

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.5
(176 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
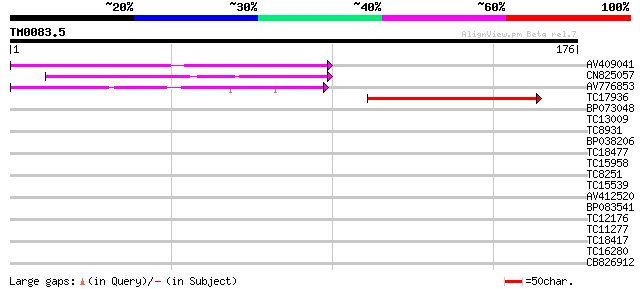
Score E
Sequences producing significant alignments: (bits) Value
AV409041 77 2e-15
CN825057 66 4e-12
AV776853 52 6e-08
TC17936 similar to UP|Q8C1C0 (Q8C1C0) PPAR gamma coactivator-1BE... 46 3e-06
BP073048 28 0.68
TC13009 similar to GB|AAL09709.1|15982737|AY057475 AT4g23750/F9D... 28 1.2
TC8931 similar to GB|AAL09731.1|15982767|AY057490 At2g32240/F22D... 27 2.0
BP038206 27 2.0
TC18477 26 3.4
TC15958 similar to GB|AAP37838.1|30725632|BT008479 At4g24220 {Ar... 26 3.4
TC8251 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Ara... 26 4.4
TC15539 25 5.8
AV412520 25 7.5
BP083541 25 7.5
TC12176 weakly similar to UP|AAQ09151 (AAQ09151) NADH dehydrogen... 25 7.5
TC11277 25 7.5
TC18417 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, pa... 25 9.8
TC16280 25 9.8
CB826912 25 9.8
>AV409041
Length = 349
Score = 76.6 bits (187), Expect = 2e-15
Identities = 38/100 (38%), Positives = 55/100 (55%)
Frame = +3
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
+EFES +AA FY EYA+ GF +S RRS + + C + G+ + +
Sbjct: 45 IEFESHEAAYAFYKEYAKSAGFGTAKLSSRRSRASKEFIDAKFSCIRYGN----KQQSDD 212
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLV 100
PR S + GCKA +H+K + GKW + FVK+HNH L+
Sbjct: 213 AINPRPSPKIGCKASMHVKRRQDGKWYVYSFVKEHNHELL 332
>CN825057
Length = 721
Score = 65.9 bits (159), Expect = 4e-12
Identities = 32/89 (35%), Positives = 50/89 (55%)
Frame = +1
Query: 12 FYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSIRRPRASTREG 71
FY EYA+ +GF + + RRS++ + + C++ G G GS RR +
Sbjct: 10 FYQEYAKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYGVTPESDG--GSNRRSSVKKTD- 180
Query: 72 CKAMIHIKYDKSGKWVITKFVKDHNHPLV 100
CKA +H+K GKW+I +F+K+HNH L+
Sbjct: 181 CKACMHVKRKPDGKWIIHEFIKEHNHELL 267
>AV776853
Length = 625
Score = 52.0 bits (123), Expect = 6e-08
Identities = 34/105 (32%), Positives = 51/105 (48%), Gaps = 6/105 (5%)
Frame = +1
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
++F SE+ A FY YA++ GF++R R + G + R+ CN++G +R K
Sbjct: 238 LDFGSEEEAYQFYQAYAKYQGFIVRKDDIGR-DYHGNVNMRQFVCNRQG----LRSKKHY 402
Query: 61 IRRPRAS-----TREGCKAMIHIKYD-KSGKWVITKFVKDHNHPL 99
R R T C A + + D K GKW + F + HNH L
Sbjct: 403 NRTDRKRDHKPVTHTNCLAKLRVHLDYKIGKWKVVSFEECHNHEL 537
>TC17936 similar to UP|Q8C1C0 (Q8C1C0) PPAR gamma coactivator-1BETA protein,
partial (3%)
Length = 552
Score = 46.2 bits (108), Expect = 3e-06
Identities = 23/54 (42%), Positives = 34/54 (62%)
Frame = -3
Query: 112 EKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKE 165
+K KKIQELT E+ I + C Y+ L + ++ +EE KLSVK+ + +LKE
Sbjct: 418 DKQKKIQELTGELEITNQRCEVYRANLLAVLRDMEEQKLKLSVKVQNARLSLKE 257
>BP073048
Length = 465
Score = 28.5 bits (62), Expect = 0.68
Identities = 14/31 (45%), Positives = 18/31 (57%)
Frame = -1
Query: 98 PLVVSPREARQTMDEKDKKIQELTAEIRIKK 128
PL+VS Q + E D+KIQE + RI K
Sbjct: 330 PLLVSSYHMDQMLKEHDRKIQEFSRRARIDK 238
>TC13009 similar to GB|AAL09709.1|15982737|AY057475 AT4g23750/F9D16_220
{Arabidopsis thaliana;}, partial (5%)
Length = 642
Score = 27.7 bits (60), Expect = 1.2
Identities = 17/61 (27%), Positives = 32/61 (51%), Gaps = 12/61 (19%)
Frame = -3
Query: 117 IQELTAEIRIKKRLCAT-------YQEQLSSFMKIVE-----EHSDKLSVKIHHIVNNLK 164
I + +E +I K++C +++ S + I+E ++ KL VK+HHI+ N K
Sbjct: 331 IPTIESEFQILKQICTRAAATTIIFEKHASFWQHIIEAVVSVQNILKLDVKLHHIIWNWK 152
Query: 165 E 165
+
Sbjct: 151 Q 149
>TC8931 similar to GB|AAL09731.1|15982767|AY057490 At2g32240/F22D22.1
{Arabidopsis thaliana;} , partial (4%)
Length = 876
Score = 26.9 bits (58), Expect = 2.0
Identities = 15/56 (26%), Positives = 28/56 (49%)
Frame = +1
Query: 100 VVSPREARQTMDEKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVK 155
+ + + A E + ++ E+ IKK Q+Q++ + ++ SDKLSVK
Sbjct: 91 IANQKVAESQKLELEASLKNSLEELEIKKNEITLLQKQVTDLEQKLQLTSDKLSVK 258
>BP038206
Length = 563
Score = 26.9 bits (58), Expect = 2.0
Identities = 12/60 (20%), Positives = 29/60 (48%)
Frame = -3
Query: 95 HNHPLVVSPREARQTMDEKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSV 154
H++PL + Q + + ++ IKK + Y+ Q + MK+++ H ++++
Sbjct: 351 HDYPLSIVDHIGFQRYSNSLQPLFQVPTRNTIKKEVIKIYEFQKTCVMKMLDSHEGRVAI 172
>TC18477
Length = 655
Score = 26.2 bits (56), Expect = 3.4
Identities = 19/66 (28%), Positives = 31/66 (46%), Gaps = 8/66 (12%)
Frame = +3
Query: 113 KDKKIQELTAEIRIKKRLCATYQEQLSSFMKI--------VEEHSDKLSVKIHHIVNNLK 164
K KKI R +++ A++ L S + + + EH ++ + I H+ NNLK
Sbjct: 273 KSKKIMVHREIERKRRQDMASFYASLRSLLPLEFIKGKRSLSEHINEAANYIKHMQNNLK 452
Query: 165 EFEPIE 170
E IE
Sbjct: 453 ELGAIE 470
>TC15958 similar to GB|AAP37838.1|30725632|BT008479 At4g24220 {Arabidopsis
thaliana;}, partial (52%)
Length = 800
Score = 26.2 bits (56), Expect = 3.4
Identities = 16/47 (34%), Positives = 23/47 (48%)
Frame = -1
Query: 27 MSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSIRRPRASTREGCK 73
+ CRR R R GC++ G + R + G RR +S R GC+
Sbjct: 434 LGCRRDRRC------RTGCSRLGDRRSRRVEGGGRRRRLSSGRRGCR 312
>TC8251 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Arabidopsis
thaliana;}, partial (41%)
Length = 909
Score = 25.8 bits (55), Expect = 4.4
Identities = 11/28 (39%), Positives = 16/28 (56%)
Frame = +1
Query: 131 CATYQEQLSSFMKIVEEHSDKLSVKIHH 158
C YQ L F+ E+HS L++ +HH
Sbjct: 1 CWQYQRLLRIFLIHAEKHSKALTLFLHH 84
>TC15539
Length = 722
Score = 25.4 bits (54), Expect = 5.8
Identities = 13/31 (41%), Positives = 20/31 (63%), Gaps = 4/31 (12%)
Frame = +2
Query: 26 VMSCRRSE---RDGRILARRLGCNKE-GHCL 52
+ SCR+ RDG + A ++GCN++ G CL
Sbjct: 128 IWSCRKGS*LCRDGGLGAGQMGCNQQ*GLCL 220
>AV412520
Length = 307
Score = 25.0 bits (53), Expect = 7.5
Identities = 11/27 (40%), Positives = 14/27 (51%)
Frame = -1
Query: 97 HPLVVSPREARQTMDEKDKKIQELTAE 123
HPL SPR+ R + E + K L E
Sbjct: 199 HPLFFSPRQFRYFLKENEGKTNSLAIE 119
>BP083541
Length = 479
Score = 25.0 bits (53), Expect = 7.5
Identities = 10/35 (28%), Positives = 23/35 (65%)
Frame = -2
Query: 87 VITKFVKDHNHPLVVSPREARQTMDEKDKKIQELT 121
VIT + + HP +V P E ++++++ K++ +L+
Sbjct: 475 VITSYALESVHPQLVVPDEVKKSVNQLLKEVVKLS 371
>TC12176 weakly similar to UP|AAQ09151 (AAQ09151) NADH dehydrogenase subunit
2, partial (6%)
Length = 478
Score = 25.0 bits (53), Expect = 7.5
Identities = 10/28 (35%), Positives = 18/28 (63%)
Frame = -3
Query: 89 TKFVKDHNHPLVVSPREARQTMDEKDKK 116
T F+K H +PL+++P+E D + K+
Sbjct: 404 TIFLKKHANPLIMNPQEKYNCED*RQKE 321
>TC11277
Length = 773
Score = 25.0 bits (53), Expect = 7.5
Identities = 9/16 (56%), Positives = 12/16 (74%)
Frame = -3
Query: 154 VKIHHIVNNLKEFEPI 169
+KIHHI+ N + F PI
Sbjct: 207 IKIHHILENCRCFRPI 160
>TC18417 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, partial (38%)
Length = 747
Score = 24.6 bits (52), Expect = 9.8
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = -2
Query: 151 KLSVKIHHIVNNLKEFEPI 169
K+S ++H +VNN E EPI
Sbjct: 668 KISNQVHSLVNNPLEVEPI 612
>TC16280
Length = 905
Score = 24.6 bits (52), Expect = 9.8
Identities = 13/37 (35%), Positives = 16/37 (43%)
Frame = +3
Query: 72 CKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQ 108
C I Y K V+ KFV+ H HP R R+
Sbjct: 24 CSE*ISWNYQKMTTCVLLKFVQCHGHPCCHQGRVPRK 134
>CB826912
Length = 597
Score = 24.6 bits (52), Expect = 9.8
Identities = 9/28 (32%), Positives = 16/28 (57%)
Frame = +1
Query: 10 KVFYDEYARHVGFVMRVMSCRRSERDGR 37
K EYA+ + + M + +C++ E GR
Sbjct: 370 KNLLQEYAQKMSYAMPLYNCKKEETPGR 453
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.135 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,983,140
Number of Sequences: 28460
Number of extensions: 37342
Number of successful extensions: 181
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 178
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 178
length of query: 176
length of database: 4,897,600
effective HSP length: 84
effective length of query: 92
effective length of database: 2,506,960
effective search space: 230640320
effective search space used: 230640320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0083.5


