

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.15
(489 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
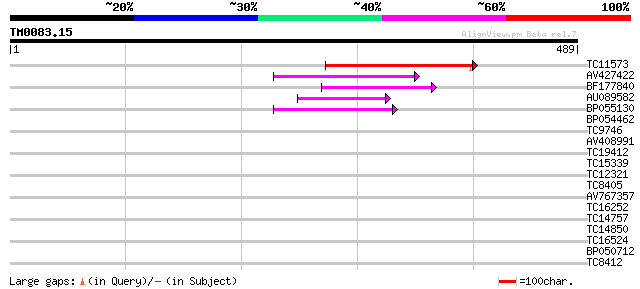
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 115 2e-26
AV427422 62 2e-10
BF177840 60 6e-10
AU089582 48 3e-06
BP055130 47 7e-06
BP054462 34 0.047
TC9746 32 0.18
AV408991 30 0.89
TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%) 28 2.6
TC15339 similar to UP|Q84VJ1 (Q84VJ1) Biostress-resistance-relat... 28 3.4
TC12321 similar to UP|Q15415 (Q15415) YRRM2, partial (5%) 28 3.4
TC8405 weakly similar to UP|O04083 (O04083) Lysophospholipase is... 28 4.4
AV767357 28 4.4
TC16252 27 5.8
TC14757 similar to UP|O04361 (O04361) Prohibitin, complete 27 7.5
TC14850 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein ... 27 7.5
TC16524 similar to GB|AAL69537.1|18491137|AY074839 AT5g45420/MFC... 27 9.8
BP050712 27 9.8
TC8412 weakly similar to UP|Q9LW00 (Q9LW00) Genomic DNA, chromos... 27 9.8
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 115 bits (288), Expect = 2e-26
Identities = 58/136 (42%), Positives = 87/136 (63%), Gaps = 5/136 (3%)
Frame = +2
Query: 273 DNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAE 332
+NG QFTSK +D DG+ I+ RF+SV+HPQTNGQ E+AN+VIL+G+++RL +A+G W +
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 333 QLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIREPSARTAGFDLNQNDQLLGEDLD 392
+L VLW+Y T P S+ E+P+ LT+G + ++P EI S T +N + +
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEISNFSW*TKARHEEENRMNMAVGWE 367
Query: 393 LLAE-----RRAIANC 403
L E R ++ NC
Sbjct: 368 KLPEGAI**RPSVRNC 415
>AV427422
Length = 417
Score = 62.0 bits (149), Expect = 2e-10
Identities = 37/127 (29%), Positives = 65/127 (51%), Gaps = 1/127 (0%)
Frame = +1
Query: 228 LVVAVDYYTKWIEAEPLATITSARV-QRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDL 286
++V VD +K+ PL +A+V F + V+R GVP +++D F S +++L
Sbjct: 37 VLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMSNFWKEL 216
Query: 287 LDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPH 346
K + ++ HP+++GQ E NR + LR + D WA + + Y T+ H
Sbjct: 217 FKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAEYWYNTSYH 396
Query: 347 STTGESP 353
+TG++P
Sbjct: 397 VSTGQTP 417
>BF177840
Length = 410
Score = 60.5 bits (145), Expect = 6e-10
Identities = 31/99 (31%), Positives = 51/99 (51%)
Frame = +2
Query: 270 LITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGD 329
+++D T+F S +R L + K +++ HPQT+GQ E N+ + LR L
Sbjct: 17 IVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLKM 196
Query: 330 WAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEI 368
W L H+ +AY HSTT SP+ + +G + P ++
Sbjct: 197 WETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>AU089582
Length = 383
Score = 48.1 bits (113), Expect = 3e-06
Identities = 23/80 (28%), Positives = 44/80 (54%)
Frame = +2
Query: 249 SARVQRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQA 308
+++ + + ++ GVP +I+D G QFTS +R L + + ++ HPQT+GQ+
Sbjct: 11 ASQYAKIYLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQS 190
Query: 309 ESANRVILRGLRKRLGDAKG 328
E +++ LR + D +G
Sbjct: 191 ERTIQILEDMLRACVXDLRG 250
>BP055130
Length = 567
Score = 47.0 bits (110), Expect = 7e-06
Identities = 29/108 (26%), Positives = 51/108 (46%), Gaps = 1/108 (0%)
Frame = +2
Query: 228 LVVAVDYYTKWIEAEP-LATITSARVQRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDL 286
++V VD +K+ P A TS++V F +++ G+P +++D FTS ++
Sbjct: 191 IIVVVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAFWKHF 370
Query: 287 LDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQL 334
+S HPQT+GQ E+ N+ + LR + + W L
Sbjct: 371 FKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLWVSYL 514
>BP054462
Length = 422
Score = 34.3 bits (77), Expect = 0.047
Identities = 37/142 (26%), Positives = 64/142 (45%), Gaps = 7/142 (4%)
Frame = +1
Query: 352 SPYRLTFGTEAVIPAEIREPSARTAGFDLNQNDQLLGEDLDLLAERRAIANCRELIAKQE 411
SP++ +G E + + ++ A ND +G D +LL++ RA L+ Q+
Sbjct: 31 SPFQALYGREPPVLLQGTTIPSKIAAV----NDLQVGRD-ELLSDLRA-----NLLKSQD 180
Query: 412 AAIRY-NKKVVLRSFAAGDLVL------QNASIGARSTERGKLAAKWEGPYRVVEATGTG 464
Y NKK + GD V + S+ + E KL+ ++ GPY +V G
Sbjct: 181 MMRTYANKKRRDVDYQIGDEVFLKLQPYRRRSLAKKMNE--KLSPRYYGPYPIVAKIGAV 354
Query: 465 AYKLETLAGKEIPRTWNATNLR 486
AY+LE A + ++ + L+
Sbjct: 355 AYRLELPAHSRVHPVFHVSLLK 420
>TC9746
Length = 637
Score = 32.3 bits (72), Expect = 0.18
Identities = 24/64 (37%), Positives = 26/64 (40%), Gaps = 1/64 (1%)
Frame = +3
Query: 9 HMLRARLRQLEEQETQSRP-CPRFPRKRRRTGARSHPLTTRKHAAGHPRPSGWPAIHRGA 67
H + R R L T P R P +RRRT SHP R H P P R A
Sbjct: 429 HEVHRRRRHLRNTPTHHPPDLRRRPLRRRRT---SHPRLHRSHVRPLPLPPSASRPRRSA 599
Query: 68 HPGA 71
PGA
Sbjct: 600 LPGA 611
>AV408991
Length = 435
Score = 30.0 bits (66), Expect = 0.89
Identities = 12/28 (42%), Positives = 19/28 (67%)
Frame = +1
Query: 71 AGAHLRLGIGLLAVQASTLSGWLPEEKA 98
+G H LG+ + V+++TL WLPE+ A
Sbjct: 259 SGIHRSLGVHISKVRSATLDTWLPEQVA 342
>TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%)
Length = 519
Score = 28.5 bits (62), Expect = 2.6
Identities = 11/29 (37%), Positives = 21/29 (71%)
Frame = +3
Query: 440 RSTERGKLAAKWEGPYRVVEATGTGAYKL 468
R ++GKL+ ++ GP+ V+E G+ +Y+L
Sbjct: 30 RFGKKGKLSPRFIGPFEVLERVGSVSYRL 116
>TC15339 similar to UP|Q84VJ1 (Q84VJ1) Biostress-resistance-related protein
(Fragment), partial (59%)
Length = 1111
Score = 28.1 bits (61), Expect = 3.4
Identities = 24/77 (31%), Positives = 31/77 (40%), Gaps = 11/77 (14%)
Frame = +3
Query: 89 LSGWLPEEKAEAKKL------VRRASWYTIV-----NDDLFKRGFSTPLLKCLAKDRAAY 137
LSGWLP K + KL RRA I+ DD+ F KCL+
Sbjct: 579 LSGWLPCAKTLSNKLQGLDEATRRAQSLPILMCHGKGDDVVPYKFGEKSSKCLSSTGFQD 758
Query: 138 VLAEIHEGSCGHHLGGR 154
V + + +C H GR
Sbjct: 759 VTFKSYYWACPLHSSGR 809
>TC12321 similar to UP|Q15415 (Q15415) YRRM2, partial (5%)
Length = 812
Score = 28.1 bits (61), Expect = 3.4
Identities = 18/51 (35%), Positives = 23/51 (44%), Gaps = 5/51 (9%)
Frame = +2
Query: 12 RARLRQLEEQETQSRPCPRFPRKRRRTGARSHPLTTRKH-----AAGHPRP 57
RA R+ +SRP P PR R R+ L++R H A G RP
Sbjct: 581 RALRRRQSRLREESRPKPLLPRPRSMAVRRTRTLSSRNHRRRSSAGGLRRP 733
>TC8405 weakly similar to UP|O04083 (O04083) Lysophospholipase isolog,
25331-24357 (At1g11090), partial (18%)
Length = 552
Score = 27.7 bits (60), Expect = 4.4
Identities = 29/83 (34%), Positives = 39/83 (46%), Gaps = 8/83 (9%)
Frame = +1
Query: 6 EEIHMLR-ARLRQLEEQETQSRPC---PRFPRKRRRTGAR-SHPLTTRKHAAGHPRPSGW 60
E++H LR LR+L + + RP PR PR+ R+ R + L R RPSG
Sbjct: 253 EDLHQLRHLGLRRLRRRPPRPRPLRRPPRLPRRPRQRRLRLALLLPPRPPQRVLLRPSGL 432
Query: 61 PAIHRGAHPGAGAH---LRLGIG 80
P R H A H L++G G
Sbjct: 433 PL--RRVHGRARHHPHVLQIGAG 495
>AV767357
Length = 458
Score = 27.7 bits (60), Expect = 4.4
Identities = 17/39 (43%), Positives = 19/39 (48%), Gaps = 7/39 (17%)
Frame = +1
Query: 41 RSHPLTTRKHAAGHP------RPSGWPAIHRG-AHPGAG 72
RS PL R+H A HP P+ P HRG HP G
Sbjct: 208 RSGPLGRRRHPAFHPACRPACHPACRPEYHRGHRHPKTG 324
>TC16252
Length = 550
Score = 27.3 bits (59), Expect = 5.8
Identities = 9/28 (32%), Positives = 13/28 (46%)
Frame = -3
Query: 43 HPLTTRKHAAGHPRPSGWPAIHRGAHPG 70
HP+ + + H P W +H HPG
Sbjct: 266 HPMLIQSRSECHLHPQAWNLLHSLVHPG 183
>TC14757 similar to UP|O04361 (O04361) Prohibitin, complete
Length = 1141
Score = 26.9 bits (58), Expect = 7.5
Identities = 19/52 (36%), Positives = 23/52 (43%)
Frame = +2
Query: 26 RPCPRFPRKRRRTGARSHPLTTRKHAAGHPRPSGWPAIHRGAHPGAGAHLRL 77
RP PR PR+ RR R +P P P G +H HP + HL L
Sbjct: 167 RPIPRHPRRHRR---RRNPF---------PHPMGPETLHL-RHPHSPPHLLL 283
>TC14850 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (25%)
Length = 577
Score = 26.9 bits (58), Expect = 7.5
Identities = 19/58 (32%), Positives = 26/58 (44%), Gaps = 2/58 (3%)
Frame = -1
Query: 157 ARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSL--VSPWPFHQWGM 212
A KV+ AG P++ A H HH PPA L++L SP P+ G+
Sbjct: 328 ATKVITAGRIVPSVHHQAYHHP------------HHMPPAELAALSSYSPLPY*HHGV 191
>TC16524 similar to GB|AAL69537.1|18491137|AY074839 AT5g45420/MFC19_9
{Arabidopsis thaliana;}, partial (41%)
Length = 997
Score = 26.6 bits (57), Expect = 9.8
Identities = 14/33 (42%), Positives = 17/33 (51%), Gaps = 2/33 (6%)
Frame = +1
Query: 33 RKRRRTGARSHPLTTRKHAAGHP--RPSGWPAI 63
R+RRR G R +K A HP +P W AI
Sbjct: 358 RRRRRGGMREDVEILKKQMAKHPVGKPGRWEAI 456
>BP050712
Length = 571
Score = 26.6 bits (57), Expect = 9.8
Identities = 19/54 (35%), Positives = 23/54 (42%)
Frame = +1
Query: 139 LAEIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHH 192
+AE HEG CG G S R+ R P + DA QR +LHH
Sbjct: 343 VAEEHEGRCGRVAGNLSSPRRRRRE----PLIAVDAK---------QRQRELHH 465
>TC8412 weakly similar to UP|Q9LW00 (Q9LW00) Genomic DNA, chromosome 3, P1
clone: MSJ11, partial (39%)
Length = 1403
Score = 26.6 bits (57), Expect = 9.8
Identities = 10/22 (45%), Positives = 14/22 (63%)
Frame = -3
Query: 191 HHAPPARLSSLVSPWPFHQWGM 212
HH PP+ L+S SPW +G+
Sbjct: 81 HHQPPSPLNSPFSPWSCFCFGL 16
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.137 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,615,792
Number of Sequences: 28460
Number of extensions: 122290
Number of successful extensions: 732
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 731
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 732
length of query: 489
length of database: 4,897,600
effective HSP length: 94
effective length of query: 395
effective length of database: 2,222,360
effective search space: 877832200
effective search space used: 877832200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0083.15


