

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0087.21
(1215 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
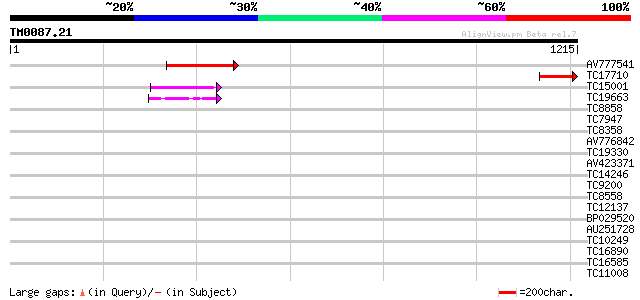
Score E
Sequences producing significant alignments: (bits) Value
AV777541 289 2e-78
TC17710 151 5e-37
TC15001 similar to UP|GLE1_YEAST (Q12315) RNA export factor GLE1... 55 7e-08
TC19663 similar to UP|Q8H6S8 (Q8H6S8) Translation initiation fac... 47 1e-05
TC8858 homologue to UP|Q84N37 (Q84N37) Potyvirus VPg interacting... 41 0.001
TC7947 similar to UP|Q9ST69 (Q9ST69) Ribosomal protein S5, parti... 36 0.042
TC8358 similar to UP|Q84T14 (Q84T14) Vacuolar ATPase subunit E (... 35 0.054
AV776842 34 0.16
TC19330 similar to UP|Q96258 (Q96258) AR791, partial (11%) 33 0.27
AV423371 33 0.35
TC14246 similar to UP|Q84LL6 (Q84LL6) Salt tolerance protein 5, ... 32 0.46
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 32 0.78
TC8558 weakly similar to UP|Q9NDI0 (Q9NDI0) 200 kDa antigen p200... 31 1.0
TC12137 similar to UP|Q8L7W3 (Q8L7W3) At2g29210/F16P2.41, partia... 30 1.7
BP029520 30 1.7
AU251728 30 2.3
TC10249 similar to UP|Q84LL9 (Q84LL9) Salt tolerance protein 3, ... 30 3.0
TC16890 30 3.0
TC16585 30 3.0
TC11008 29 3.9
>AV777541
Length = 466
Score = 289 bits (739), Expect = 2e-78
Identities = 155/155 (100%), Positives = 155/155 (100%)
Frame = +1
Query: 336 KKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKE 395
KKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKE
Sbjct: 1 KKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKE 180
Query: 396 EAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIY 455
EAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIY
Sbjct: 181 EAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIY 360
Query: 456 LEQIRERANLRDQSSPLLRRSLNKDGQGRSTPNNS 490
LEQIRERANLRDQSSPLLRRSLNKDGQGRSTPNNS
Sbjct: 361 LEQIRERANLRDQSSPLLRRSLNKDGQGRSTPNNS 465
>TC17710
Length = 593
Score = 151 bits (382), Expect = 5e-37
Identities = 76/80 (95%), Positives = 77/80 (96%)
Frame = +2
Query: 1136 KGIRAFLGKSGVLGNSLKNGRLKSLRDGKTTKSSEEVVPRQNLSTSETSHSMLHCRFPQS 1195
K IRAFLGKSGVLGNSLKN RLKSLRD KTT+SSEEVVPRQNLSTSETSHSMLHCRFPQS
Sbjct: 2 KAIRAFLGKSGVLGNSLKNARLKSLRDAKTTESSEEVVPRQNLSTSETSHSMLHCRFPQS 181
Query: 1196 FIDKVEQFFSAEIANGVDGV 1215
FIDKVEQFFSAEIANGVDGV
Sbjct: 182 FIDKVEQFFSAEIANGVDGV 241
>TC15001 similar to UP|GLE1_YEAST (Q12315) RNA export factor GLE1, partial
(4%)
Length = 988
Score = 55.1 bits (131), Expect = 7e-08
Identities = 46/155 (29%), Positives = 73/155 (46%), Gaps = 3/155 (1%)
Frame = +2
Query: 303 RHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQ 362
R EA L + + + R SLN E ++ ++++L R ++ V K+
Sbjct: 362 RKEAELKLIEEETAKRVEEAIRKRVEESLNSEEVQVEIQRRLEEGRKRLIVEVAVQLEKE 541
Query: 363 KEDLAREEAVLERRKLIEAEKLQRLAEMQRKK-EEAQIRR--EEERKASSAAREARAIEQ 419
KE E E + E E L R+ E RKK EEAQ R E++R+ RE I++
Sbjct: 542 KEAALIEAREKEEQARKEKEDLDRMLEENRKKIEEAQRREALEQQRREEERYRELEEIQR 721
Query: 420 LRRKEERAKAQQEEAELLAQKLAERLNESEQRRKI 454
+ + R K Q+EE E L Q + L +++ R K+
Sbjct: 722 QKEEAMRRKKQEEEQERLNQ--IKLLGKNKSRPKL 820
>TC19663 similar to UP|Q8H6S8 (Q8H6S8) Translation initiation factor,
partial (10%)
Length = 485
Score = 47.4 bits (111), Expect = 1e-05
Identities = 47/158 (29%), Positives = 74/158 (46%), Gaps = 1/158 (0%)
Frame = +3
Query: 298 QRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEK-LQ 356
Q+ + + E + A A S+ NE E K ++ K + S+ + A+K L
Sbjct: 36 QKKKKKKEKEKEKKAAAAAAGSAPANET-------VEVKAEVVESKKNESKTKAADKKLP 194
Query: 357 VIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARA 416
+ +E LAR + ER+K E EKL +KEE + RR+EE + A EAR
Sbjct: 195 KHVREMQEALARRKEAEERKKREEEEKL--------RKEEEERRRQEELERQ--AEEARR 344
Query: 417 IEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKI 454
++ R KE+ K +QE L ++ E+ RR+I
Sbjct: 345 RKKEREKEKLLKKKQEGKLLTGKQKEEQRRLEAMRRQI 458
Score = 40.8 bits (94), Expect = 0.001
Identities = 31/118 (26%), Positives = 56/118 (47%), Gaps = 12/118 (10%)
Frame = +3
Query: 353 EKLQVIKSKQKEDLAR---------EEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREE 403
+K + K K+KE A E V + +++E++K + + KK +R +
Sbjct: 36 QKKKKKKEKEKEKKAAAAAAGSAPANETVEVKAEVVESKKNESKTKAADKKLPKHVREMQ 215
Query: 404 E---RKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIYLEQ 458
E R+ + R+ R E+ RKEE + +QEE E A++ R E E+ + + +Q
Sbjct: 216 EALARRKEAEERKKREEEEKLRKEEEERRRQEELERQAEEARRRKKEREKEKLLKKKQ 389
>TC8858 homologue to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), partial (65%)
Length = 1272
Score = 41.2 bits (95), Expect = 0.001
Identities = 36/146 (24%), Positives = 75/146 (50%), Gaps = 14/146 (9%)
Frame = +1
Query: 339 ILRQKLHGSELRRAEKLQVIKS---------KQKEDLAREEAVLE---RRKLIEAEKLQR 386
++++ + E+ EK+++ K ++ D ARE L+ ++K ++ E+L+R
Sbjct: 493 VVQEAIRKMEMVADEKMRMFKKARLALEACDRELADKAREAGELKTERQKKKLQIEELER 672
Query: 387 LAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLN 446
+ + K EA + + KA+ A REA ++++ AK+ + E E + L ++L+
Sbjct: 673 IVRL--KNAEADMF---QLKANEAKREAERLQRI----ALAKSDKSEEEYTSNYLKQKLS 825
Query: 447 ESEQRRKIYLEQIR--ERANLRDQSS 470
E+E ++ E+I+ E + L SS
Sbjct: 826 EAEAEKQYLYEKIKLQESSRLSQSSS 903
>TC7947 similar to UP|Q9ST69 (Q9ST69) Ribosomal protein S5, partial (81%)
Length = 1294
Score = 35.8 bits (81), Expect = 0.042
Identities = 25/57 (43%), Positives = 32/57 (55%), Gaps = 1/57 (1%)
Frame = -3
Query: 370 EAVLERRKLIEAEKLQRLA-EMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEE 425
E V+ERR L+ K L E +R+K+E + R EER A +A AIE RKEE
Sbjct: 242 EEVVERRGLVRRGKKGSLVVEGRREKDEEERREREERAAEGTTEDAVAIE--GRKEE 78
>TC8358 similar to UP|Q84T14 (Q84T14) Vacuolar ATPase subunit E (Fragment),
complete
Length = 1321
Score = 35.4 bits (80), Expect = 0.054
Identities = 27/85 (31%), Positives = 43/85 (49%)
Frame = +1
Query: 367 AREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEER 426
A EE +E+ +L+EAEK + E +RK+ + +IRR+ E S A I+ L+ +++
Sbjct: 181 AEEEFNIEKLQLVEAEKKKIRQEYERKERQIEIRRKIE---YSMQLNASRIKVLQAQDDL 351
Query: 427 AKAQQEEAELLAQKLAERLNESEQR 451
+E A E LN S R
Sbjct: 352 VNTMKEAAS------KELLNVSHHR 408
>AV776842
Length = 380
Score = 33.9 bits (76), Expect = 0.16
Identities = 25/84 (29%), Positives = 40/84 (46%)
Frame = -2
Query: 198 LSSPFRVSSRMSYSPSLGRKSAERVRTLHDKLMSPEKKKKTTSDLKKEAEEKHARALRIR 257
L++ R +++ +P + + +R+RTLH KL K KT S + K A + I
Sbjct: 313 LNTQQRSYTQILQNPKISTRKRKRIRTLHGKL----GKMKTLSSVAKRAP-----TMAIE 161
Query: 258 SELETERVQKLQRTSQKLNRVSEW 281
+ +E E + L QK R S W
Sbjct: 160 ASIEREEEESLMENLQK--RCSLW 95
>TC19330 similar to UP|Q96258 (Q96258) AR791, partial (11%)
Length = 507
Score = 33.1 bits (74), Expect = 0.27
Identities = 39/156 (25%), Positives = 71/156 (45%), Gaps = 9/156 (5%)
Frame = +2
Query: 333 EENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQ-RLAEMQ 391
E+NK+ I+ K +R KLQ ++S+ E L R K +EAE Q R ++
Sbjct: 56 EQNKEQIINLK------QRVSKLQDLESQATASDQEIETKLRRLKDLEAEAEQLRKTNLR 217
Query: 392 RKKEEAQIRR--EEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKL-AERLNES 448
+ + + + R + + ++A E + LR + ER + + E ++L A+R ++
Sbjct: 218 LQMDNSDLARRLDSTQILANAVLEDPEADALREEGERLRLENEGLTKEVEQLQADRCSDV 397
Query: 449 EQRRKIYLEQI-----RERANLRDQSSPLLRRSLNK 479
E+ +YL I E N + + R L+K
Sbjct: 398 EE--LVYLRWINACLRHEMRNYQPPPGKTVARDLSK 499
>AV423371
Length = 437
Score = 32.7 bits (73), Expect = 0.35
Identities = 28/88 (31%), Positives = 43/88 (48%)
Frame = +3
Query: 365 DLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKE 424
D R + VL + K E + + + E+ +K +EA EE +A+ A AR E+LRR +
Sbjct: 78 DKTRSDIVLAQAK--ENDGREMIIELPKKLKEA---AEEANQANLHAEAAR--EELRRVK 236
Query: 425 ERAKAQQEEAELLAQKLAERLNESEQRR 452
E A+ + A + KL E E R
Sbjct: 237 EEAEQAKAGASTMQSKLLAAQKEIEAAR 320
>TC14246 similar to UP|Q84LL6 (Q84LL6) Salt tolerance protein 5, partial
(57%)
Length = 1280
Score = 32.3 bits (72), Expect = 0.46
Identities = 25/74 (33%), Positives = 41/74 (54%)
Frame = +1
Query: 360 SKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQ 419
S++ + LA E A E L+ A A +RKK +REEE KAS +A+A ++
Sbjct: 268 SRESDFLATEAAEKEVASLVRA------AREKRKK-----KREEELKASE--EKAKAEKR 408
Query: 420 LRRKEERAKAQQEE 433
L+ E++ KA++ +
Sbjct: 409 LKEAEKKPKAEEND 450
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 31.6 bits (70), Expect = 0.78
Identities = 27/129 (20%), Positives = 61/129 (46%), Gaps = 1/129 (0%)
Frame = +1
Query: 335 NKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLER-RKLIEAEKLQRLAEMQRK 393
N+ + ++KL + AE +++K + ++ E V E + +E E+ + E + +
Sbjct: 94 NESITKKRKLVPKSSQSAELPFKLQNKLEAEVQEYEEVEEEVEEEVEEEEEEEEEEEEGE 273
Query: 394 KEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLNESEQRRK 453
+EE + EEE +A + + I++L +++ LLA + + +++ RK
Sbjct: 274 EEEEEEEEEEEVEAEAEEEDDEPIQKLLEP----MGKEQIISLLADAASNHRDVADRIRK 441
Query: 454 IYLEQIRER 462
+ E + R
Sbjct: 442 VADEDVSHR 468
>TC8558 weakly similar to UP|Q9NDI0 (Q9NDI0) 200 kDa antigen p200
(Fragment), partial (4%)
Length = 585
Score = 31.2 bits (69), Expect = 1.0
Identities = 45/169 (26%), Positives = 73/169 (42%), Gaps = 4/169 (2%)
Frame = +1
Query: 374 ERRKLI----EAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKA 429
ERR I E + R A + E Q R E+ER AREA E+ R++EE +A
Sbjct: 46 ERRNNIRLREEQDAAYRAALEADQARERQRREEQER----LAREAAEAERKRKEEE--EA 207
Query: 430 QQEEAELLAQKLAERLNESEQRRKIYLEQIRERANLRDQSSPLLRRSLNKDGQGRSTPNN 489
++ EA A+K A +Q+ + E+ + N+ + +L R + + R N
Sbjct: 208 REREAREAAEKQAALAKIRQQKAESLGEEPAKGPNV----TQVLVRFPTGERKERRFNNT 375
Query: 490 SSDDSQTNIASGLGSSLGIGSITLQHSMKRRIKRIRQKLMALKYEFLEP 538
+ S + LG L S +L + R + + ++LK L P
Sbjct: 376 VTIQSVYDYVDSLG-CLEADSYSLVSNFPRVVYGQEKLTLSLKEAGLHP 519
Score = 29.6 bits (65), Expect = 3.0
Identities = 34/127 (26%), Positives = 58/127 (44%), Gaps = 10/127 (7%)
Frame = +1
Query: 333 EENKKLILRQKLHGSELR-----RAEKLQVIKSKQKEDLAREEAVLER-----RKLIEAE 382
EE+ ++ +L E R R E+ ++ + D ARE E R+ EAE
Sbjct: 1 EESSPTLVAARLDAEERRNNIRLREEQDAAYRAALEADQARERQRREEQERLAREAAEAE 180
Query: 383 KLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLA 442
+ ++KEE + R ER+A AA + A+ AK +Q++AE L ++ A
Sbjct: 181 R--------KRKEEEEAR---EREAREAAEKQAAL---------AKIRQQKAESLGEEPA 300
Query: 443 ERLNESE 449
+ N ++
Sbjct: 301 KGPNVTQ 321
>TC12137 similar to UP|Q8L7W3 (Q8L7W3) At2g29210/F16P2.41, partial (9%)
Length = 579
Score = 30.4 bits (67), Expect = 1.7
Identities = 28/96 (29%), Positives = 48/96 (49%), Gaps = 1/96 (1%)
Frame = +3
Query: 369 EEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAK 428
+++ LE RK + +K + E + +KEE + RREE R RE R E+L+ K +
Sbjct: 27 DDSELEDRKETQRKKRE---EKKLRKEEKRRRREERRH----RREERRAEKLKLKSKTDY 185
Query: 429 -AQQEEAELLAQKLAERLNESEQRRKIYLEQIRERA 463
+ EEAE + + +K+ +E +R +A
Sbjct: 186 ISDDEEAERMDDHPSNNEETQVDPKKLEIE-LRNKA 290
Score = 28.9 bits (63), Expect = 5.1
Identities = 26/93 (27%), Positives = 42/93 (44%)
Frame = +3
Query: 331 LNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEKLQRLAEM 390
L +E K+ ++ H E RRAEKL K K K D ++ E+ +R+ +
Sbjct: 87 LRKEEKRRRREERRHRREERRAEKL---KLKSKTDYISDD-----------EEAERMDDH 224
Query: 391 QRKKEEAQIRREEERKASSAAREARAIEQLRRK 423
EE Q+ + +K R +A+E L+ K
Sbjct: 225 PSNNEETQV---DPKKLEIELRN-KALESLKAK 311
>BP029520
Length = 381
Score = 30.4 bits (67), Expect = 1.7
Identities = 15/40 (37%), Positives = 20/40 (49%)
Frame = +1
Query: 152 INEKNPRSTDNLRRQMQLPEKDKEKRSTSLGKSMNAWKEK 191
IN D R Q Q P++ K++ GK+ N WKEK
Sbjct: 19 INRAQDVLVDTNRIQPQGPKRQKQRNIERQGKARNIWKEK 138
>AU251728
Length = 397
Score = 30.0 bits (66), Expect = 2.3
Identities = 16/33 (48%), Positives = 23/33 (69%), Gaps = 1/33 (3%)
Frame = +2
Query: 906 SNLAQPVVFLLSALSET-GLVSLPSLLTAVLLQ 937
S+L P +FLL+ L + L+S+PS LT +LLQ
Sbjct: 131 SSLESPFIFLLTPLQASLSLLSIPSFLTFLLLQ 229
>TC10249 similar to UP|Q84LL9 (Q84LL9) Salt tolerance protein 3, partial
(35%)
Length = 1096
Score = 29.6 bits (65), Expect = 3.0
Identities = 25/81 (30%), Positives = 41/81 (49%)
Frame = +1
Query: 387 LAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKEERAKAQQEEAELLAQKLAERLN 446
++E Q KE+A EEER ARE A E+ +RKE+ + ++ +K ++
Sbjct: 118 ISEFQTAKEKAI---EEER----LAREREAEERRKRKEQEYERKRRGDSSDREKHRDKDR 276
Query: 447 ESEQRRKIYLEQIRERANLRD 467
+ E+ R + RER+ RD
Sbjct: 277 DRERDRYRDRDSDRERSRERD 339
Score = 29.3 bits (64), Expect = 3.9
Identities = 25/109 (22%), Positives = 49/109 (44%)
Frame = +1
Query: 359 KSKQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKASSAAREARAIE 418
K+ ++E LARE ERRK E E ++ +E+ R+++R R +
Sbjct: 145 KAIEEERLAREREAEERRKRKEQEYERKRRGDSSDREK---HRDKDRDRERDRYRDRDSD 315
Query: 419 QLRRKEERAKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRERANLRD 467
+ R +E + ++ + +L R N R Y ++ R R+ +++
Sbjct: 316 RERSRERDVRGYRDGGRGVDSRLGNRRNGGRDR---YRDRSRSRSPVKN 453
>TC16890
Length = 965
Score = 29.6 bits (65), Expect = 3.0
Identities = 43/194 (22%), Positives = 78/194 (40%), Gaps = 14/194 (7%)
Frame = +2
Query: 297 HQRSESRHEAFLAQVAKRAGDESSKVNEVRFITSLNEENKKLILRQKLHGSELRRAEKLQ 356
H +++++H+ + K G E + E N + K L K+ G ++ +L
Sbjct: 38 HFKTKTKHK----EKEKEKGKEREENKEE------NSKKKGF*LLNKMQGGGEQQNNQLV 187
Query: 357 VIKS--------KQKEDLAREEAVLERRKLIEAEKLQRLAEMQRKKEEAQIRREEERKAS 408
V S KED + L K E E ++ E++ K + R EEE K
Sbjct: 188 VQNSGSLSFSSQMSKEDEEMSRSALSNFKAKEEEIERKKMEVREKVQLQLGRVEEETKRL 367
Query: 409 SAAR-EARAIEQLRRKE-----ERAKAQQEEAELLAQKLAERLNESEQRRKIYLEQIRER 462
+ R E ++ RKE +R + +E + L ++ E + + + E+ RE+
Sbjct: 368 ATIREELESLADPMRKEVAIVRKRIDSVNKELKPLGHTCQKKEKEYKDALEAFNEKNREK 547
Query: 463 ANLRDQSSPLLRRS 476
L + L+ S
Sbjct: 548 VQLITRLMELVSES 589
>TC16585
Length = 426
Score = 29.6 bits (65), Expect = 3.0
Identities = 18/51 (35%), Positives = 29/51 (56%)
Frame = -1
Query: 485 STPNNSSDDSQTNIASGLGSSLGIGSITLQHSMKRRIKRIRQKLMALKYEF 535
++PNNS+ +S+T S S IG+ T S++ R +RIR ++ Y F
Sbjct: 387 ASPNNSTGNSKTTAQS----SNRIGNTTTSQSLQFRHRRIRFSVVIRYYRF 247
>TC11008
Length = 796
Score = 29.3 bits (64), Expect = 3.9
Identities = 45/221 (20%), Positives = 79/221 (35%), Gaps = 10/221 (4%)
Frame = +3
Query: 214 LGRKSAERVRTLHDKLMSPEKKKKTTSDLKKEAEEKHAR-----ALRIRSELETERVQKL 268
L K RVR D L +K+++ D KK AEE + AL++ SE ++
Sbjct: 135 LEEKRKRRVREKEDDLADKQKEEEEIVDAKKRAEEDQRQKQQRDALKLLSEHVVNSGEET 314
Query: 269 QRTSQKLNRV----SEWHAVRHLKLREGMYARHQRSESRH-EAFLAQVAKRAGDESSKVN 323
T + V +E V H REG + H E+ +A V
Sbjct: 315 MATEEIAKEVKYIDAEQDTVAHYS-REGHIGDGNSVNAIHDESAMASVGTTDTQSGGNGP 491
Query: 324 EVRFITSLNEENKKLILRQKLHGSELRRAEKLQVIKSKQKEDLAREEAVLERRKLIEAEK 383
+ L K+ + H E A K + ++ D + EE + + +
Sbjct: 492 LKKLGFGLVGSGKRAAVASAFHEEEEDEAHKDKKLRPWVPIDYSTEELQAVQPTISGSTP 671
Query: 384 LQRLAEMQRKKEEAQIRREEERKASSAAREARAIEQLRRKE 424
+A + K + +E++ R R+ E+ ++
Sbjct: 672 PNLVAAAEFAKRISSTNFKEDKPDGERDRSRRSNEKSNHRD 794
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.129 0.350
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,927,429
Number of Sequences: 28460
Number of extensions: 210506
Number of successful extensions: 932
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 911
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 924
length of query: 1215
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1114
effective length of database: 2,023,140
effective search space: 2253777960
effective search space used: 2253777960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0087.21


