

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0089a.1
(119 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
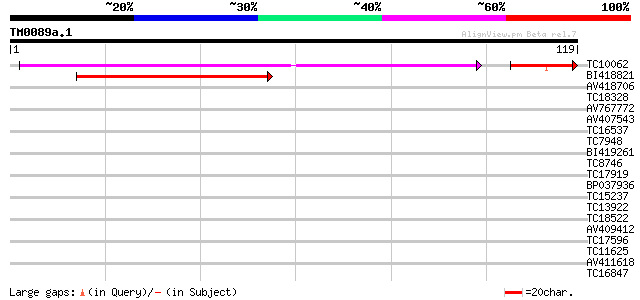
Score E
Sequences producing significant alignments: (bits) Value
TC10062 similar to BAD08700 (BAD08700) Cold shock domain protein... 67 3e-13
BI418821 39 2e-04
AV418706 30 0.11
TC18328 weakly similar to UP|HSP2_PANPA (P35299) Sperm histone P... 27 0.91
AV767772 27 1.2
AV407543 26 2.0
TC16537 26 2.0
TC7948 25 2.6
BI419261 25 3.5
TC8746 similar to PIR|T06396|T06396 isoprenylated protein - soyb... 25 3.5
TC17919 similar to UP|KPYA_RICCO (Q43117) Pyruvate kinase isozym... 25 3.5
BP037936 25 4.5
TC15237 similar to UP|AAQ24632 (AAQ24632) GPRP, partial (64%) 25 4.5
TC13922 UP|Q99NH6 (Q99NH6) bZIP protein ATF7, partial (5%) 25 4.5
TC18522 weakly similar to UP|HSP1_OCTVU (P83214) Sperm protamine... 25 4.5
AV409412 22 5.1
TC17596 similar to UP|Q42768 (Q42768) Acetohydroxyacid synthase ... 24 5.9
TC11625 weakly similar to UP|O81015 (O81015) At2g26920 protein, ... 24 5.9
AV411618 24 5.9
TC16847 weakly similar to PIR|T51733|T51733 cytidine deaminase ... 24 7.7
>TC10062 similar to BAD08700 (BAD08700) Cold shock domain protein 2, partial
(66%)
Length = 481
Score = 67.0 bits (162), Expect(2) = 3e-13
Identities = 43/97 (44%), Positives = 55/97 (56%)
Frame = +3
Query: 3 MSVSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTILI 62
MS S V K+K F+DQKGF IT DDGG+DLF+ S+IKS+GF CL E E V+ ++
Sbjct: 33 MSDSRVNGKVKWFNDQKGFGFITPDDGGDDLFVHQSEIKSEGFRCLGEGEDVEFIVA-SD 209
Query: 63 PMAALRR*RD*RPD*API*GTCCGGNDSSGYGRGGGY 99
P + P+ + GT GG G GGGY
Sbjct: 210 PDGRAKAIEVTGPNGQAVQGTRRGGGAGGGGRGGGGY 320
Score = 21.6 bits (44), Expect(2) = 3e-13
Identities = 9/16 (56%), Positives = 10/16 (62%), Gaps = 2/16 (12%)
Frame = +2
Query: 106 RIWRNG--RQKRWRWF 119
RIWR R KRW W+
Sbjct: 374 RIWRRRRIRWKRWLWW 421
>BI418821
Length = 614
Score = 38.9 bits (89), Expect = 2e-04
Identities = 19/41 (46%), Positives = 26/41 (63%)
Frame = +2
Query: 15 FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVD 55
F++ KGF I DDGG DLF+ S I+S G+ L E + V+
Sbjct: 101 FNNVKGFGFIKPDDGGEDLFVHQSSIRSDGYRTLLEGDLVE 223
>AV418706
Length = 374
Score = 30.0 bits (66), Expect = 0.11
Identities = 17/41 (41%), Positives = 18/41 (43%), Gaps = 6/41 (14%)
Frame = -3
Query: 84 CCGGND-SSGYGRGGGYQEVVDI-----RIWRNGRQKRWRW 118
CCGG G GR GG+ V I R WR RW W
Sbjct: 147 CCGGEGVQHGDGRWGGWGRGVGILSAVMRCWRRRCHCRWAW 25
>TC18328 weakly similar to UP|HSP2_PANPA (P35299) Sperm histone P2 precursor
(Protamine P2), partial (25%)
Length = 517
Score = 26.9 bits (58), Expect = 0.91
Identities = 13/37 (35%), Positives = 19/37 (51%), Gaps = 4/37 (10%)
Frame = -3
Query: 82 GTCCGGNDSSGYGRG----GGYQEVVDIRIWRNGRQK 114
G CCGG D RG G+ V + +WR+G ++
Sbjct: 146 GRCCGGGDWEVAARGKTGIHGFVHGVVLFLWRSGGER 36
>AV767772
Length = 460
Score = 26.6 bits (57), Expect = 1.2
Identities = 11/25 (44%), Positives = 15/25 (60%)
Frame = -3
Query: 92 GYGRGGGYQEVVDIRIWRNGRQKRW 116
G+G GGG+ E + W R+KRW
Sbjct: 380 GFG*GGGFWEEKERGYWN*LRKKRW 306
>AV407543
Length = 227
Score = 25.8 bits (55), Expect = 2.0
Identities = 9/17 (52%), Positives = 10/17 (57%)
Frame = -3
Query: 82 GTCCGGNDSSGYGRGGG 98
GTCCGG G+G G
Sbjct: 93 GTCCGGGGGGGHGGSVG 43
>TC16537
Length = 622
Score = 25.8 bits (55), Expect = 2.0
Identities = 8/15 (53%), Positives = 11/15 (73%)
Frame = -1
Query: 103 VDIRIWRNGRQKRWR 117
+D RIWR +K+WR
Sbjct: 238 LDTRIWRRDNEKKWR 194
>TC7948
Length = 1230
Score = 25.4 bits (54), Expect = 2.6
Identities = 15/43 (34%), Positives = 21/43 (47%), Gaps = 8/43 (18%)
Frame = -2
Query: 83 TCCGGNDSSGYGRGGGYQEVV--------DIRIWRNGRQKRWR 117
TCCGG GGG EV+ ++R+WR +R+R
Sbjct: 347 TCCGGV-------GGGGCEVIQVKYGE*SEVRVWRRIIMRRFR 240
>BI419261
Length = 591
Score = 25.0 bits (53), Expect = 3.5
Identities = 9/18 (50%), Positives = 11/18 (61%)
Frame = +2
Query: 87 GNDSSGYGRGGGYQEVVD 104
G D+ GYG G GY + D
Sbjct: 410 GRDAGGYGSGXGYSDGCD 463
>TC8746 similar to PIR|T06396|T06396 isoprenylated protein - soybean
(fragment) {Glycine max;}, partial (30%)
Length = 594
Score = 25.0 bits (53), Expect = 3.5
Identities = 10/17 (58%), Positives = 11/17 (63%)
Frame = +1
Query: 82 GTCCGGNDSSGYGRGGG 98
GTC GG S G+ GGG
Sbjct: 538 GTCGGGGISLGFSDGGG 588
>TC17919 similar to UP|KPYA_RICCO (Q43117) Pyruvate kinase isozyme A,
chloroplast precursor , partial (11%)
Length = 566
Score = 25.0 bits (53), Expect = 3.5
Identities = 11/25 (44%), Positives = 14/25 (56%)
Frame = -3
Query: 94 GRGGGYQEVVDIRIWRNGRQKRWRW 118
G GGG+ R+W+ G RWRW
Sbjct: 276 GGGGGW--CGGRRVWKMGGGCRWRW 208
>BP037936
Length = 518
Score = 24.6 bits (52), Expect = 4.5
Identities = 11/24 (45%), Positives = 13/24 (53%)
Frame = -1
Query: 82 GTCCGGNDSSGYGRGGGYQEVVDI 105
G CCGGN G GGG + +I
Sbjct: 146 GRCCGGN-----GGGGGVETAFEI 90
>TC15237 similar to UP|AAQ24632 (AAQ24632) GPRP, partial (64%)
Length = 814
Score = 24.6 bits (52), Expect = 4.5
Identities = 9/16 (56%), Positives = 10/16 (62%)
Frame = -2
Query: 84 CCGGNDSSGYGRGGGY 99
CCGG + GY GGY
Sbjct: 321 CCGGYPAGGYPCCGGY 274
>TC13922 UP|Q99NH6 (Q99NH6) bZIP protein ATF7, partial (5%)
Length = 511
Score = 24.6 bits (52), Expect = 4.5
Identities = 12/22 (54%), Positives = 12/22 (54%), Gaps = 4/22 (18%)
Frame = -2
Query: 85 CGGNDSS----GYGRGGGYQEV 102
CG N S G G GGGYQ V
Sbjct: 165 CGSNSGSSGGGGGGEGGGYQAV 100
>TC18522 weakly similar to UP|HSP1_OCTVU (P83214) Sperm protamine P1 (Po1)
[Contains: Sperm protamine P2 (Po2) (Main protamine)],
partial (68%)
Length = 320
Score = 24.6 bits (52), Expect = 4.5
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = -2
Query: 82 GTCCGGNDSSGYGRGGGYQEVVD 104
G GG + G GGG++EVV+
Sbjct: 220 GEDSGGETLAAVGGGGGFEEVVE 152
>AV409412
Length = 426
Score = 21.6 bits (44), Expect(2) = 5.1
Identities = 11/25 (44%), Positives = 12/25 (48%)
Frame = -1
Query: 94 GRGGGYQEVVDIRIWRNGRQKRWRW 118
GRG E I WR G + R RW
Sbjct: 279 GRGREVME*RGIS*WRKGGRGRNRW 205
Score = 21.2 bits (43), Expect(2) = 5.1
Identities = 9/17 (52%), Positives = 10/17 (57%)
Frame = -2
Query: 82 GTCCGGNDSSGYGRGGG 98
G GG+D G G GGG
Sbjct: 419 GVVEGGDDVEGGGGGGG 369
>TC17596 similar to UP|Q42768 (Q42768) Acetohydroxyacid synthase , partial
(16%)
Length = 537
Score = 24.3 bits (51), Expect = 5.9
Identities = 15/52 (28%), Positives = 18/52 (33%), Gaps = 17/52 (32%)
Frame = -2
Query: 84 CCGGNDSSGYGRGG--------------GYQEVVDIRIWRNG---RQKRWRW 118
CC G+ G+G G G E DI W G R+ W W
Sbjct: 200 CCSGSGGGGFGEGA*DSDGMRFVENGREG*GEEFDIAFWEVGG*RRR*EW*W 45
>TC11625 weakly similar to UP|O81015 (O81015) At2g26920 protein, partial
(8%)
Length = 410
Score = 24.3 bits (51), Expect = 5.9
Identities = 12/32 (37%), Positives = 15/32 (46%)
Frame = +2
Query: 86 GGNDSSGYGRGGGYQEVVDIRIWRNGRQKRWR 117
GG G G GGG + R WR ++R R
Sbjct: 260 GGGGGGGGGGGGGITVEWENRRWRERERERER 355
>AV411618
Length = 320
Score = 24.3 bits (51), Expect = 5.9
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = +2
Query: 90 SSGYGRGGGYQEVVDIRIWRNGRQKRW 116
SSGY R + +RIWRN + RW
Sbjct: 218 SSGYERR---KTTARVRIWRNEVKDRW 289
>TC16847 weakly similar to PIR|T51733|T51733 cytidine deaminase [validated]
- Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (74%)
Length = 1098
Score = 23.9 bits (50), Expect = 7.7
Identities = 16/56 (28%), Positives = 27/56 (47%)
Frame = +1
Query: 56 LLLTILIPMAALRR*RD*RPD*API*GTCCGGNDSSGYGRGGGYQEVVDIRIWRNG 111
L++T+L P RR D P+ G CC + S R ++++++ RNG
Sbjct: 790 LIMTVLSPPCWWRR-MGRWLDRRPLQGCCCKKSPLSASSRLSFVLQLIELKFVRNG 954
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.359 0.166 0.627
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,003,595
Number of Sequences: 28460
Number of extensions: 23826
Number of successful extensions: 557
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 529
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 556
length of query: 119
length of database: 4,897,600
effective HSP length: 79
effective length of query: 40
effective length of database: 2,649,260
effective search space: 105970400
effective search space used: 105970400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 49 (23.5 bits)
Lotus: description of TM0089a.1


