

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.10
(353 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
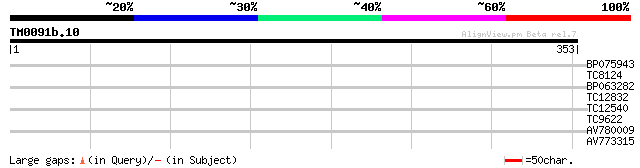
Score E
Sequences producing significant alignments: (bits) Value
BP075943 39 0.001
TC8124 weakly similar to UP|IF5Z_ARATH (Q9S825) Probable eukaryo... 32 0.21
BP063282 32 0.21
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 31 0.27
TC12540 29 1.4
TC9622 weakly similar to UP|AAQ54501 (AAQ54501) Glucosyltransfer... 29 1.4
AV780009 28 1.8
AV773315 28 3.0
>BP075943
Length = 547
Score = 38.9 bits (89), Expect = 0.001
Identities = 29/75 (38%), Positives = 41/75 (54%), Gaps = 5/75 (6%)
Frame = -3
Query: 131 YLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNGKQVF-RYLKGALDMSLFFPF 189
Y + LLYL + TR DI+FA+N L+++ S+PT H F +YL G+ LF+P
Sbjct: 224 YRRIVGRLLYLNT-TRPDITFAVNQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGLFYPA 48
Query: 190 MSKL----NLIGYAD 200
S +LIG D
Sbjct: 47 SSSTL*LPSLIGGPD 3
>TC8124 weakly similar to UP|IF5Z_ARATH (Q9S825) Probable eukaryotic
translation initiation factor 5-2 (eIF-5 2), partial
(5%)
Length = 884
Score = 31.6 bits (70), Expect = 0.21
Identities = 14/21 (66%), Positives = 17/21 (80%)
Frame = +1
Query: 99 LCTLLVLRSLNVNKDPFRP*E 119
LCT +V+RSLNVN+ PF P E
Sbjct: 718 LCTSIVVRSLNVNQGPFIPQE 780
>BP063282
Length = 519
Score = 31.6 bits (70), Expect = 0.21
Identities = 14/21 (66%), Positives = 17/21 (80%)
Frame = -1
Query: 99 LCTLLVLRSLNVNKDPFRP*E 119
LCT +V+RSLNVN+ PF P E
Sbjct: 177 LCTSIVVRSLNVNQGPFIPQE 115
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 31.2 bits (69), Expect = 0.27
Identities = 21/66 (31%), Positives = 35/66 (52%), Gaps = 1/66 (1%)
Frame = +2
Query: 76 VFMYQRTYIAKVLKGFYMDKSCPLCTLLVLR-SLNVNKDPFRP*EKNEELLDPEVPYLSA 134
+++ Q+ Y+ KVL+ F M P+ T L + L+ + P E+ E VPY SA
Sbjct: 254 IWLSQKNYLQKVLRRFNMQDCNPISTPLPVNYKLSSSMIPSSEAERMEM---SRVPYASA 424
Query: 135 IKALLY 140
+ +L+Y
Sbjct: 425 VGSLMY 442
>TC12540
Length = 538
Score = 28.9 bits (63), Expect = 1.4
Identities = 15/45 (33%), Positives = 22/45 (48%)
Frame = -1
Query: 130 PYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNGKQVF 174
PY I Y SH+ LD+S ++ A +S HWN Q++
Sbjct: 442 PYFLYIFFFFYHISHSLLDVSSRVHSFANLASV*VM-HWNHNQIY 311
>TC9622 weakly similar to UP|AAQ54501 (AAQ54501) Glucosyltransferase
(Fragment), partial (84%)
Length = 584
Score = 28.9 bits (63), Expect = 1.4
Identities = 14/41 (34%), Positives = 25/41 (60%)
Frame = -2
Query: 72 LKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNK 112
L + V +Y+R + ++LKG+ M S PL +++ + N NK
Sbjct: 562 LLNGVIIYRR*FYIELLKGYMMKNSMPLIA*ILIDNNNNNK 440
>AV780009
Length = 529
Score = 28.5 bits (62), Expect = 1.8
Identities = 14/32 (43%), Positives = 19/32 (58%)
Frame = -1
Query: 172 QVFRYLKGALDMSLFFPFMSKLNLIGYADAGY 203
+V RY+KGA LFF S L L Y+D+ +
Sbjct: 517 RVLRYVKGAPAQGLFFSADSPLKLQAYSDSDW 422
>AV773315
Length = 519
Score = 27.7 bits (60), Expect = 3.0
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = +3
Query: 281 LHALLNQKKDTSWEMDKANITG 302
LH +N K T W++ K NITG
Sbjct: 12 LHT*MNHHKVTKWKISKMNITG 77
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.357 0.160 0.555
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,661,958
Number of Sequences: 28460
Number of extensions: 69891
Number of successful extensions: 774
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 771
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 774
length of query: 353
length of database: 4,897,600
effective HSP length: 91
effective length of query: 262
effective length of database: 2,307,740
effective search space: 604627880
effective search space used: 604627880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0091b.10


