

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0079c.14
(171 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
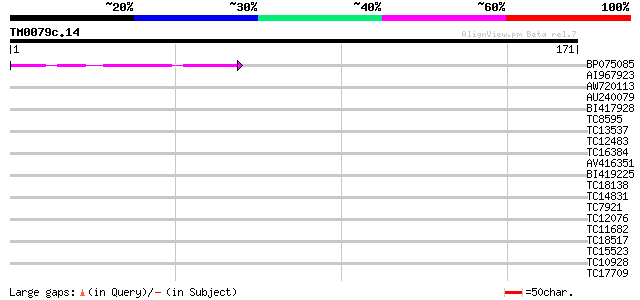
Score E
Sequences producing significant alignments: (bits) Value
BP075085 48 1e-06
AI967923 32 0.076
AW720113 30 0.22
AU240079 30 0.29
BI417928 29 0.38
TC8595 similar to PIR|T09390|T09390 21K protein precursor - alfa... 29 0.38
TC13537 similar to GB|AAP21222.1|30102608|BT006414 At3g06035 {Ar... 29 0.49
TC12483 weakly similar to UP|Q40552 (Q40552) Pistil extensin lik... 29 0.49
TC16384 homologue to UP|Q8L5G3 (Q8L5G3) Proline rich protein 3 (... 29 0.49
AV416351 28 0.64
BI419225 28 0.64
TC18138 UP|ACT1_TOBAC (Q05214) Actin, partial (39%) 28 0.64
TC14831 homologue to UP|P93499 (P93499) DnaJ-like protein (Fragm... 28 0.84
TC7921 homologue to UP|Q8LHP9 (Q8LHP9) OJ1008_F01.11 protein, pa... 28 0.84
TC12076 similar to GB|AAL69531.1|18491125|AY074833 At2g47490/T30... 28 1.1
TC11682 homologue to UP|Q7PZ83 (Q7PZ83) AgCP9540 (Fragment), par... 28 1.1
TC18517 weakly similar to UP|AAQ82688 (AAQ82688) Epa4p, partial ... 27 1.4
TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-bindi... 27 1.4
TC10928 weakly similar to PIR|G96668|G96668 protein F1N19.7 [imp... 27 1.4
TC17709 weakly similar to UP|Q9MA09 (Q9MA09) F20B17.9 (At1g79660... 27 1.9
>BP075085
Length = 407
Score = 47.8 bits (112), Expect = 1e-06
Identities = 33/70 (47%), Positives = 42/70 (59%)
Frame = -2
Query: 1 MRGAESEGGGGGGGCAVRGGNIYWGRKHATDFRGIVVIFAWVSVPQTLLQDFVNLYSSLG 60
MRGAESEG GG CAVR GNIY + + ++ F + LL V+LYSSLG
Sbjct: 370 MRGAESEGDGG---CAVRXGNIY-----SEESL*SLLGFLFHRHCHRLL---VDLYSSLG 224
Query: 61 WNSLVCYAHY 70
WNSLV ++ +
Sbjct: 223 WNSLVSFSFF 194
>AI967923
Length = 302
Score = 31.6 bits (70), Expect = 0.076
Identities = 18/38 (47%), Positives = 22/38 (57%), Gaps = 7/38 (18%)
Frame = -1
Query: 4 AESEG---GGGGGGCAVRGGN----IYWGRKHATDFRG 34
AE+EG GGGG G A RGG+ + GR+H RG
Sbjct: 176 AEAEGVHGGGGGEGVAARGGSGSREVVGGRRHVRQARG 63
>AW720113
Length = 512
Score = 30.0 bits (66), Expect = 0.22
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = -1
Query: 8 GGGGGGGCAVRGGNIYWGRKHATDFRGIVVIFAWV 42
GGGGG GC GG+ G + +VV +W+
Sbjct: 167 GGGGGCGCGCGGGSGGGGGSSGEGWNAVVVGVSWI 63
>AU240079
Length = 300
Score = 29.6 bits (65), Expect = 0.29
Identities = 16/54 (29%), Positives = 25/54 (45%)
Frame = -1
Query: 105 AFSAGSKACLYKVLQLIDGRCETPHCLHNYHLLRNCVSGHIYDSGPLDVTSDFG 158
+F + A Y ++ + C C H+Y RN SGH+ S P+ +T G
Sbjct: 186 SFLISTSAFHYGIVCCLVNAC----CFHSYXKKRNKTSGHLKRSSPIVITCPSG 37
>BI417928
Length = 509
Score = 29.3 bits (64), Expect = 0.38
Identities = 12/18 (66%), Positives = 13/18 (71%)
Frame = -3
Query: 3 GAESEGGGGGGGCAVRGG 20
G E GGGGGGG +V GG
Sbjct: 237 GVEERGGGGGGGESVGGG 184
>TC8595 similar to PIR|T09390|T09390 21K protein precursor - alfalfa
{Medicago sativa;}, partial (93%)
Length = 811
Score = 29.3 bits (64), Expect = 0.38
Identities = 12/26 (46%), Positives = 14/26 (53%)
Frame = -2
Query: 1 MRGAESEGGGGGGGCAVRGGNIYWGR 26
++G E E GG GC GGN W R
Sbjct: 87 IQGGEEEQGGEEKGCVCHGGNEDWAR 10
>TC13537 similar to GB|AAP21222.1|30102608|BT006414 At3g06035 {Arabidopsis
thaliana;} , partial (19%)
Length = 765
Score = 28.9 bits (63), Expect = 0.49
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = +2
Query: 80 VPLAFCVVDELIEELKTKSCPVVFAAFSAGS 110
VP A+C+ DE+ EE++ C V + GS
Sbjct: 146 VPKAYCLADEIAEEMEDIPCEQVSQYYPGGS 238
>TC12483 weakly similar to UP|Q40552 (Q40552) Pistil extensin like protein
(Fragment), partial (6%)
Length = 534
Score = 28.9 bits (63), Expect = 0.49
Identities = 10/15 (66%), Positives = 11/15 (72%)
Frame = -2
Query: 10 GGGGGCAVRGGNIYW 24
GGGGGC V GG+ W
Sbjct: 302 GGGGGCLVAGGDCMW 258
>TC16384 homologue to UP|Q8L5G3 (Q8L5G3) Proline rich protein 3 (Fragment),
partial (43%)
Length = 610
Score = 28.9 bits (63), Expect = 0.49
Identities = 24/75 (32%), Positives = 31/75 (41%), Gaps = 1/75 (1%)
Frame = -3
Query: 8 GGGGGGGCAVRGGNIYWGRKHATDFRGIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLVCY 67
GGG GGG A GG+ +W + W+ +P D NL SL
Sbjct: 212 GGGVGGGTAAAGGHCWW--------------YGWMPLP---AMDEFNLSVSL------TQ 102
Query: 68 AHYLSAFR-DESTVP 81
+H FR DE+T P
Sbjct: 101 SHSFPWFRTDETTKP 57
>AV416351
Length = 428
Score = 28.5 bits (62), Expect = 0.64
Identities = 11/16 (68%), Positives = 13/16 (80%)
Frame = -2
Query: 5 ESEGGGGGGGCAVRGG 20
E +GGGGGGG +RGG
Sbjct: 130 ELDGGGGGGGRGLRGG 83
>BI419225
Length = 205
Score = 28.5 bits (62), Expect = 0.64
Identities = 11/17 (64%), Positives = 12/17 (69%)
Frame = +3
Query: 8 GGGGGGGCAVRGGNIYW 24
GGGGGGG V GG+ W
Sbjct: 141 GGGGGGGAXVGGGSGAW 191
>TC18138 UP|ACT1_TOBAC (Q05214) Actin, partial (39%)
Length = 580
Score = 28.5 bits (62), Expect = 0.64
Identities = 11/17 (64%), Positives = 13/17 (75%)
Frame = -1
Query: 4 AESEGGGGGGGCAVRGG 20
AE EGGGGGGG ++ G
Sbjct: 130 AEEEGGGGGGGGGIKEG 80
>TC14831 homologue to UP|P93499 (P93499) DnaJ-like protein (Fragment),
partial (63%)
Length = 851
Score = 28.1 bits (61), Expect = 0.84
Identities = 11/19 (57%), Positives = 13/19 (67%)
Frame = -1
Query: 3 GAESEGGGGGGGCAVRGGN 21
G S GGGGGGGC+ G+
Sbjct: 191 GGSSIGGGGGGGCSCGEGD 135
>TC7921 homologue to UP|Q8LHP9 (Q8LHP9) OJ1008_F01.11 protein, partial (7%)
Length = 1202
Score = 28.1 bits (61), Expect = 0.84
Identities = 10/15 (66%), Positives = 12/15 (79%)
Frame = -2
Query: 5 ESEGGGGGGGCAVRG 19
E++GGGGGGGC G
Sbjct: 466 ETDGGGGGGGCRKSG 422
>TC12076 similar to GB|AAL69531.1|18491125|AY074833 At2g47490/T30B22.21
{Arabidopsis thaliana;}, partial (20%)
Length = 506
Score = 27.7 bits (60), Expect = 1.1
Identities = 14/35 (40%), Positives = 19/35 (54%), Gaps = 7/35 (20%)
Frame = -2
Query: 6 SEGGGGGGGCA-------VRGGNIYWGRKHATDFR 33
+ GGG GGG A +RGG + GR+H +R
Sbjct: 322 NSGGGTGGGIAENTFGVDIRGGGMGIGRRHGGSWR 218
>TC11682 homologue to UP|Q7PZ83 (Q7PZ83) AgCP9540 (Fragment), partial (11%)
Length = 822
Score = 27.7 bits (60), Expect = 1.1
Identities = 11/14 (78%), Positives = 11/14 (78%)
Frame = -1
Query: 8 GGGGGGGCAVRGGN 21
GGGGGGG RGGN
Sbjct: 402 GGGGGGGGGSRGGN 361
>TC18517 weakly similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (4%)
Length = 493
Score = 27.3 bits (59), Expect = 1.4
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = -1
Query: 10 GGGGGCAVRGGNIYWGRK 27
G GGGCA RGG + G K
Sbjct: 262 GSGGGCASRGGRLRRGSK 209
>TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-binding,
abscisic acid-inducible protein, complete
Length = 972
Score = 27.3 bits (59), Expect = 1.4
Identities = 12/25 (48%), Positives = 13/25 (52%)
Frame = +2
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRK 27
G GGGGGG GG Y GR+
Sbjct: 665 GGSRYSGGGGGGYGGSGGGGYGGRR 739
>TC10928 weakly similar to PIR|G96668|G96668 protein F1N19.7 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 590
Score = 27.3 bits (59), Expect = 1.4
Identities = 11/17 (64%), Positives = 11/17 (64%)
Frame = +2
Query: 3 GAESEGGGGGGGCAVRG 19
GA GGGGGGG A G
Sbjct: 239 GAAGGGGGGGGGAAAEG 289
>TC17709 weakly similar to UP|Q9MA09 (Q9MA09) F20B17.9
(At1g79660/F20B17_9), partial (39%)
Length = 576
Score = 26.9 bits (58), Expect = 1.9
Identities = 12/22 (54%), Positives = 13/22 (58%)
Frame = -1
Query: 3 GAESEGGGGGGGCAVRGGNIYW 24
G E+ GGGGGGG GG W
Sbjct: 90 GREANGGGGGGG----GGGFGW 37
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.141 0.461
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,655,030
Number of Sequences: 28460
Number of extensions: 57620
Number of successful extensions: 1498
Number of sequences better than 10.0: 130
Number of HSP's better than 10.0 without gapping: 1023
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1300
length of query: 171
length of database: 4,897,600
effective HSP length: 84
effective length of query: 87
effective length of database: 2,506,960
effective search space: 218105520
effective search space used: 218105520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0079c.14


