

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0079a.1
(139 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
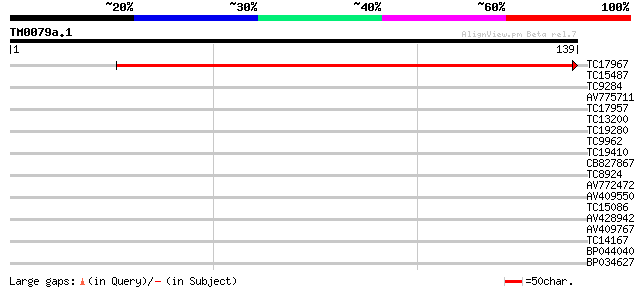
Score E
Sequences producing significant alignments: (bits) Value
TC17967 similar to UP|Q7XA66 (Q7XA66) At1g49590, partial (17%) 225 2e-60
TC15487 similar to UP|O23144 (O23144) Proton pump interactor (AT... 33 0.023
TC9284 weakly similar to UP|Q9LPY0 (Q9LPY0) T23J18.22, partial (... 29 0.34
AV775711 28 0.44
TC17957 weakly similar to GB|AAM98310.1|22655436|AY142046 At2g19... 28 0.44
TC13200 similar to UP|RIR2_TOBAC (P49730) Ribonucleoside-diphosp... 28 0.58
TC19280 homologue to PIR|JQ2256|JQ2256 phosphoribosylformylglyci... 27 1.3
TC9962 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial... 27 1.3
TC19410 weakly similar to AAR24668 (AAR24668) At1g24040, partial... 27 1.3
CB827867 27 1.7
TC8924 similar to UP|Q9SXS8 (Q9SXS8) Ethylene responsive element... 26 2.2
AV772472 26 2.2
AV409550 26 2.9
TC15086 similar to UP|Q8VZF6 (Q8VZF6) AT5g45560/MFC19_23, partia... 26 2.9
AV428942 26 2.9
AV409767 25 3.7
TC14167 similar to UP|O48642 (O48642) Class III acidic endochiti... 25 4.9
BP044040 25 4.9
BP034627 25 4.9
>TC17967 similar to UP|Q7XA66 (Q7XA66) At1g49590, partial (17%)
Length = 662
Score = 225 bits (573), Expect = 2e-60
Identities = 113/113 (100%), Positives = 113/113 (100%)
Frame = +2
Query: 27 YSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVK 86
YSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVK
Sbjct: 2 YSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVK 181
Query: 87 AAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQEREKSLLGLYSKP 139
AAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQEREKSLLGLYSKP
Sbjct: 182 AAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQEREKSLLGLYSKP 340
>TC15487 similar to UP|O23144 (O23144) Proton pump interactor
(AT4g27500/F27G19_100), partial (16%)
Length = 1153
Score = 32.7 bits (73), Expect = 0.023
Identities = 33/118 (27%), Positives = 53/118 (43%), Gaps = 14/118 (11%)
Frame = +2
Query: 27 YSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGR-----VVTTSLNP 81
+S AI K +EE SP T+K S + KP L V P
Sbjct: 233 FSKAIPKQ-PKEEPKPSPQETLPTQKESKNKVRDLKAKPDKKDLEEADEYEFEVPRKETP 409
Query: 82 TRNVKAAPSSLS-------LGKRKRPGEKPKVVSEEE--KSALKAREAARKRVQEREK 130
+ + P+ L + K K+ E+ K ++E+ K+ALKA++ A K++++REK
Sbjct: 410 PKEPEIDPAKLKEMKREEEIAKAKQAWERKKKLAEKAAAKAALKAQKEAEKKLKDREK 583
>TC9284 weakly similar to UP|Q9LPY0 (Q9LPY0) T23J18.22, partial (33%)
Length = 898
Score = 28.9 bits (63), Expect = 0.34
Identities = 19/54 (35%), Positives = 27/54 (49%)
Frame = +1
Query: 47 ASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRP 100
ASTTK +++TS +KPH P P +T L+ T N +A S + P
Sbjct: 214 ASTTKPSAATSPP---SKPH-APTPSSSSSTPLSSTTNPNSASPSFATSSPSSP 363
>AV775711
Length = 255
Score = 28.5 bits (62), Expect = 0.44
Identities = 16/49 (32%), Positives = 24/49 (48%)
Frame = +3
Query: 15 NGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGN 63
N FY P F+Y+ KW + STT+K++++S NK N
Sbjct: 27 NSFYLKPTKTFFYTMEYNKWHWRRN--------STTQKSTNSSCLNKIN 149
>TC17957 weakly similar to GB|AAM98310.1|22655436|AY142046
At2g19080/T20K24.9 {Arabidopsis thaliana;}, partial
(10%)
Length = 648
Score = 28.5 bits (62), Expect = 0.44
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = +3
Query: 42 ASPHFASTTKKASSTSQSNKGNKPHNGP 69
+ PHF + ++S SQS K +KP + P
Sbjct: 87 SGPHFHTNASSSASRSQSTKSSKPKSKP 170
>TC13200 similar to UP|RIR2_TOBAC (P49730) Ribonucleoside-diphosphate
reductase small chain (Ribonucleotide reductase) (R2
subunit) , partial (40%)
Length = 624
Score = 28.1 bits (61), Expect = 0.58
Identities = 14/58 (24%), Positives = 26/58 (44%)
Frame = -2
Query: 42 ASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKR 99
+ PH + TT A+ +Q+ + PH+ P R + +P N + + +R R
Sbjct: 242 SDPHSSPTTHTAARRTQTPSQSPPHHAQSPSRAPR*ASSPEENTTSTRNRSEAPRRVR 69
>TC19280 homologue to PIR|JQ2256|JQ2256 phosphoribosylformylglycinamidine
cyclo-ligase precursor - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (18%)
Length = 572
Score = 26.9 bits (58), Expect = 1.3
Identities = 12/33 (36%), Positives = 18/33 (54%)
Frame = -1
Query: 42 ASPHFASTTKKASSTSQSNKGNKPHNGPLPGRV 74
AS F T ++ +S +GNKP P+PG +
Sbjct: 533 ASITFVPTPSVPATRYESPRGNKPPKPPIPGAI 435
>TC9962 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial (21%)
Length = 585
Score = 26.9 bits (58), Expect = 1.3
Identities = 16/58 (27%), Positives = 26/58 (44%)
Frame = +3
Query: 34 WVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSS 91
W+ + YAS +F +K+ +T + TT+ NP++N APSS
Sbjct: 66 WMAKNNKYASINFNHIYEKSITTKNT----------------TTTTNPSKNPSLAPSS 191
>TC19410 weakly similar to AAR24668 (AAR24668) At1g24040, partial (28%)
Length = 606
Score = 26.9 bits (58), Expect = 1.3
Identities = 20/76 (26%), Positives = 31/76 (40%), Gaps = 2/76 (2%)
Frame = +3
Query: 47 ASTTKKASSTSQSNKGNKPHNGPLPG--RVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKP 104
A+TT T PH PLP + S + + +PSS +L P
Sbjct: 117 ATTTANLYKTRPFTLSTTPHTNPLPSFHPHPSPSFRISSSQSQSPSSATLDDPSHPFLPG 296
Query: 105 KVVSEEEKSALKAREA 120
+ ++ EE L++ EA
Sbjct: 297 RFLTNEELQQLESLEA 344
>CB827867
Length = 471
Score = 26.6 bits (57), Expect = 1.7
Identities = 24/95 (25%), Positives = 40/95 (41%)
Frame = +3
Query: 35 VTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSL 94
V EE H S+ +ASS + GN+ G LP + PT+ P +
Sbjct: 60 VELEEQDVLKHPTSSAPEASSPAS---GNRALGGHLPSTATREVMTPTQ-----PETRI* 215
Query: 95 GKRKRPGEKPKVVSEEEKSALKAREAARKRVQERE 129
GKR+ G++ + E + K R++ E++
Sbjct: 216 GKRRADGQQRQ---GEARERPKIRDSLTTEASEQD 311
>TC8924 similar to UP|Q9SXS8 (Q9SXS8) Ethylene responsive element binding
factor, partial (28%)
Length = 778
Score = 26.2 bits (56), Expect = 2.2
Identities = 27/95 (28%), Positives = 37/95 (38%), Gaps = 2/95 (2%)
Frame = +2
Query: 18 YYDPKSG--FYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVV 75
Y+ P + FYY+ V Q+ + S T+ SST +S G +P
Sbjct: 23 YHHPAAADPFYYAAGNNGGVVQDHHHVSNPQRPTSSGMSSTVESFSGPRP---------- 172
Query: 76 TTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEE 110
PT AAP S S +R P VV E+
Sbjct: 173 -----PTVKPAAAPPSSS----RRYPRTPPVVPED 250
>AV772472
Length = 423
Score = 26.2 bits (56), Expect = 2.2
Identities = 8/22 (36%), Positives = 11/22 (49%)
Frame = +2
Query: 1 WDFESSSGYYYHKTNGFYYDPK 22
W GY++HK GF+ K
Sbjct: 47 WHLHEMDGYFHHKLTGFFLHTK 112
>AV409550
Length = 413
Score = 25.8 bits (55), Expect = 2.9
Identities = 11/38 (28%), Positives = 16/38 (41%)
Frame = +3
Query: 34 WVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLP 71
W+T E A P + T ++ S S + P P P
Sbjct: 282 WLTPSETSARPSSSPTASGSALASSSAPSSSPEGNPTP 395
>TC15086 similar to UP|Q8VZF6 (Q8VZF6) AT5g45560/MFC19_23, partial (38%)
Length = 1542
Score = 25.8 bits (55), Expect = 2.9
Identities = 23/110 (20%), Positives = 49/110 (43%), Gaps = 5/110 (4%)
Frame = +2
Query: 33 KWVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTR--NVKAAPS 90
+W Q + ++P SST+ S+KG KP++ S++PT + A
Sbjct: 26 EWFAQADERSAPPRIPVMVNMSSTAVSSKGQKPND---------LSMHPTSLDQINFASR 178
Query: 91 SLSLGKRKRPGEKPKVVSEEEKSALKA---REAARKRVQEREKSLLGLYS 137
+ +L ++ ++E E+ A ++ + R ++E +++ L S
Sbjct: 179 NSALMDEYSDEDEDFQIAEPEQEAFQSDLDNDVRRIALEEEPATVIDLSS 328
>AV428942
Length = 362
Score = 25.8 bits (55), Expect = 2.9
Identities = 20/64 (31%), Positives = 28/64 (43%)
Frame = +2
Query: 65 PHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKR 124
P P+ TSL R+ +AP L L +R+R ++ K VSE + R A
Sbjct: 41 PCRRPISSITTETSLLLRRHFFSAPKPLFLHRRRRFQKEEKQVSESR*RGGRGRG*AALE 220
Query: 125 VQER 128
V R
Sbjct: 221 VTRR 232
>AV409767
Length = 411
Score = 25.4 bits (54), Expect = 3.7
Identities = 11/25 (44%), Positives = 14/25 (56%)
Frame = -2
Query: 79 LNPTRNVKAAPSSLSLGKRKRPGEK 103
LNPT+ A + KR+RPG K
Sbjct: 407 LNPTQKAIAINKCIVCNKRRRPGNK 333
>TC14167 similar to UP|O48642 (O48642) Class III acidic endochitinase
precursor , partial (88%)
Length = 1336
Score = 25.0 bits (53), Expect = 4.9
Identities = 16/56 (28%), Positives = 26/56 (45%), Gaps = 2/56 (3%)
Frame = +2
Query: 43 SPHFASTTKKASS--TSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGK 96
+PH T + S +S S+ +PH+G L VVT P N+ + ++ K
Sbjct: 173 APHVTPETMRLCSLLSSTSSAAAEPHHGTLQAIVVTGVPVPNSNLTSKTANEKASK 340
>BP044040
Length = 491
Score = 25.0 bits (53), Expect = 4.9
Identities = 14/39 (35%), Positives = 21/39 (52%)
Frame = +3
Query: 53 ASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSS 91
+SS+S S+ P+ P P +T + NP N K P+S
Sbjct: 159 SSSSSSSSAPINPNRNPTPR--LTPTQNPNPNSKPRPAS 269
>BP034627
Length = 357
Score = 25.0 bits (53), Expect = 4.9
Identities = 12/35 (34%), Positives = 19/35 (54%)
Frame = +1
Query: 56 TSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPS 90
TS+ N+GNK +N T + P+RN + P+
Sbjct: 91 TSERNRGNKANNQKKEAHN*TKWMLPSRNYEHTPT 195
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.306 0.125 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,342,758
Number of Sequences: 28460
Number of extensions: 29979
Number of successful extensions: 197
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 195
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 197
length of query: 139
length of database: 4,897,600
effective HSP length: 81
effective length of query: 58
effective length of database: 2,592,340
effective search space: 150355720
effective search space used: 150355720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 50 (23.9 bits)
Lotus: description of TM0079a.1


