

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091a.6
(242 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
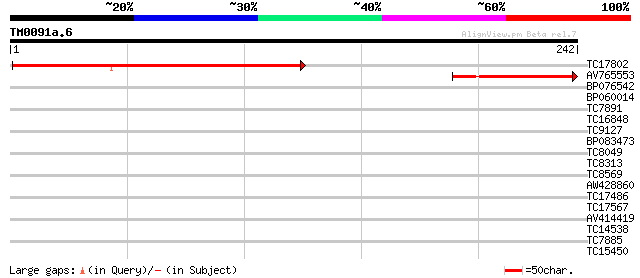
Score E
Sequences producing significant alignments: (bits) Value
TC17802 similar to UP|Q9XII2 (Q9XII2) At2g16050 protein, partial... 249 4e-67
AV765553 83 5e-17
BP076542 31 0.17
BP060014 28 1.9
TC7891 similar to UP|DCAM_VICFA (Q9M4D8) S-adenosylmethionine de... 27 2.4
TC16848 homologue to UP|BAD08451 (BAD08451) Glucose-6-phosphate ... 27 3.2
TC9127 weakly similar to UP|LYM2_ARATH (O23006) LysM-domain GPI-... 27 4.1
BP083473 27 4.1
TC8049 similar to UP|Q8LPQ5 (Q8LPQ5) AT3g18290/MIE15_8, partial ... 26 5.4
TC8313 26 5.4
TC8569 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal protein... 26 7.0
AW428860 26 7.0
TC17486 similar to UP|RU17_ARATH (Q42404) U1 small nuclear ribon... 26 7.0
TC17567 similar to UP|Q8RXD3 (Q8RXD3) ABI3-interacting protein 2... 26 7.0
AV414419 25 9.2
TC14538 similar to GB|AAH54382.1|32450414|BC054382 feminization ... 25 9.2
TC7885 similar to UP|RL4B_ARATH (Q9SF40) 60S ribosomal protein L... 25 9.2
TC15450 similar to GB|AAD04752.1|4098989|ATU81498 phenylalkylami... 25 9.2
>TC17802 similar to UP|Q9XII2 (Q9XII2) At2g16050 protein, partial (72%)
Length = 571
Score = 249 bits (635), Expect = 4e-67
Identities = 108/126 (85%), Positives = 116/126 (91%), Gaps = 1/126 (0%)
Frame = +2
Query: 2 KYSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCS-ICDYDLHMQCAIPSPTLV 60
K SEISHFSHP+HKLRFE++ FPFKCDGCKE+GIGSRYKCS ICDYDLHM CAIPSP+L
Sbjct: 194 KCSEISHFSHPKHKLRFEYSEFPFKCDGCKEVGIGSRYKCSSICDYDLHMHCAIPSPSLY 373
Query: 61 HPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEV 120
HPFY KCSFQF+S PPGNIPRYCNACEK V GFVYHC +CGFDLHPCCAKLPMVLDDGE+
Sbjct: 374 HPFYPKCSFQFLSKPPGNIPRYCNACEKGVTGFVYHCMTCGFDLHPCCAKLPMVLDDGEI 553
Query: 121 KLYLYR 126
KLYLYR
Sbjct: 554 KLYLYR 571
>AV765553
Length = 354
Score = 82.8 bits (203), Expect = 5e-17
Identities = 41/53 (77%), Positives = 49/53 (92%)
Frame = -1
Query: 190 NTLYAAHNSRRSKGKVKKCCEIAGLAVQFVISAVLGDPTALIAGVVGSLMSRA 242
NTL A+HNSR KGKV+KCCEIAG+AVQ V+SAVLG+PTALIAG++GS+MSRA
Sbjct: 354 NTLQASHNSR-IKGKVRKCCEIAGMAVQVVVSAVLGNPTALIAGIMGSVMSRA 199
>BP076542
Length = 376
Score = 31.2 bits (69), Expect = 0.17
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = -1
Query: 95 YHCKSCGFDLHPCCAKL 111
Y+C+ C FDLHP CA +
Sbjct: 367 YYCEECDFDLHPKCASV 317
Score = 27.3 bits (59), Expect = 2.4
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = -1
Query: 39 YKCSICDYDLHMQCA 53
Y C CD+DLH +CA
Sbjct: 367 YYCEECDFDLHPKCA 323
>BP060014
Length = 366
Score = 27.7 bits (60), Expect = 1.9
Identities = 19/69 (27%), Positives = 25/69 (35%), Gaps = 5/69 (7%)
Frame = +3
Query: 104 LHPCCAK-LPMVLDDGE----VKLYLYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV 158
+H CC P V E +K + R C CG KG + K HV C
Sbjct: 147 VHRCCVDWAPQVYFVDETCKNLKAEVARGAKLKCSTCGLKGAALGCYVKSCKRTYHVPCA 326
Query: 159 REMVVENWD 167
++ WD
Sbjct: 327 MDVSTCRWD 353
>TC7891 similar to UP|DCAM_VICFA (Q9M4D8) S-adenosylmethionine
decarboxylase proenzyme (AdoMetDC) (SamDC) [Contains:
S-adenosylmethionine decarboxylase alpha chain;
S-adenosylmethionine decarboxylase beta chain] ,
complete
Length = 1889
Score = 27.3 bits (59), Expect = 2.4
Identities = 20/78 (25%), Positives = 35/78 (44%)
Frame = -3
Query: 133 HRCGRKGRSWSYRSKCKSYNLHVACVREMVVENWDHVYVAGGKGRNKVVETSVPGLKNTL 192
H C RK SWS + +CK + + +M+ E + + G K K K+ L
Sbjct: 546 HGCDRKECSWSKKDRCKESTIAKRGIGKMLFEGE**LILEGNKNG*K---------KHWL 394
Query: 193 YAAHNSRRSKGKVKKCCE 210
+A + +S +++ CE
Sbjct: 393 HA--RNYQSVSRIRMVCE 346
>TC16848 homologue to UP|BAD08451 (BAD08451) Glucose-6-phosphate isomerase,
partial (7%)
Length = 723
Score = 26.9 bits (58), Expect = 3.2
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = -1
Query: 60 VHPFYTKCSFQFMSSPP 76
+H TKCSFQF +S P
Sbjct: 345 LHDILTKCSFQFQNSTP 295
>TC9127 weakly similar to UP|LYM2_ARATH (O23006) LysM-domain GPI-anchored
protein 2 precursor, partial (46%)
Length = 1016
Score = 26.6 bits (57), Expect = 4.1
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = -2
Query: 2 KYSEISHFSHPQHKLRFEHNA 22
K S ++HFSH +KL EH A
Sbjct: 1015 KGSPVNHFSHSMNKLGAEHRA 953
>BP083473
Length = 214
Score = 26.6 bits (57), Expect = 4.1
Identities = 10/28 (35%), Positives = 17/28 (60%)
Frame = -3
Query: 181 VETSVPGLKNTLYAAHNSRRSKGKVKKC 208
++ S+P +KN++YA R KV+ C
Sbjct: 134 IDPSLPDIKNSIYATDEFRMFSFKVRPC 51
>TC8049 similar to UP|Q8LPQ5 (Q8LPQ5) AT3g18290/MIE15_8, partial (8%)
Length = 639
Score = 26.2 bits (56), Expect = 5.4
Identities = 14/29 (48%), Positives = 14/29 (48%), Gaps = 5/29 (17%)
Frame = +2
Query: 132 CHRCGRKGRS---WSYR--SKCKSYNLHV 155
CH C RKG S W Y C SYN V
Sbjct: 218 CHDCNRKGTSRFHWLYHKCGFCGSYNTRV 304
>TC8313
Length = 1087
Score = 26.2 bits (56), Expect = 5.4
Identities = 13/36 (36%), Positives = 18/36 (49%)
Frame = +1
Query: 49 HMQCAIPSPTLVHPFYTKCSFQFMSSPPGNIPRYCN 84
+M C P TLV C+FQF+S P + + N
Sbjct: 907 YMYCV*PGRTLVCLLAALCTFQFVSLNPMCVELFVN 1014
>TC8569 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal protein S14,
complete
Length = 633
Score = 25.8 bits (55), Expect = 7.0
Identities = 13/39 (33%), Positives = 16/39 (40%)
Frame = +3
Query: 130 SPCHRCGRKGRSWSYRSKCKSYNLHVACVREMVVENWDH 168
+PCH + SWS+ S C C ENW H
Sbjct: 351 APCHWWKQDKNSWSWCSGCS------PCPCSFRNENWSH 449
>AW428860
Length = 347
Score = 25.8 bits (55), Expect = 7.0
Identities = 11/33 (33%), Positives = 18/33 (54%), Gaps = 1/33 (3%)
Frame = -2
Query: 127 KVSSPCHRC-GRKGRSWSYRSKCKSYNLHVACV 158
+V +P H C G + + W+ R+ HVAC+
Sbjct: 214 EVQAPTHGCSGSESKDWAVRTAS*Q*RCHVACL 116
>TC17486 similar to UP|RU17_ARATH (Q42404) U1 small nuclear
ribonucleoprotein 70 kDa (U1 snRNP 70 kDa) (snRNP70)
(U1-70K), partial (39%)
Length = 595
Score = 25.8 bits (55), Expect = 7.0
Identities = 12/29 (41%), Positives = 13/29 (44%)
Frame = -1
Query: 27 CDGCKEIGIGSRYKCSICDYDLHMQCAIP 55
C G GIG Y S +L CAIP
Sbjct: 373 CAGVSATGIGGEYSASPGSANLPTNCAIP 287
>TC17567 similar to UP|Q8RXD3 (Q8RXD3) ABI3-interacting protein 2, partial
(25%)
Length = 511
Score = 25.8 bits (55), Expect = 7.0
Identities = 8/28 (28%), Positives = 15/28 (53%)
Frame = +2
Query: 139 GRSWSYRSKCKSYNLHVACVREMVVENW 166
G++W +KCKSY+ + ++ W
Sbjct: 11 GKTWL*MTKCKSYHASTRSIHHVLSRGW 94
>AV414419
Length = 396
Score = 25.4 bits (54), Expect = 9.2
Identities = 11/19 (57%), Positives = 11/19 (57%)
Frame = -2
Query: 102 FDLHPCCAKLPMVLDDGEV 120
FDLHP C KL V EV
Sbjct: 227 FDLHPACPKLQPVRSGDEV 171
>TC14538 similar to GB|AAH54382.1|32450414|BC054382 feminization 1 homolog a
{Mus musculus;}, partial (3%)
Length = 2631
Score = 25.4 bits (54), Expect = 9.2
Identities = 12/39 (30%), Positives = 19/39 (47%), Gaps = 1/39 (2%)
Frame = +2
Query: 9 FSHP-QHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDY 46
F HP ++ R + +P+ C C E G+ K C+Y
Sbjct: 839 FVHPGENARRRDPRKYPYSCVPCPEFRKGACQKGDSCEY 955
>TC7885 similar to UP|RL4B_ARATH (Q9SF40) 60S ribosomal protein L4-2 (L1),
complete
Length = 1554
Score = 25.4 bits (54), Expect = 9.2
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +2
Query: 105 HPCCAKLPMVLDDGEVKLYLYRKVSSPCHRCGRKGR 140
+P C+ L ++L L+ + SSPCHR GR
Sbjct: 35 NPSCSPLTLLL-----LLHFNGRSSSPCHRASPGGR 127
>TC15450 similar to GB|AAD04752.1|4098989|ATU81498 phenylalkylamine binding
protein homolog {Arabidopsis thaliana;} , partial (70%)
Length = 1088
Score = 25.4 bits (54), Expect = 9.2
Identities = 26/104 (25%), Positives = 43/104 (41%), Gaps = 10/104 (9%)
Frame = -2
Query: 65 TKCSFQFMSSPPGNIPRYCNACE----------KDVNGFVYHCKSCGFDLHPCCAKLPMV 114
+KCS + G+ Y N E K V G + + CG+ +H LP
Sbjct: 694 SKCSRSLPPAADGDDRGYHNPGEIGSNIVCIIVK*VGGKIITFQDCGYVIHSSSI*LPQ- 518
Query: 115 LDDGEVKLYLYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV 158
+G++K SS C + SW+ + C S+N + +C+
Sbjct: 517 -RNGKLKHVAI*LSSSYSVYC*KA--SWTLYNCC*SFNSNYSCI 395
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.138 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,112,864
Number of Sequences: 28460
Number of extensions: 118648
Number of successful extensions: 793
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 785
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 790
length of query: 242
length of database: 4,897,600
effective HSP length: 88
effective length of query: 154
effective length of database: 2,393,120
effective search space: 368540480
effective search space used: 368540480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0091a.6


