

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.16
(294 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
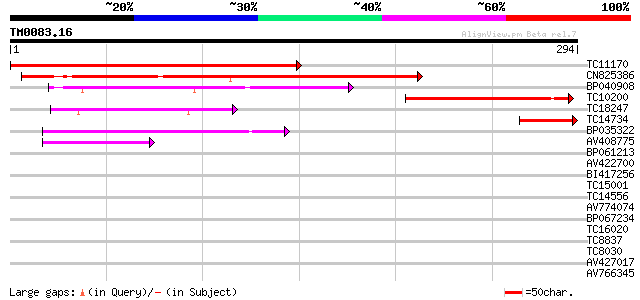
Sequences producing significant alignments: (bits) Value
TC11170 weakly similar to UP|Q9FKT4 (Q9FKT4) RNA-binding protein... 300 1e-82
CN825386 177 2e-45
BP040908 126 4e-30
TC10200 similar to UP|AAR01750 (AAR01750) Expressed protein, par... 119 6e-28
TC18247 similar to UP|BTG1_HUMAN (P31607) BTG1 protein (B-cell t... 72 1e-13
TC14734 UP|Q9FKT4 (Q9FKT4) RNA-binding protein-like, partial (9%) 64 2e-11
BP035322 55 2e-08
AV408775 47 5e-06
BP061213 32 0.097
AV422700 30 0.48
BI417256 28 1.8
TC15001 similar to UP|GLE1_YEAST (Q12315) RNA export factor GLE1... 28 2.4
TC14556 homologue to UP|Q9ZTW5 (Q9ZTW5) GDP-mannose pyrophosphor... 27 3.1
AV774074 27 4.1
BP067234 27 5.3
TC16020 similar to UP|Q9SLX0 (Q9SLX0) Importin alpha 1b, partial... 27 5.3
TC8837 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein ki... 26 7.0
TC8030 26 7.0
AV427017 26 7.0
AV766345 26 9.1
>TC11170 weakly similar to UP|Q9FKT4 (Q9FKT4) RNA-binding protein-like,
partial (32%)
Length = 613
Score = 300 bits (769), Expect = 1e-82
Identities = 151/151 (100%), Positives = 151/151 (100%)
Frame = +2
Query: 1 MMSSSGAGRYMAFPPSPSAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVL 60
MMSSSGAGRYMAFPPSPSAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVL
Sbjct: 161 MMSSSGAGRYMAFPPSPSAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVL 340
Query: 61 PHCYRLINQEILRVTTLLGNASVLGQSRLEHASPLATGGIFSNGGADVNGWASRFQSEMP 120
PHCYRLINQEILRVTTLLGNASVLGQSRLEHASPLATGGIFSNGGADVNGWASRFQSEMP
Sbjct: 341 PHCYRLINQEILRVTTLLGNASVLGQSRLEHASPLATGGIFSNGGADVNGWASRFQSEMP 520
Query: 121 SLLQSSATPSWLSPQGSSSGLVVKKTMRVDI 151
SLLQSSATPSWLSPQGSSSGLVVKKTMRVDI
Sbjct: 521 SLLQSSATPSWLSPQGSSSGLVVKKTMRVDI 613
>CN825386
Length = 670
Score = 177 bits (449), Expect = 2e-45
Identities = 105/215 (48%), Positives = 131/215 (60%), Gaps = 7/215 (3%)
Frame = +3
Query: 7 AGRYMAFPP-SPSAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVLPHCYR 65
+G Y FPP P PPSP +RS SS +++++YL ELL ER LGPF+ VLP C R
Sbjct: 48 SGTYFQFPPPGPGLPPSP----IRS--SSIPSDRERYLAELLAERQKLGPFVQVLPQCTR 209
Query: 66 LINQEILRVTTLLGNASVLGQSRLEHASPLATGGIFSNGGA-DVNGWAS---RFQSEMPS 121
LI QEI R+T N + R E SP + N D+ GW + +
Sbjct: 210 LITQEIRRITGF--NQGFIEHDRPESDSPFRSLSQHPNSRPMDLEGWPAVPIEDNGNLQR 383
Query: 122 LLQSSATP-SWLSPQGSSSGLVVKKTMRVDIPVEVYPN-FNFVGRLLGPRGNSLKRVEAS 179
+ A P W QG + VVK+ +R+D+PVE YPN +NFVGR+LGPRGNSLKRVEA
Sbjct: 384 IASFQAPPMGWPGTQGVPTTPVVKRVIRLDVPVEKYPNQYNFVGRILGPRGNSLKRVEAM 563
Query: 180 TDCRVLIRGRGSIKDPAKEEMMRGKPGYEHLNEPL 214
T+CRV IRG GS+KD KEE ++ KPGYEHLNEPL
Sbjct: 564 TECRVYIRGCGSVKDSIKEEKLKDKPGYEHLNEPL 668
>BP040908
Length = 523
Score = 126 bits (317), Expect = 4e-30
Identities = 79/168 (47%), Positives = 100/168 (59%), Gaps = 10/168 (5%)
Frame = -3
Query: 21 PSPHLSGLRSAASSAL------AEQD-KYLTELLGERHILGPFMAVLPHCYRLINQEILR 73
PSP +R+AA+SA +E D +YL+ELL ER LGPFM VLP C RL+NQEILR
Sbjct: 521 PSP----VRAAAASAQIRGTPSSEMDSQYLSELLAERQKLGPFMQVLPTCNRLLNQEILR 354
Query: 74 VTTLLGNASVLGQSRLEHASP--LATGGIFSN-GGADVNGWASRFQSEMPSLLQSSATPS 130
V+ +L N RL H SP +A + SN G + GW Q + T
Sbjct: 353 VSGMLSNQGFGDFDRLRHRSPSPMAPSNLMSNVPGTGLGGWNGLQQERLCGT--PGMTMD 180
Query: 131 WLSPQGSSSGLVVKKTMRVDIPVEVYPNFNFVGRLLGPRGNSLKRVEA 178
W S + V++ +R++IPV+ +PNFNFVGRLLGPRGNSLKRVEA
Sbjct: 179 WQVAPASPTSYTVRRILRLEIPVDTFPNFNFVGRLLGPRGNSLKRVEA 36
>TC10200 similar to UP|AAR01750 (AAR01750) Expressed protein, partial (30%)
Length = 615
Score = 119 bits (298), Expect = 6e-28
Identities = 58/87 (66%), Positives = 71/87 (80%)
Frame = +3
Query: 206 GYEHLNEPLHVLVEAEFPAEIIDARLMQAREILEDLLKPVDESQDYYKKQQLRELALLNG 265
GYEH +EPLH+L+EAE PA I+D RL QA+EI+E++LKPVDESQD+YK+QQLRE A+LN
Sbjct: 3 GYEHXSEPLHILIEAELPAXIVDVRLRQAQEIIEEILKPVDESQDFYKRQQLRERAMLNS 182
Query: 266 TLREEGSPMSGSVSPFHNSLGMKRAKT 292
REE +SGSVSPF S +KRAKT
Sbjct: 183 NFREESPQLSGSVSPF-TSNEIKRAKT 260
>TC18247 similar to UP|BTG1_HUMAN (P31607) BTG1 protein (B-cell
translocation gene 1 protein), partial (8%)
Length = 543
Score = 71.6 bits (174), Expect = 1e-13
Identities = 45/102 (44%), Positives = 57/102 (55%), Gaps = 5/102 (4%)
Frame = +2
Query: 22 SPHLSGLRSAASS--ALAEQDKYLTELLGERHILGPFMAVLPHCYRLINQEILRVTTLLG 79
+P+ S R+A+ + E +YL ELL E LGPF VLP C RLINQEILRV+ +L
Sbjct: 170 NPNYSPARAASPQIRSTPEDSQYLQELLAEHQKLGPFTQVLPICNRLINQEILRVSRMLS 349
Query: 80 NASVLGQSRLEH--ASPLATGGIFSN-GGADVNGWASRFQSE 118
N RL H SP+A+ I SN G + GW S Q +
Sbjct: 350 NQGFGDFDRLRHRSPSPMASSNIMSNVSGTGLGGWNSLQQEQ 475
>TC14734 UP|Q9FKT4 (Q9FKT4) RNA-binding protein-like, partial (9%)
Length = 598
Score = 64.3 bits (155), Expect = 2e-11
Identities = 30/30 (100%), Positives = 30/30 (100%)
Frame = +2
Query: 265 GTLREEGSPMSGSVSPFHNSLGMKRAKTRG 294
GTLREEGSPMSGSVSPFHNSLGMKRAKTRG
Sbjct: 2 GTLREEGSPMSGSVSPFHNSLGMKRAKTRG 91
>BP035322
Length = 556
Score = 54.7 bits (130), Expect = 2e-08
Identities = 42/129 (32%), Positives = 57/129 (43%), Gaps = 1/129 (0%)
Frame = +1
Query: 18 SAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVLPHCYRLINQEILRVTTL 77
S+P S + S +YL ELL E L PFM VLP C RL+NQEILR +
Sbjct: 172 SSPSSQRANSPNINMRSNFDVDSQYLAELLAEHQKLRPFMQVLPLCSRLLNQEILRASGK 351
Query: 78 LGNASVLGQSRLEHASPLATGGIFSNG-GADVNGWASRFQSEMPSLLQSSATPSWLSPQG 136
G G S + L+ G + S+ + +GW S + + Q W +P
Sbjct: 352 NGMLQNQGFSDFDRLQVLSPGHMTSSELTPNFSGWNSLSHEQRLAGAQ-GLNMDWQAPPA 528
Query: 137 SSSGLVVKK 145
S +VKK
Sbjct: 529 VPSSHIVKK 555
>AV408775
Length = 427
Score = 46.6 bits (109), Expect = 5e-06
Identities = 27/58 (46%), Positives = 32/58 (54%)
Frame = +1
Query: 18 SAPPSPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVLPHCYRLINQEILRVT 75
S+P + S S +YL ELL E L PFM VLP C RL+NQEILRV+
Sbjct: 205 SSPATARASSPNINIRSNFNVDSQYLAELLEEYQKLRPFMQVLPLCTRLLNQEILRVS 378
>BP061213
Length = 498
Score = 32.3 bits (72), Expect = 0.097
Identities = 17/39 (43%), Positives = 19/39 (48%)
Frame = -3
Query: 13 FPPSPSAPPSPHLSGLRSAASSALAEQDKYLTELLGERH 51
FPP P APP P G AAS+ + Y T L E H
Sbjct: 202 FPPPPEAPPHPPTPGTL*AASN*CSHLSPYTTSLT*ELH 86
>AV422700
Length = 472
Score = 30.0 bits (66), Expect = 0.48
Identities = 13/21 (61%), Positives = 15/21 (70%)
Frame = +3
Query: 14 PPSPSAPPSPHLSGLRSAASS 34
PPSP+APP H G R A+SS
Sbjct: 42 PPSPAAPPLLHCCGARLASSS 104
>BI417256
Length = 449
Score = 28.1 bits (61), Expect = 1.8
Identities = 15/40 (37%), Positives = 22/40 (54%)
Frame = +2
Query: 22 SPHLSGLRSAASSALAEQDKYLTELLGERHILGPFMAVLP 61
S +GL + +A EQD Y+ L E H++ P M V+P
Sbjct: 164 SARQAGLLRYSYAASEEQDLYVAGDLYENHVVTPAMIVVP 283
>TC15001 similar to UP|GLE1_YEAST (Q12315) RNA export factor GLE1, partial
(4%)
Length = 988
Score = 27.7 bits (60), Expect = 2.4
Identities = 28/95 (29%), Positives = 47/95 (49%), Gaps = 8/95 (8%)
Frame = +2
Query: 173 LKRVEASTDCRVLIRGRGSIKDPAKEEMM------RGKPGYEHLNEPLHVLVEAEFPAEI 226
LK +E T RV R +++ E + R + G + L + V +E E A +
Sbjct: 377 LKLIEEETAKRVEEAIRKRVEESLNSEEVQVEIQRRLEEGRKRLIVEVAVQLEKEKEAAL 556
Query: 227 IDARLM--QAREILEDLLKPVDESQDYYKKQQLRE 259
I+AR QAR+ EDL + ++E++ ++ Q RE
Sbjct: 557 IEAREKEEQARKEKEDLDRMLEENRKKIEEAQRRE 661
>TC14556 homologue to UP|Q9ZTW5 (Q9ZTW5) GDP-mannose pyrophosphorylase,
complete
Length = 1521
Score = 27.3 bits (59), Expect = 3.1
Identities = 31/110 (28%), Positives = 50/110 (45%), Gaps = 28/110 (25%)
Frame = +3
Query: 24 HLSGLRSAASSALAEQDKYL-------TELLGERHILGPFMAVLPHC-----YRLINQEI 71
+L LR +SS LA + T +GE ++GP +A+ P C RL I
Sbjct: 855 YLDSLRKKSSSKLASGSNIVGNVIVDETAKIGEGCLIGPDVAIGPGCIVESGVRLSRCTI 1034
Query: 72 LR----------VTTLLGNASVLGQ-SRLEHASPL-----ATGGIFSNGG 105
+R ++++G S +GQ +R+E+ + L I+SNGG
Sbjct: 1035MRGVRIKKHACISSSIIGWHSTVGQWARVENMTILGEDVHVCDEIYSNGG 1184
>AV774074
Length = 434
Score = 26.9 bits (58), Expect = 4.1
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = +3
Query: 16 SPSAPPSPHLSGLRSAASSALA 37
SPSAP PH +RSAA + A
Sbjct: 333 SPSAPSDPHQRTMRSAAGTTTA 398
>BP067234
Length = 493
Score = 26.6 bits (57), Expect = 5.3
Identities = 17/49 (34%), Positives = 23/49 (46%), Gaps = 1/49 (2%)
Frame = -3
Query: 8 GRYMAFPPSPSAPPSP-HLSGLRSAASSALAEQDKYLTELLGERHILGP 55
G+++AFP +P SP H + A Q+K T LLG H P
Sbjct: 377 GKFLAFPGNPPRYQSPYHRRSSWCGWN*PSAPQEKGCTHLLGASHPRNP 231
>TC16020 similar to UP|Q9SLX0 (Q9SLX0) Importin alpha 1b, partial (55%)
Length = 951
Score = 26.6 bits (57), Expect = 5.3
Identities = 24/62 (38%), Positives = 28/62 (44%), Gaps = 1/62 (1%)
Frame = -3
Query: 68 NQEILRVTTLLGNASVLGQSRLEHASPLATGGIFSNGGADVNGWASRFQS-EMPSLLQSS 126
NQ + R+ T SVL S L IFSN A WA RF S E+P L +S
Sbjct: 862 NQYVSRIFTAFS*ISVL--------S*L*RFSIFSNPSASSIIWAYRFTSPELPVFLSAS 707
Query: 127 AT 128
T
Sbjct: 706 PT 701
>TC8837 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(15%)
Length = 606
Score = 26.2 bits (56), Expect = 7.0
Identities = 10/13 (76%), Positives = 11/13 (83%)
Frame = -3
Query: 14 PPSPSAPPSPHLS 26
PPSPS+PPSP S
Sbjct: 283 PPSPSSPPSPPAS 245
>TC8030
Length = 991
Score = 26.2 bits (56), Expect = 7.0
Identities = 18/53 (33%), Positives = 27/53 (49%), Gaps = 2/53 (3%)
Frame = +1
Query: 11 MAFPPSPSA-PPSPHLSGLRS-AASSALAEQDKYLTELLGERHILGPFMAVLP 61
+ + PSP A PP P S + AA+S Q++ T +G L PF ++ P
Sbjct: 100 LPWKPSPRAKPPPPRASSSHAHAATSMYPSQERATT--MGNHRRLSPFRSLWP 252
>AV427017
Length = 429
Score = 26.2 bits (56), Expect = 7.0
Identities = 16/40 (40%), Positives = 23/40 (57%), Gaps = 1/40 (2%)
Frame = +3
Query: 3 SSSGAGRYMAFPPSPSAPPSP-HLSGLRSAASSALAEQDK 41
++S A + PPS S+PPSP SG SA +S+ + K
Sbjct: 147 TASSASSARSSPPSSSSPPSPASSSG*SSAPTSSNSASPK 266
>AV766345
Length = 349
Score = 25.8 bits (55), Expect = 9.1
Identities = 13/24 (54%), Positives = 13/24 (54%)
Frame = -2
Query: 14 PPSPSAPPSPHLSGLRSAASSALA 37
PPSP APP R AA S LA
Sbjct: 93 PPSPQAPPPRQHLTTREAAHSPLA 22
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,825,785
Number of Sequences: 28460
Number of extensions: 66922
Number of successful extensions: 889
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 818
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 883
length of query: 294
length of database: 4,897,600
effective HSP length: 90
effective length of query: 204
effective length of database: 2,336,200
effective search space: 476584800
effective search space used: 476584800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0083.16


