

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.2
(84 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
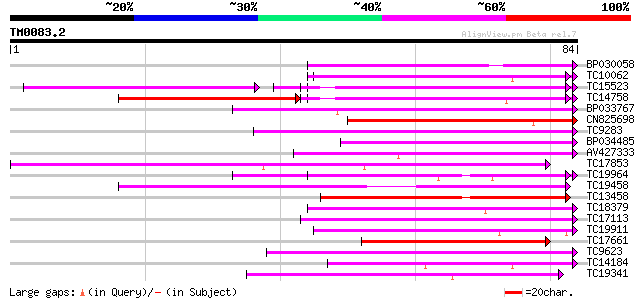
Score E
Sequences producing significant alignments: (bits) Value
BP030058 49 2e-07
TC10062 similar to BAD08700 (BAD08700) Cold shock domain protein... 46 1e-06
TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-bindi... 42 1e-05
TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding p... 42 1e-05
BP033767 40 6e-05
CN825698 40 7e-05
TC9283 39 2e-04
BP034485 39 2e-04
AV427333 39 2e-04
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 38 3e-04
TC19964 similar to UP|Q40363 (Q40363) NuM1 protein, partial (11%) 38 3e-04
TC19458 weakly similar to UP|Q9STE8 (Q9STE8) Chloroplast import-... 38 3e-04
TC13458 weakly similar to UP|Z282_HUMAN (Q9UDV7) Zinc finger pro... 38 4e-04
TC18379 similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich glycop... 38 4e-04
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 38 4e-04
TC19911 weakly similar to PIR|A44805|A44805 eggshell protein pre... 37 5e-04
TC17661 weakly similar to UP|Q09085 (Q09085) Hydroxyproline-rich... 37 5e-04
TC9623 homologue to UP|Q8H1K8 (Q8H1K8) Stromal ascorbate peroxid... 37 5e-04
TC14184 similar to UP|Q93WZ6 (Q93WZ6) Abscisic stress ripening-l... 37 6e-04
TC19341 similar to UP|Q9CAD1 (Q9CAD1) GMP synthase, partial (28%) 37 6e-04
>BP030058
Length = 455
Score = 48.9 bits (115), Expect = 2e-07
Identities = 23/40 (57%), Positives = 24/40 (59%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG D GG + G GGG GG GD GGGGG
Sbjct: 298 GGGGEGGGGGGGDGGGGGGDGHGSGGG--GGEGDGGGGGG 411
Score = 32.7 bits (73), Expect = 0.012
Identities = 18/42 (42%), Positives = 20/42 (46%), Gaps = 7/42 (16%)
Frame = +1
Query: 50 GGGG-------GSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GGGG G GG GG GGG+ GG G +G G G
Sbjct: 247 GGGGDKGLQFCGQEGYFGGGGEGGGGGGGDGGGGGGDGHGSG 372
>TC10062 similar to BAD08700 (BAD08700) Cold shock domain protein 2, partial
(66%)
Length = 481
Score = 45.8 bits (107), Expect = 1e-06
Identities = 20/39 (51%), Positives = 21/39 (53%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG + GGG G GG GG GGG GG G GGGG
Sbjct: 279 GGGAGGGGRGGGGYGGGGYGGGGYGGGGRGGGGYGGGGG 395
Score = 43.5 bits (101), Expect = 7e-06
Identities = 20/40 (50%), Positives = 20/40 (50%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGG G GG GG GGG GG G GG GG
Sbjct: 294 GGGRGGGGYGGGGYGGGGYGGGGRGGGGYGGGGGYGGSGG 413
Score = 43.1 bits (100), Expect = 9e-06
Identities = 22/41 (53%), Positives = 22/41 (53%), Gaps = 1/41 (2%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNS-GGSGDNGGGGG 84
GG GGG G GG GG GGG GGSG GGGGG
Sbjct: 309 GGGYGGGGYGGGGYGGGGRGGGGYGGGGGYGGSGGYGGGGG 431
Score = 42.4 bits (98), Expect = 2e-05
Identities = 19/39 (48%), Positives = 19/39 (48%)
Frame = +3
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
G GGG G GG GG GGG GG G GGG G
Sbjct: 267 GTRRGGGAGGGGRGGGGYGGGGYGGGGYGGGGRGGGGYG 383
Score = 39.3 bits (90), Expect = 1e-04
Identities = 20/40 (50%), Positives = 21/40 (52%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG +GG GG GG G G GGGGG
Sbjct: 366 GGGGYGGGGGYG--GSGGYGGGGGYGGGGRGGGRGGGGGG 479
Score = 39.3 bits (90), Expect = 1e-04
Identities = 20/40 (50%), Positives = 20/40 (50%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGG GG GG G G SGG G GG GG
Sbjct: 321 GGGGYGGGGYGGGGRGGGGYGGGGGYGGSGGYGGGGGYGG 440
Score = 25.8 bits (55), Expect = 1.5
Identities = 12/25 (48%), Positives = 13/25 (52%)
Frame = +3
Query: 60 NGGINDGGIGGGNSGGSGDNGGGGG 84
NG G GG +GG G GGG G
Sbjct: 249 NGQAVQGTRRGGGAGGGGRGGGGYG 323
>TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-binding,
abscisic acid-inducible protein, complete
Length = 972
Score = 42.4 bits (98), Expect = 2e-05
Identities = 22/40 (55%), Positives = 23/40 (57%)
Frame = +2
Query: 44 EGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
EGG GGGGGS GG GG GGG GG + GGGG
Sbjct: 644 EGG--YGGGGGSRYSGGGGGGYGGSGGGGYGGRREGGGGG 757
Score = 41.2 bits (95), Expect = 3e-05
Identities = 21/45 (46%), Positives = 25/45 (54%)
Frame = +2
Query: 40 IAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
+ V+E GGGG GG +GG GGG GGS +GGGGG
Sbjct: 569 VTVNEAQARGRGGGGGGGGYGGGRREGGYGGG--GGSRYSGGGGG 697
Score = 35.4 bits (80), Expect = 0.002
Identities = 19/41 (46%), Positives = 21/41 (50%), Gaps = 3/41 (7%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGI---GGGNSGGSGDNGGG 82
GG GGGG +GG GG GGG GGS + GGG
Sbjct: 665 GGSRYSGGGGGGYGGSGGGGYGGRREGGGGGYGGSREGGGG 787
Score = 35.4 bits (80), Expect = 0.002
Identities = 21/43 (48%), Positives = 21/43 (48%), Gaps = 3/43 (6%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGG---GGG 84
G SGGGGG GG GG GG GG G GG GGG
Sbjct: 668 GSRYSGGGGG----GYGGSGGGGYGGRREGGGGGYGGSREGGG 784
Score = 34.3 bits (77), Expect = 0.004
Identities = 16/36 (44%), Positives = 16/36 (44%)
Frame = +2
Query: 48 SSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
S GGGGG GG GGG G GGGG
Sbjct: 680 SGGGGGGYGGSGGGGYGGRREGGGGGYGGSREGGGG 787
Score = 33.9 bits (76), Expect(2) = 1e-05
Identities = 17/40 (42%), Positives = 17/40 (42%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGG GG GG GG GG G GG
Sbjct: 695 GGYGGSGGGGYGGRREGG--GGGYGGSREGGGGSRYSSGG 808
Score = 30.8 bits (68), Expect = 0.045
Identities = 20/44 (45%), Positives = 26/44 (58%)
Frame = +3
Query: 6 VVAAVVVVVWRSVVVVAATMVVLAAAVAVVVEVAIAVDEGGDSS 49
V AA VVV VVV T+VV+ A +AVV A+ V E D++
Sbjct: 600 VEAAAVVVDTAVVVVKVDTVVVVVAVIAVVAVEAMVVAEEEDTA 731
Score = 29.6 bits (65), Expect = 0.10
Identities = 18/42 (42%), Positives = 24/42 (56%)
Frame = +3
Query: 1 MVAIKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVVEVAIAV 42
+V +KV VVVV +VV V A +V AVVV+V + V
Sbjct: 633 VVVVKVDTVVVVVAVIAVVAVEAMVVAEEEDTAVVVKVVVVV 758
Score = 28.1 bits (61), Expect = 0.29
Identities = 17/46 (36%), Positives = 20/46 (42%)
Frame = +2
Query: 39 AIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
AI G D G ++ G GG GGG GG + G GGG
Sbjct: 527 AIEGMNGQDIDGRNVTVNEAQARGRGGGGGGGGYGGGRREGGYGGG 664
Score = 28.1 bits (61), Expect(2) = 1e-05
Identities = 17/35 (48%), Positives = 21/35 (59%)
Frame = +3
Query: 3 AIKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVVE 37
A+ V AVVVV +VVVV A + V+A VV E
Sbjct: 612 AVVVDTAVVVVKVDTVVVVVAVIAVVAVEAMVVAE 716
Score = 23.9 bits (50), Expect = 5.6
Identities = 17/35 (48%), Positives = 18/35 (50%)
Frame = +3
Query: 8 AAVVVVVWRSVVVVAATMVVLAAAVAVVVEVAIAV 42
A VVV V VVVVA VV A+ V E AV
Sbjct: 630 AVVVVKVDTVVVVVAVIAVVAVEAMVVAEEEDTAV 734
Score = 23.5 bits (49), Expect = 7.3
Identities = 14/34 (41%), Positives = 18/34 (52%)
Frame = +3
Query: 3 AIKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVV 36
A VV VVVV VVV ++ + A A+VV
Sbjct: 609 AAVVVDTAVVVVKVDTVVVVVAVIAVVAVEAMVV 710
>TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding protein 2,
partial (93%)
Length = 725
Score = 42.4 bits (98), Expect = 2e-05
Identities = 21/45 (46%), Positives = 22/45 (48%), Gaps = 5/45 (11%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGN-----SGGSGDNGGGGG 84
GG GG GG GG +GG GGG GG G GGGGG
Sbjct: 325 GGGGGGGRGGGGYGGGGGRREGGYGGGGGSRGYGGGGGGYGGGGG 459
Score = 38.5 bits (88), Expect(2) = 1e-05
Identities = 19/40 (47%), Positives = 21/40 (52%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGGG GG +GG GG G +GGGGG
Sbjct: 430 GGGGYGGGGGGG---YGGRREGGYGGDGGGSRYGSGGGGG 540
Score = 38.1 bits (87), Expect = 3e-04
Identities = 20/39 (51%), Positives = 21/39 (53%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG GGGGG G GG GGG+ GSG GGGG
Sbjct: 433 GGGYGGGGGGGYGGRREG-GYGGDGGGSRYGSGGGGGGG 546
Score = 37.7 bits (86), Expect = 4e-04
Identities = 21/40 (52%), Positives = 22/40 (54%)
Frame = +1
Query: 44 EGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
EGG GGGGGS GG GG GGG GG + G GG
Sbjct: 385 EGG--YGGGGGSRGYGGGGGGYGGGGGGGYGGRREGGYGG 498
Score = 37.0 bits (84), Expect = 6e-04
Identities = 18/40 (45%), Positives = 18/40 (45%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GGGG GG G GGG G G GGG G
Sbjct: 352 GGGYGGGGGRREGGYGGGGGSRGYGGGGGGYGGGGGGGYG 471
Score = 33.1 bits (74), Expect = 0.009
Identities = 19/40 (47%), Positives = 19/40 (47%), Gaps = 1/40 (2%)
Frame = +1
Query: 45 GGDSSGG-GGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
GG GG GGG GG GG GGG GG G GG
Sbjct: 373 GGRREGGYGGGGGSRGYGG-GGGGYGGGGGGGYGGRREGG 489
Score = 32.0 bits (71), Expect = 0.020
Identities = 17/39 (43%), Positives = 19/39 (48%), Gaps = 5/39 (12%)
Frame = +1
Query: 50 GGGGGSSDDNNGGIND-----GGIGGGNSGGSGDNGGGG 83
G G D N +N+ GG GGG GG G GGGG
Sbjct: 262 GMNGNELDGRNITVNEAQARGGGGGGGGRGGGGYGGGGG 378
Score = 29.3 bits (64), Expect = 0.13
Identities = 15/35 (42%), Positives = 17/35 (47%), Gaps = 3/35 (8%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDG---GIGGGNSGGS 76
GG GG GG + GG G G GGG GG+
Sbjct: 445 GGGGGGGYGGRREGGYGGDGGGSRYGSGGGGGGGN 549
Score = 28.1 bits (61), Expect = 0.29
Identities = 18/45 (40%), Positives = 24/45 (53%)
Frame = +2
Query: 2 VAIKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVVEVAIAVDEGG 46
VA+ + VV+ VVVV MVV+ A A+ V AV+ GG
Sbjct: 431 VAVDMEEEAAVVM---VVVVKVDMVVMVVAPAMAAAVVAAVETGG 556
Score = 23.5 bits (49), Expect(2) = 1e-05
Identities = 12/27 (44%), Positives = 18/27 (66%)
Frame = +2
Query: 17 SVVVVAATMVVLAAAVAVVVEVAIAVD 43
+ V V + + AAAV V V++A+AVD
Sbjct: 332 AAVAVVEEVDMEAAAVVVKVDMAVAVD 412
>BP033767
Length = 541
Score = 40.4 bits (93), Expect = 6e-05
Identities = 23/57 (40%), Positives = 28/57 (48%), Gaps = 6/57 (10%)
Frame = -3
Query: 34 VVVEVAIAVDEGGD------SSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
VV A AVDEGGD + GG G +++ GG GG+ G G GG GG
Sbjct: 503 VVFAAASAVDEGGDRDDGDGAGDGGNGDAEELVGGFGFGGLREDEGGDVGGEGGEGG 333
>CN825698
Length = 704
Score = 40.0 bits (92), Expect = 7e-05
Identities = 18/35 (51%), Positives = 23/35 (65%), Gaps = 1/35 (2%)
Frame = -1
Query: 51 GGGGSSDDNNGGINDGGIGGGNSGGS-GDNGGGGG 84
GGGG S+ GG + GG GGG GG+ ++G GGG
Sbjct: 206 GGGGRSESEGGGSDGGGGGGGGGGGAEAEDGDGGG 102
Score = 28.1 bits (61), Expect = 0.29
Identities = 14/35 (40%), Positives = 16/35 (45%)
Frame = -1
Query: 50 GGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG S + GG G S G G +GGGGG
Sbjct: 254 GGRSKSRSSRLSLLRAGGGGRSESEGGGSDGGGGG 150
Score = 27.7 bits (60), Expect = 0.38
Identities = 14/34 (41%), Positives = 18/34 (52%)
Frame = -1
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGS 76
D GG GGGGG+ ++ G GG SGG+
Sbjct: 167 DGGGGGGGGGGGAEAEDGDG---GGDLSSESGGA 75
Score = 26.9 bits (58), Expect = 0.66
Identities = 14/36 (38%), Positives = 18/36 (49%)
Frame = -1
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGG 81
G S GGGG G + G GGG+ S ++GG
Sbjct: 179 GGGSDGGGGGGGGGGGAEAEDGDGGGDL--SSESGG 78
>TC9283
Length = 1307
Score = 38.9 bits (89), Expect = 2e-04
Identities = 21/48 (43%), Positives = 22/48 (45%)
Frame = -2
Query: 37 EVAIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
EV A D G GGGGG S G G +SG G GGGGG
Sbjct: 211 EVIFAEDGAGGVRGGGGGVSGHFQGVGGSGATDSNDSGEPGGGGGGGG 68
>BP034485
Length = 485
Score = 38.9 bits (89), Expect = 2e-04
Identities = 17/35 (48%), Positives = 19/35 (53%)
Frame = +3
Query: 50 GGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GGGGG + + N N GG N G G GGGGG
Sbjct: 153 GGGGGETANKNKNKNKNNGGGNNKGKEGKKGGGGG 257
Score = 32.7 bits (73), Expect = 0.012
Identities = 18/40 (45%), Positives = 19/40 (47%), Gaps = 1/40 (2%)
Frame = +3
Query: 46 GDSSGGGGGSSDDNNGGINDGGI-GGGNSGGSGDNGGGGG 84
G GGGGG D + D I G SGG G GGG G
Sbjct: 234 GKKGGGGGGKLDKIDFDFQDFDIPSHGKSGGKGGGGGGKG 353
Score = 30.8 bits (68), Expect = 0.045
Identities = 18/47 (38%), Positives = 19/47 (40%), Gaps = 9/47 (19%)
Frame = +3
Query: 46 GDSSGGGGGSS---------DDNNGGINDGGIGGGNSGGSGDNGGGG 83
G GGGGG S D N GG +G G G GGGG
Sbjct: 27 GKQKGGGGGDSSSDKKKKTKDKNGGGFLVKFLGLGKKSKKGGGGGGG 167
Score = 30.8 bits (68), Expect = 0.045
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = +3
Query: 50 GGGGGSSDDNNGGINDGGIGGGNSGGSG 77
GGGGG + NG + GIGG +G G
Sbjct: 330 GGGGGGKGNKNGHDHGHGIGGNMNGNGG 413
Score = 30.4 bits (67), Expect = 0.059
Identities = 14/39 (35%), Positives = 17/39 (42%)
Frame = +3
Query: 44 EGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGG 82
+GG GG + + N N GG G G G GGG
Sbjct: 144 KGGGGGGGETANKNKNKNKNNGGGNNKGKEGKKGGGGGG 260
>AV427333
Length = 387
Score = 38.5 bits (88), Expect = 2e-04
Identities = 20/43 (46%), Positives = 25/43 (57%), Gaps = 1/43 (2%)
Frame = -3
Query: 43 DEGGDSSGGGGGSS-DDNNGGINDGGIGGGNSGGSGDNGGGGG 84
D GG + GGGG + DD GG + GG G GG+ + GGGG
Sbjct: 298 DGGGRTCGGGGVNG*DDGGGGDDGGGEGCPRGGGANE*EGGGG 170
Score = 33.9 bits (76), Expect = 0.005
Identities = 18/50 (36%), Positives = 22/50 (44%), Gaps = 11/50 (22%)
Frame = -3
Query: 46 GDSSGGGGGSSDDNNGGI-----------NDGGIGGGNSGGSGDNGGGGG 84
G+ GGGG G I GG+ G + GG GD+GGG G
Sbjct: 364 GEGGSGGGGGRTCGGGVIG*CGDGGGRTCGGGGVNG*DDGGGGDDGGGEG 215
Score = 30.0 bits (66), Expect = 0.078
Identities = 18/54 (33%), Positives = 20/54 (36%), Gaps = 13/54 (24%)
Frame = -3
Query: 44 EGGDSSGGG-------------GGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
EGG GGG GG GG+N GGG G G+ GG
Sbjct: 361 EGGSGGGGGRTCGGGVIG*CGDGGGRTCGGGGVNG*DDGGGGDDGGGEGCPRGG 200
Score = 30.0 bits (66), Expect = 0.078
Identities = 21/57 (36%), Positives = 26/57 (44%), Gaps = 15/57 (26%)
Frame = -3
Query: 43 DEGGDSSGGG------GGSSDDNNGGIND----GGIGGGNSGGSGDNGG-----GGG 84
D+GG GG GG +++ GG D G+GGG G G GG GGG
Sbjct: 253 DDGGGGDDGGGEGCPRGGGANE*EGGGGDLK*WEGVGGGLKGWEGGVGGL*GWEGGG 83
Score = 28.1 bits (61), Expect = 0.29
Identities = 19/44 (43%), Positives = 20/44 (45%), Gaps = 3/44 (6%)
Frame = -3
Query: 44 EGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGG---GGG 84
EG SGGGGG + GGG G GD GG GGG
Sbjct: 367 EGEGGSGGGGGRT-----------CGGGVIG*CGDGGGRTCGGG 269
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 38.1 bits (87), Expect = 3e-04
Identities = 27/89 (30%), Positives = 40/89 (44%), Gaps = 9/89 (10%)
Frame = +3
Query: 1 MVAIKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVV---EVAIAVDEGGDSSGG------ 51
+V I ++ A+V+V VVV V + V++V+ V + GG GG
Sbjct: 339 VVVIVMIVAIVIVAEAMEVVVDLMEVNALSVVSLVILLGSVLVKGKRGGGRYGGRESRXS 518
Query: 52 GGGSSDDNNGGINDGGIGGGNSGGSGDNG 80
GGG D N + GG + G GD+G
Sbjct: 519 GGGYGPDRNADRSSGGRSSRDGGSHGDSG 605
>TC19964 similar to UP|Q40363 (Q40363) NuM1 protein, partial (11%)
Length = 562
Score = 38.1 bits (87), Expect = 3e-04
Identities = 21/43 (48%), Positives = 23/43 (52%), Gaps = 4/43 (9%)
Frame = +2
Query: 45 GGDSSGGGGGSSDDNNGG----INDGGIGGGNSGGSGDNGGGG 83
GG S GG GG D GG GG GG+ GG G +GGGG
Sbjct: 158 GGRSGGGRGGRFDSGRGGGRGRFGSGG-RGGDRGGRGGSGGGG 283
Score = 37.4 bits (85), Expect = 5e-04
Identities = 22/56 (39%), Positives = 25/56 (44%), Gaps = 5/56 (8%)
Frame = +2
Query: 34 VVVEVAIAVDEGGDSSGGGGGSSDDNNGGINDGGIGG-----GNSGGSGDNGGGGG 84
+ V+ A D G GGG S GG D G GG G+ G GD GG GG
Sbjct: 101 LAVDEAKPRDNQGSGGRSGGGRSGGGRGGRFDSGRGGGRGRFGSGGRGGDRGGRGG 268
>TC19458 weakly similar to UP|Q9STE8 (Q9STE8) Chloroplast import-associated
channel homolog (AT3g46740/T6H20_230), partial (10%)
Length = 469
Score = 38.1 bits (87), Expect = 3e-04
Identities = 24/67 (35%), Positives = 33/67 (48%)
Frame = +3
Query: 17 SVVVVAATMVVLAAAVAVVVEVAIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGS 76
S+ +T V L++A A +V + + + GGGG GG GG GGG GG
Sbjct: 120 SLFKTISTTVALSSAAASIVLHSTPLTPFLSTGGGGG-------GGTTGGGGGGGWFGGG 278
Query: 77 GDNGGGG 83
G +G GG
Sbjct: 279 GGDGNGG 299
Score = 32.3 bits (72), Expect = 0.016
Identities = 19/49 (38%), Positives = 24/49 (48%)
Frame = -3
Query: 36 VEVAIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
+ VA A E +S GG+S + G GG GG D+GGGGG
Sbjct: 293 ISVASAAAEPTSTSSSAGGASTSASAR----GEEGGERGGVEDDGGGGG 159
Score = 28.1 bits (61), Expect = 0.29
Identities = 13/21 (61%), Positives = 14/21 (65%), Gaps = 2/21 (9%)
Frame = +3
Query: 66 GGIGGGNSGGSGDNG--GGGG 84
GG GGG +GG G G GGGG
Sbjct: 219 GGGGGGTTGGGGGGGWFGGGG 281
Score = 27.7 bits (60), Expect = 0.38
Identities = 12/36 (33%), Positives = 15/36 (41%)
Frame = -3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNG 80
G +S G GG+ D G GGG G +G
Sbjct: 239 GASTSASARGEEGGERGGVEDDGGGGGGECDGGRDG 132
Score = 26.9 bits (58), Expect = 0.66
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = +3
Query: 69 GGGNSGGSGDNGGGGG 84
GGG GG+ GGGGG
Sbjct: 216 GGGGGGGTTGGGGGGG 263
>TC13458 weakly similar to UP|Z282_HUMAN (Q9UDV7) Zinc finger protein 282
(HTLV-I U5RE binding protein 1) (HUB-1), partial (3%)
Length = 810
Score = 37.7 bits (86), Expect = 4e-04
Identities = 18/37 (48%), Positives = 23/37 (61%)
Frame = -1
Query: 47 DSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
D +GGGG + D +GG G GGG+ GG +GGGG
Sbjct: 384 DGAGGGGQYTVDGDGGFFGDG-GGGDGGGIAGDGGGG 277
Score = 25.8 bits (55), Expect = 1.5
Identities = 17/45 (37%), Positives = 21/45 (45%), Gaps = 2/45 (4%)
Frame = -1
Query: 42 VDEGGDSSGGGGGSSDDNNGGINDGG--IGGGNSGGSGDNGGGGG 84
V G +S GG D++ G GG G+ G GD GGG G
Sbjct: 441 VSMGYNSPGGPVMYMYDDDDGAGGGGQYTVDGDGGFFGDGGGGDG 307
>TC18379 similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich glycoprotein
DZ-HRGP precursor, partial (8%)
Length = 584
Score = 37.7 bits (86), Expect = 4e-04
Identities = 21/44 (47%), Positives = 23/44 (51%), Gaps = 4/44 (9%)
Frame = -1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIG----GGNSGGSGDNGGGGG 84
GG GG GG D GG G G GG +GG+G GGGGG
Sbjct: 386 GGAGLGGRGGKVDVKRGGP*GRGGGAAQYGGGAGGAGAGGGGGG 255
Score = 29.3 bits (64), Expect = 0.13
Identities = 15/32 (46%), Positives = 16/32 (49%)
Frame = -1
Query: 51 GGGGSSDDNNGGINDGGIGGGNSGGSGDNGGG 82
G GG + GG G GGG GG G GGG
Sbjct: 326 GRGGGAAQYGGGAGGAGAGGG--GGGGAIGGG 237
Score = 27.7 bits (60), Expect = 0.38
Identities = 16/41 (39%), Positives = 18/41 (43%), Gaps = 1/41 (2%)
Frame = -1
Query: 40 IAVDEGGD-SSGGGGGSSDDNNGGINDGGIGGGNSGGSGDN 79
+ V GG GGG GG GG GGG + G G N
Sbjct: 353 VDVKRGGP*GRGGGAAQYGGGAGGAGAGGGGGGGAIGGG*N 231
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 37.7 bits (86), Expect = 4e-04
Identities = 18/41 (43%), Positives = 20/41 (47%)
Frame = -2
Query: 44 EGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
+GG + GGGGG GI GG G GG D GG G
Sbjct: 788 KGGKNGGGGGGGGGGVKPGITIGGKGNKGEGGCFDGGGWNG 666
Score = 34.7 bits (78), Expect = 0.003
Identities = 20/43 (46%), Positives = 23/43 (52%), Gaps = 3/43 (6%)
Frame = -2
Query: 45 GGDSSGGGGGSSDD---NNGGINDGGIGGGNSGGSGDNGGGGG 84
GG + G GG D N GIN GG+GGG +G GG GG
Sbjct: 722 GGKGNKGEGGCFDGGGWNGLGINGGGLGGG*NGLGISGGGLGG 594
Score = 33.5 bits (75), Expect = 0.007
Identities = 14/22 (63%), Positives = 15/22 (67%)
Frame = -2
Query: 61 GGINDGGIGGGNSGGSGDNGGG 82
GG N+GG GG N GG G GGG
Sbjct: 809 GGGNNGGKGGKNGGGGGGGGGG 744
Score = 32.7 bits (73), Expect = 0.012
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 7/49 (14%)
Frame = -2
Query: 42 VDEGGDSSGGGGGSSDDNNGGINDGGIGGGNS-------GGSGDNGGGG 83
V+ GG GG G NGG GG GGG GG G+ G GG
Sbjct: 830 VEIGGLPGGGNNGGKGGKNGG---GGGGGGGGVKPGITIGGKGNKGEGG 693
Score = 32.3 bits (72), Expect = 0.016
Identities = 21/47 (44%), Positives = 24/47 (50%), Gaps = 7/47 (14%)
Frame = -2
Query: 45 GGDSSGG-------GGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
GG GG GGG N G + GG+GG N GSG +GGG G
Sbjct: 653 GGGLGGG*NGLGISGGGLGG*NGLGKSGGGLGGWN--GSGKSGGGLG 519
Score = 32.3 bits (72), Expect = 0.016
Identities = 13/24 (54%), Positives = 15/24 (62%)
Frame = -2
Query: 61 GGINDGGIGGGNSGGSGDNGGGGG 84
GG+ GG GG G +G GGGGG
Sbjct: 821 GGLPGGGNNGGKGGKNGGGGGGGG 750
Score = 31.6 bits (70), Expect = 0.027
Identities = 23/50 (46%), Positives = 26/50 (52%), Gaps = 9/50 (18%)
Frame = -2
Query: 44 EGGDSSGGG--------GGSSDDNNG-GINDGGIGGGNSGGSGDNGGGGG 84
EGG GGG GG NG GI+ GG+GG N G G +GGG G
Sbjct: 701 EGGCFDGGGWNGLGINGGGLGGG*NGLGISGGGLGG*N--GLGKSGGGLG 558
Score = 30.4 bits (67), Expect = 0.059
Identities = 19/41 (46%), Positives = 22/41 (53%), Gaps = 4/41 (9%)
Frame = -2
Query: 46 GDSSGGGGG--SSDDNNGGINDGGIG--GGNSGGSGDNGGG 82
G S GG GG S + GG+ G+G GG G G NGGG
Sbjct: 581 GKSGGGLGGWNGSGKSGGGLG*KGLGIRGGG*NGLGINGGG 459
Score = 25.4 bits (54), Expect = 1.9
Identities = 13/27 (48%), Positives = 13/27 (48%)
Frame = -2
Query: 51 GGGGSSDDNNGGINDGGIGGGNSGGSG 77
GGG NGG N GI GG G G
Sbjct: 467 GGG*KGLGTNGG*NGLGIRGGGKNGIG 387
>TC19911 weakly similar to PIR|A44805|A44805 eggshell protein precursor -
fluke (Schistosoma haematobium) {Schistosoma
haematobium;} , partial (24%)
Length = 350
Score = 37.4 bits (85), Expect = 5e-04
Identities = 20/43 (46%), Positives = 25/43 (57%), Gaps = 4/43 (9%)
Frame = +1
Query: 46 GDSSGGGGGSSDDNNGGINDGGIGGG--NSGGSGDNGG--GGG 84
G GGG D+ +GG GG G G ++GG GD+GG GGG
Sbjct: 115 GHGLSGGGYDDDNRSGGRTGGGYGDGGHSAGGYGDHGGRSGGG 243
Score = 36.2 bits (82), Expect = 0.001
Identities = 20/46 (43%), Positives = 23/46 (49%), Gaps = 4/46 (8%)
Frame = +1
Query: 43 DEGGDSSGGGGGSSDDNNGGIND-GGIGGGNSGGSGDNG---GGGG 84
+ G +GGG G + GG D GG GG GG GD G G GG
Sbjct: 151 NRSGGRTGGGYGDGGHSAGGYGDHGGRSGGGGGGYGDGGLSSGSGG 288
Score = 35.4 bits (80), Expect = 0.002
Identities = 19/41 (46%), Positives = 23/41 (55%)
Frame = +1
Query: 43 DEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
D GG S GGGGG +GG++ G G G+ G SG G G
Sbjct: 217 DHGGRSGGGGGG---YGDGGLSSGSGGYGDGGRSGVGYGDG 330
Score = 34.3 bits (77), Expect = 0.004
Identities = 20/43 (46%), Positives = 23/43 (52%), Gaps = 3/43 (6%)
Frame = +1
Query: 44 EGGDSSGG---GGGSSDDNNGGINDGGIGGGNSGGSGDNGGGG 83
+GG S+GG GG S GG DGG+ S GSG G GG
Sbjct: 187 DGGHSAGGYGDHGGRSGGGGGGYGDGGL----SSGSGGYGDGG 303
Score = 28.1 bits (61), Expect = 0.29
Identities = 17/41 (41%), Positives = 19/41 (45%), Gaps = 2/41 (4%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGG--SGDNGGGG 83
GG S G SD+ +GG G G SGG DN GG
Sbjct: 43 GGGSHGNDHRHSDNEHGGRRSETTGHGLSGGGYDDDNRSGG 165
Score = 25.0 bits (53), Expect = 2.5
Identities = 12/38 (31%), Positives = 17/38 (44%)
Frame = +1
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGG 82
G GGG +D+ N+ G + G G +GGG
Sbjct: 25 GNTGHSGGGSHGNDHRHSDNEHGGRRSETTGHGLSGGG 138
Score = 24.3 bits (51), Expect = 4.3
Identities = 14/36 (38%), Positives = 21/36 (57%)
Frame = +2
Query: 1 MVAIKVVAAVVVVVWRSVVVVAATMVVLAAAVAVVV 36
M+ I V+AAV VV ++ A M ++ +AVVV
Sbjct: 140 MMMITVLAAVPAVVTAMEDILLAAMGIMGVDLAVVV 247
>TC17661 weakly similar to UP|Q09085 (Q09085) Hydroxyproline-rich
glycoprotein (HRGP) (Fragment), partial (8%)
Length = 417
Score = 37.4 bits (85), Expect = 5e-04
Identities = 15/28 (53%), Positives = 18/28 (63%)
Frame = +3
Query: 53 GGSSDDNNGGINDGGIGGGNSGGSGDNG 80
GG+ D+NNGG GG GG G GD+G
Sbjct: 282 GGNDDNNNGGDGSGGGGGDEGSGGGDDG 365
Score = 35.4 bits (80), Expect = 0.002
Identities = 16/31 (51%), Positives = 18/31 (57%)
Frame = +3
Query: 54 GSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
G +DDNN G G G+ GG GD G GGG
Sbjct: 282 GGNDDNNNG------GDGSGGGGGDEGSGGG 356
Score = 31.6 bits (70), Expect = 0.027
Identities = 14/24 (58%), Positives = 18/24 (74%)
Frame = +3
Query: 43 DEGGDSSGGGGGSSDDNNGGINDG 66
+ GGD SGGGGG D+ +GG +DG
Sbjct: 300 NNGGDGSGGGGG--DEGSGGGDDG 365
Score = 23.1 bits (48), Expect = 9.5
Identities = 8/17 (47%), Positives = 11/17 (64%)
Frame = +3
Query: 68 IGGGNSGGSGDNGGGGG 84
IGG + +G +G GGG
Sbjct: 279 IGGNDDNNNGGDGSGGG 329
>TC9623 homologue to UP|Q8H1K8 (Q8H1K8) Stromal ascorbate peroxidase,
partial (42%)
Length = 640
Score = 37.4 bits (85), Expect = 5e-04
Identities = 19/46 (41%), Positives = 24/46 (51%)
Frame = -3
Query: 39 AIAVDEGGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGGGGG 84
A A++ G GGGGG ++ GG+ G G SGG G N GG
Sbjct: 209 AQALEGGTGRGGGGGGE*EEGTGGV*GGRE*EGGSGGGGRNHASGG 72
>TC14184 similar to UP|Q93WZ6 (Q93WZ6) Abscisic stress ripening-like
protein, partial (57%)
Length = 1141
Score = 37.0 bits (84), Expect = 6e-04
Identities = 23/49 (46%), Positives = 24/49 (48%), Gaps = 12/49 (24%)
Frame = +3
Query: 48 SSGGGGGSSDDNN----------GGINDGGIGGGNS--GGSGDNGGGGG 84
+SGGGG S N GG GG GGG S GG G GGGGG
Sbjct: 300 NSGGGGYDSGYNKTAYSTDEPSGGGYGTGGAGGGYSTGGGYGTGGGGGG 446
Score = 34.7 bits (78), Expect = 0.003
Identities = 17/36 (47%), Positives = 20/36 (55%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNG 80
GG S+GGG G+ G + GG GGG S G G G
Sbjct: 396 GGYSTGGGYGTGGGGGGEYSTGGGGGGYSTGGGGYG 503
Score = 33.9 bits (76), Expect = 0.005
Identities = 23/56 (41%), Positives = 27/56 (48%), Gaps = 8/56 (14%)
Frame = +3
Query: 37 EVAIAVDE----GGDSSGGGGGSSDDNNGGINDGGIGGG----NSGGSGDNGGGGG 84
+ A + DE G + G GGG S GG GG GGG GG G + GGGG
Sbjct: 336 KTAYSTDEPSGGGYGTGGAGGGYS--TGGGYGTGGGGGGEYSTGGGGGGYSTGGGG 497
Score = 29.6 bits (65), Expect = 0.10
Identities = 14/36 (38%), Positives = 18/36 (49%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNG 80
GG GGGG + + GG G GG G+ D+G
Sbjct: 411 GGGYGTGGGGGGEYSTGGGGGGYSTGGGGYGNDDSG 518
Score = 27.3 bits (59), Expect = 0.50
Identities = 14/37 (37%), Positives = 16/37 (42%)
Frame = +3
Query: 45 GGDSSGGGGGSSDDNNGGINDGGIGGGNSGGSGDNGG 81
G + GGGGG GG GG G G D+ G
Sbjct: 417 GYGTGGGGGGEYSTGGGG---GGYSTGGGGYGNDDSG 518
>TC19341 similar to UP|Q9CAD1 (Q9CAD1) GMP synthase, partial (28%)
Length = 484
Score = 37.0 bits (84), Expect = 6e-04
Identities = 19/49 (38%), Positives = 27/49 (54%), Gaps = 2/49 (4%)
Frame = -3
Query: 36 VEVAIAVDEGGDSSGGGGGSSDDNNGGIN--DGGIGGGNSGGSGDNGGG 82
VE A+ G S G G + +D++GG+ DGG GGG G++G G
Sbjct: 176 VEEAVREGRGVGSVDGVGAAGEDDDGGVEFGDGGEGGGAGDAEGEDGEG 30
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.310 0.136 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,126,805
Number of Sequences: 28460
Number of extensions: 19669
Number of successful extensions: 1990
Number of sequences better than 10.0: 533
Number of HSP's better than 10.0 without gapping: 875
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1568
length of query: 84
length of database: 4,897,600
effective HSP length: 60
effective length of query: 24
effective length of database: 3,190,000
effective search space: 76560000
effective search space used: 76560000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0083.2


