

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
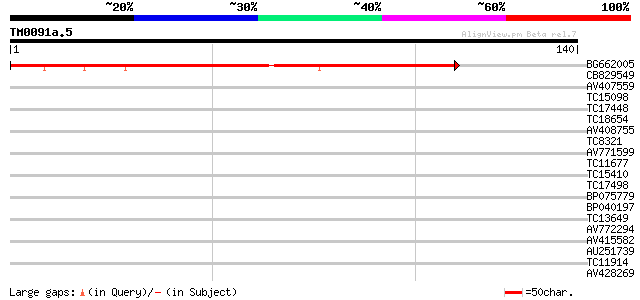
Query= TM0091a.5
(140 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG662005 81 8e-17
CB829549 31 0.091
AV407559 29 0.26
TC15098 similar to UP|Q8VYV7 (Q8VYV7) AT5g66120/K2A18_20, partia... 28 0.59
TC17448 homologue to UP|O23628 (O23628) Histone H2A.F/Z (At3g545... 27 1.0
TC18654 homologue to UP|Q8GZU9 (Q8GZU9) Ribonucleotide reductase... 27 1.7
AV408755 26 2.2
TC8321 similar to UP|AAS46230 (AAS46230) Peroxiredoxin Q, partia... 26 2.2
AV771599 26 2.9
TC11677 weakly similar to UP|Q87R67 (Q87R67) Naphthoate synthase... 26 2.9
TC15410 25 3.8
TC17498 similar to GB|AAO44081.1|28466945|BT004815 At3g57000 {Ar... 25 3.8
BP075779 25 3.8
BP040197 25 5.0
TC13649 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, ... 25 5.0
AV772294 25 5.0
AV415582 25 5.0
AU251739 25 6.5
TC11914 homologue to UP|Q7PDW6 (Q7PDW6) ENSANGP00000024926 (Frag... 25 6.5
AV428269 25 6.5
>BG662005
Length = 458
Score = 80.9 bits (198), Expect = 8e-17
Identities = 54/126 (42%), Positives = 80/126 (62%), Gaps = 15/126 (11%)
Frame = +2
Query: 1 MGICAST-----QYAGKGGNHG--------MQSTVNIVHF-DGKLQQLKEPVKAWHVLSQ 46
MG+CAS + +GKGG G S+V +++ G +++ K+PV A V+S+
Sbjct: 77 MGVCASKTNEKPRASGKGGRGGGGAAESFRRPSSVMVMNLAGGGIKEYKQPVPARVVVSE 256
Query: 47 NPNHFLCSSESMYVGSPMTPVVPNEELQL-NHIYFLVPRSKSRLPLSLEDLCALAIKANA 105
NP+++LC+SESM VG+ M P VP+EE+ L IYFLVP S S LSL LC LA+KA++
Sbjct: 257 NPDYYLCNSESMCVGTCM-PRVPDEEVLLPGRIYFLVPLSHSHHQLSLPLLCDLAVKASS 433
Query: 106 ALAHSS 111
L +++
Sbjct: 434 VLDNAN 451
>CB829549
Length = 534
Score = 30.8 bits (68), Expect = 0.091
Identities = 16/46 (34%), Positives = 26/46 (55%), Gaps = 1/46 (2%)
Frame = +2
Query: 46 QNPNHFLCSSESM-YVGSPMTPVVPNEELQLNHIYFLVPRSKSRLP 90
+ P H L S+++ + G P+ PN+EL+ IYF+V K + P
Sbjct: 122 KTPGHVLLDSQAVKHFGLRARPLEPNQELKPKKIYFVVELPKVQEP 259
>AV407559
Length = 429
Score = 29.3 bits (64), Expect = 0.26
Identities = 23/86 (26%), Positives = 38/86 (43%)
Frame = +3
Query: 44 LSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKA 103
LS +P H S + + +P P E + F +S+ P C+LA
Sbjct: 24 LSLSPTHSDRHLFSAQIAAVNSPATPRHE*MNDIFIFNQSQSQCSSP------CSLA*SP 185
Query: 104 NAALAHSSSVFEPSSSVPQINFKTHP 129
N ++ +SS+F P++ + Q THP
Sbjct: 186 NPLISVASSLFHPNNPISQ----THP 251
>TC15098 similar to UP|Q8VYV7 (Q8VYV7) AT5g66120/K2A18_20, partial (66%)
Length = 1272
Score = 28.1 bits (61), Expect = 0.59
Identities = 14/40 (35%), Positives = 21/40 (52%), Gaps = 7/40 (17%)
Frame = -2
Query: 97 CALAIKANAALAHSSSVF-------EPSSSVPQINFKTHP 129
C L + A AHS +F +PSS +P ++F +HP
Sbjct: 602 CQLQQPLHEATAHSQLLFLLHDQMYDPSSMLPSVHFPSHP 483
>TC17448 homologue to UP|O23628 (O23628) Histone H2A.F/Z (At3g54560),
partial (88%)
Length = 597
Score = 27.3 bits (59), Expect = 1.0
Identities = 16/71 (22%), Positives = 30/71 (41%)
Frame = -1
Query: 67 VVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSSSVPQINFK 126
+ P+ +LQ+ Y L + +R+P ++LC+ +K + SS P P
Sbjct: 267 ISPDSKLQMPRSYPLHLQILARIPSQFKNLCSQVLKNGGQVHCSSCTNTPIRLNPLFQLS 88
Query: 127 THPVHSNVSLG 137
H ++ G
Sbjct: 87 MDTTHRKLNSG 55
>TC18654 homologue to UP|Q8GZU9 (Q8GZU9) Ribonucleotide reductase large
subunit A, partial (31%)
Length = 753
Score = 26.6 bits (57), Expect = 1.7
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = +1
Query: 27 FDGKLQQLKEPVKAWH 42
F G LQQ K+P K+WH
Sbjct: 235 F*GHLQQEKQPAKSWH 282
>AV408755
Length = 351
Score = 26.2 bits (56), Expect = 2.2
Identities = 20/52 (38%), Positives = 27/52 (51%), Gaps = 1/52 (1%)
Frame = +1
Query: 43 VLSQN-PNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSL 93
++S N P H L SS + P +P ELQL L R+++RLP SL
Sbjct: 67 IISPNSPLHLLSSSSATMFTLNAPPQLP--ELQLTSPSELTWRNRTRLPASL 216
>TC8321 similar to UP|AAS46230 (AAS46230) Peroxiredoxin Q, partial (75%)
Length = 873
Score = 26.2 bits (56), Expect = 2.2
Identities = 16/44 (36%), Positives = 23/44 (51%), Gaps = 4/44 (9%)
Frame = +1
Query: 80 FLVPRSKSRLPLSLEDLCA----LAIKANAALAHSSSVFEPSSS 119
FL P S S LP + + +K +++ +HSSS PSSS
Sbjct: 49 FLHPPSSSNLPFPFPQSSSNSQFIGLKLSSSSSHSSSFSVPSSS 180
>AV771599
Length = 535
Score = 25.8 bits (55), Expect = 2.9
Identities = 30/124 (24%), Positives = 49/124 (39%), Gaps = 6/124 (4%)
Frame = +3
Query: 16 HGMQSTVNIVHFDGKLQQLKEPVKA-WHVLSQNPNHFLCSSESMYVGSPMT-----PVVP 69
HG T+ I H+ K K+ KA W +L H E++Y + +
Sbjct: 72 HGENHTLGI-HYPKKESSQKKKTKATWSILKCTQTH---EQEAVYSERGLNINNCIMIYS 239
Query: 70 NEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSSSVPQINFKTHP 129
N E + + + P + + +++ LA + + SVF PSSS +K
Sbjct: 240 NSESHKSTVLLIYPETVAVDLVTIRHGTPLA----SMVPSGFSVFPPSSSTRDPVYKPSN 407
Query: 130 VHSN 133
HSN
Sbjct: 408 KHSN 419
>TC11677 weakly similar to UP|Q87R67 (Q87R67) Naphthoate synthase, partial
(41%)
Length = 499
Score = 25.8 bits (55), Expect = 2.9
Identities = 16/53 (30%), Positives = 26/53 (48%)
Frame = -2
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLC 53
MG +S +Y G + ++S++ I+H LK P ++W V P F C
Sbjct: 297 MGFLSSIKYQGCIPTNFLKSSMPIIHCIKS*--LKNPNRSWAVPEYLPTPFYC 145
>TC15410
Length = 610
Score = 25.4 bits (54), Expect = 3.8
Identities = 13/54 (24%), Positives = 20/54 (36%)
Frame = +3
Query: 83 PRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSSSVPQINFKTHPVHSNVSL 136
P S SLE L L + + + P +P IN +HS + +
Sbjct: 174 PSDNSEFVNSLEKLLQLCQELGKQIDEIGACLYPPQEIPSINAALEKIHSTIDV 335
>TC17498 similar to GB|AAO44081.1|28466945|BT004815 At3g57000 {Arabidopsis
thaliana;}, partial (28%)
Length = 589
Score = 25.4 bits (54), Expect = 3.8
Identities = 13/39 (33%), Positives = 23/39 (58%)
Frame = +2
Query: 102 KANAALAHSSSVFEPSSSVPQINFKTHPVHSNVSLGYSH 140
+A L+ SSS +P+ ++ Q+N KT + +V + SH
Sbjct: 20 RALVVLSSSSSEIQPTINLAQLNPKTRQLSCSVVVEGSH 136
>BP075779
Length = 476
Score = 25.4 bits (54), Expect = 3.8
Identities = 9/27 (33%), Positives = 19/27 (70%)
Frame = +1
Query: 52 LCSSESMYVGSPMTPVVPNEELQLNHI 78
LC+ + ++ S +P++P+ +LQ NH+
Sbjct: 271 LCNLKLLFCSS*FSPLLPHFKLQSNHL 351
>BP040197
Length = 466
Score = 25.0 bits (53), Expect = 5.0
Identities = 13/37 (35%), Positives = 19/37 (51%), Gaps = 1/37 (2%)
Frame = +2
Query: 22 VNIVHFDGKLQQLKEPVKAWHVLSQNPNHF-LCSSES 57
V++ HF + Q P+ W V Q P H+ LC + S
Sbjct: 236 VSVTHFRTFMDQ*CFPIL*WPVC*QEPEHYLLCKTHS 346
>TC13649 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, partial
(68%)
Length = 690
Score = 25.0 bits (53), Expect = 5.0
Identities = 16/56 (28%), Positives = 25/56 (44%)
Frame = -1
Query: 81 LVPRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSSSVPQINFKTHPVHSNVSL 136
+V +KSR +S+ + S+ FEPSSS I H + +V+L
Sbjct: 558 IVEHTKSRTHISVHIPTTRRVLILFFFTQSTKTFEPSSSAEPIYPSKHSLRLSVTL 391
>AV772294
Length = 494
Score = 25.0 bits (53), Expect = 5.0
Identities = 17/47 (36%), Positives = 21/47 (44%), Gaps = 1/47 (2%)
Frame = -2
Query: 46 QNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSR-LPL 91
Q PN F CS + + P+ P+ L L L RS S LPL
Sbjct: 460 QQPNMFFCSRSV*RIPLTLPPICPSSSLNLMGRNILGRRSCSESLPL 320
>AV415582
Length = 424
Score = 25.0 bits (53), Expect = 5.0
Identities = 10/31 (32%), Positives = 16/31 (51%)
Frame = -1
Query: 30 KLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
+L L+ + WH P H LC S+ +Y+
Sbjct: 172 QLLLLETAISQWHDGLLFPRHHLCQSKLLYI 80
>AU251739
Length = 338
Score = 24.6 bits (52), Expect = 6.5
Identities = 10/33 (30%), Positives = 17/33 (51%)
Frame = -3
Query: 10 AGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWH 42
+G G +HG+ S+ +HFD + P + H
Sbjct: 162 SGPGFSHGIASSTKTLHFDSYVTLHMHPGRPLH 64
>TC11914 homologue to UP|Q7PDW6 (Q7PDW6) ENSANGP00000024926 (Fragment),
partial (6%)
Length = 599
Score = 24.6 bits (52), Expect = 6.5
Identities = 12/31 (38%), Positives = 19/31 (60%), Gaps = 3/31 (9%)
Frame = -2
Query: 62 SPMTPVVPN---EELQLNHIYFLVPRSKSRL 89
S + +VPN +E Q+NH + +P KSR+
Sbjct: 223 STLQGLVPN*T*QEQQMNHPFLCLPTLKSRI 131
>AV428269
Length = 418
Score = 24.6 bits (52), Expect = 6.5
Identities = 23/96 (23%), Positives = 39/96 (39%)
Frame = +3
Query: 45 SQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKAN 104
+Q NH S +S + M+ + ++ELQL + ++ L L +L +KA+
Sbjct: 102 NQETNHSTMSEQSQLRRTRMSSMQTDQELQLKASRDVAMAMAAKAKLLLREL--KTVKAD 275
Query: 105 AALAHSSSVFEPSSSVPQINFKTHPVHSNVSLGYSH 140
A A Q+ + +H N G SH
Sbjct: 276 LAFA--------KDRCAQLEEENRILHENRERGDSH 359
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.130 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,938,803
Number of Sequences: 28460
Number of extensions: 46601
Number of successful extensions: 256
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 255
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 256
length of query: 140
length of database: 4,897,600
effective HSP length: 81
effective length of query: 59
effective length of database: 2,592,340
effective search space: 152948060
effective search space used: 152948060
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 50 (23.9 bits)
Lotus: description of TM0091a.5


