIKAWEESEKSIADNKAHKKLSDISA 547
>TC8424 weakly similar to GB|BAC22265.1|24414015|AP003803 arm repeat
containing protein homolog-like {Oryza sativa (japonica
cultivar-group);} , partial (49%)
Length = 1586
Score = 35.8 bits (81), Expect = 0.008
Identities = 26/89 (29%), Positives = 40/89 (44%), Gaps = 2/89 (2%)
Frame = -3
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+P +P PS A P SP PP +++ P DPP +SP + + +
Sbjct: 834 HPSHPEPQPSC*APP---SPSPPD------QSLAPPQDPPAASPLQRSPPSPCSASQSTA 682
Query: 190 KKALSQHVSAP--KKVTPSKRPATQPLRR 216
ALS H S+P ++ +K A P +R
Sbjct: 681 CTALSAHPSSPYWNHLSAAKPAAVSPRKR 595
>BP047269
Length = 619
Score = 35.4 bits (80), Expect = 0.010
Identities = 29/93 (31%), Positives = 41/93 (43%), Gaps = 10/93 (10%)
Frame = +1
Query: 137 SPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPN---TAPMKKAL 193
S S P+Q +T + +P P PP SSP + A + + PN +AP+ + L
Sbjct: 85 SESQPSQSRTSVATTTVVGAGKPPLKTPSRLPPPSSPPSTAASPNSSPNWRVSAPIAREL 264
Query: 194 SQHVSAPK--KVTPSKRP-----ATQPLRRSNR 219
+ S + K TP P AT LRR R
Sbjct: 265 ATRSSRKQAHKTTPFAEPSSSTAATMTLRRRCR 363
>TC7994 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomerase
precursor , partial (81%)
Length = 1355
Score = 35.4 bits (80), Expect = 0.010
Identities = 27/102 (26%), Positives = 43/102 (41%), Gaps = 9/102 (8%)
Frame = +2
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKA---DPPKSSPSNAAE 178
+P T R +PP + + S PA +P PP P + P + PP +SP+++A
Sbjct: 395 SPKTTSRDSPPTRPSTTSSPAWSSASAPAPP------PPSSSPSSASFSPPANSPTSSA- 553
Query: 179 AIRAPPNTA------PMKKALSQHVSAPKKVTPSKRPATQPL 214
+PP +A P + P ++PS P L
Sbjct: 554 ---SPPPSARRSRRDPSASRSPSSTTIPASISPSTAPTRSTL 670
Score = 30.4 bits (67), Expect = 0.33
Identities = 27/92 (29%), Positives = 37/92 (39%), Gaps = 6/92 (6%)
Frame = +2
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKN------VLPKADPPKSSPSNA 176
P+ + PP PS P+ PKT S P + + P PP SSPS+A
Sbjct: 338 PLPLNYAPPPLPNPSVPSPS-PKTTSRDSPPTRPSTTSSPAWSSASAPAPPPPSSSPSSA 514
Query: 177 AEAIRAPPNTAPMKKALSQHVSAPKKVTPSKR 208
+ +PP +P A S P S+R
Sbjct: 515 S---FSPPANSPTSSA-----SPPPSARRSRR 586
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.133 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,248,786
Number of Sequences: 28460
Number of extensions: 95016
Number of successful extensions: 1784
Number of sequences better than 10.0: 372
Number of HSP's better than 10.0 without gapping: 1247
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1536
length of query: 267
length of database: 4,897,600
effective HSP length: 89
effective length of query: 178
effective length of database: 2,364,660
effective search space: 420909480
effective search space used: 420909480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0091b.13


