

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
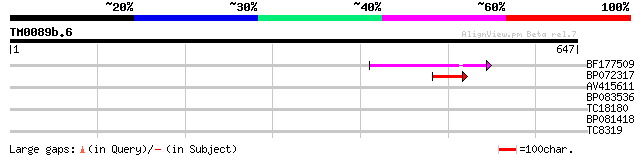
Query= TM0089b.6
(647 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF177509 50 1e-06
BP072317 49 2e-06
AV415611 35 0.038
BP083536 34 0.084
TC18180 weakly similar to GB|AAF04891.1|6175165|ATAC011437 Mutat... 30 1.6
BP081418 28 4.6
TC8319 weakly similar to UP|RK4_TOBAC (O80361) 50S ribosomal pro... 27 7.8
>BF177509
Length = 472
Score = 49.7 bits (117), Expect = 1e-06
Identities = 37/141 (26%), Positives = 62/141 (43%), Gaps = 1/141 (0%)
Frame = +1
Query: 411 WLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFA 470
W+ +IP W F + N + ES N IL A P++ M++ IR +M F
Sbjct: 19 WIRRIPPRLWATAYFEGQRFGHLTAN-IVESLNTWILEASGLPIIQMMECIRRQLMTWFN 195
Query: 471 TMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHD-QFIVDLKKH 529
E ++ ++P + ++ FEV +SH+ IVD++
Sbjct: 196 ERRETSMQWASILVPSAERCVAEALERARTYQVLRANEAEFEV---ISHEGSNIVDIRNR 366
Query: 530 TCMCNFWELVGIPFRHEIAAI 550
C+C W+L G+P H +AA+
Sbjct: 367 CCLCRGWQLYGLPCAHAVAAL 429
>BP072317
Length = 436
Score = 49.3 bits (116), Expect = 2e-06
Identities = 21/40 (52%), Positives = 31/40 (77%)
Frame = +3
Query: 483 VMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQF 522
VMPKPL+RL WE++ TS+ +PQ G ++EVKH+V+ + F
Sbjct: 315 VMPKPLKRLAWEVKGTSNGLPQASGEGLYEVKHVVTLEGF 434
>AV415611
Length = 219
Score = 35.0 bits (79), Expect = 0.038
Identities = 14/33 (42%), Positives = 19/33 (57%)
Frame = +3
Query: 523 IVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQ 555
+VD+ + C C W+L G+P H IA I GQ
Sbjct: 108 VVDIDRWECSCKTWQLTGVPCCHAIAVIVGIGQ 206
>BP083536
Length = 377
Score = 33.9 bits (76), Expect = 0.084
Identities = 12/22 (54%), Positives = 15/22 (67%)
Frame = +1
Query: 531 CMCNFWELVGIPFRHEIAAISH 552
C CN WELV IP RH +A + +
Sbjct: 307 CCCNLWELVEIPCRHALAGVHY 372
>TC18180 weakly similar to GB|AAF04891.1|6175165|ATAC011437 Mutator-like
transposase {Arabidopsis thaliana;} , partial (4%)
Length = 513
Score = 29.6 bits (65), Expect = 1.6
Identities = 23/89 (25%), Positives = 35/89 (38%), Gaps = 3/89 (3%)
Frame = +3
Query: 559 DFVHAYYRRAAYRATYEHLVSPISGQKLWPTTNDEYILS---PLYRRALGRPKKLRMRDP 615
D+ Y +YR TY V+PI + + + + +++ P RR GRP
Sbjct: 12 DYCSRYCTTESYRLTYSERVNPIPIMAVPESKDSQPVVTVTPPFTRRPPGRP-------- 167
Query: 616 HEDTTNNHTRLHIGVSRNKFSRCGQFGHN 644
T + +I SRC GHN
Sbjct: 168 ---ATKRYVSQNIVKRELHCSRCKGLGHN 245
>BP081418
Length = 459
Score = 28.1 bits (61), Expect = 4.6
Identities = 18/56 (32%), Positives = 28/56 (49%), Gaps = 6/56 (10%)
Frame = +2
Query: 531 CMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRA------AYRATYEHLVSP 580
C+C+ V F+HE + +H+ P DFV+ RR + RA++ L SP
Sbjct: 290 CLCS----VNCNFQHEDSIHTHSSSLPSDFVNMSCRRCSNSNSHSTRASHWFLSSP 445
>TC8319 weakly similar to UP|RK4_TOBAC (O80361) 50S ribosomal protein L4,
chloroplast precursor (R-protein L4), partial (30%)
Length = 650
Score = 27.3 bits (59), Expect = 7.8
Identities = 13/37 (35%), Positives = 20/37 (53%)
Frame = -1
Query: 97 ISLCFVVRLVESIHLVSKQCMDYIPVGECSTTSLHQV 133
+ L ++ L++ IHL+ C+ Y P S T HQV
Sbjct: 293 LHLLLLLPLLQHIHLLHHHCLGY*PHTSDSKTPQHQV 183
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.341 0.148 0.503
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,352,629
Number of Sequences: 28460
Number of extensions: 181700
Number of successful extensions: 1260
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1249
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1258
length of query: 647
length of database: 4,897,600
effective HSP length: 96
effective length of query: 551
effective length of database: 2,165,440
effective search space: 1193157440
effective search space used: 1193157440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0089b.6


