

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
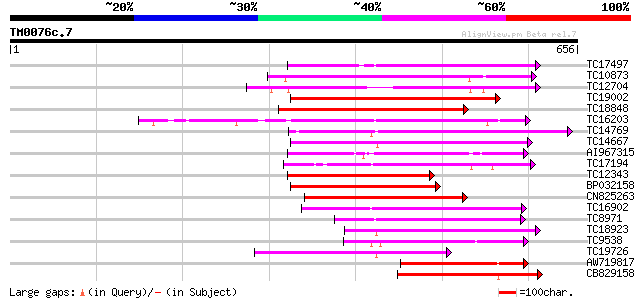
Query= TM0076c.7
(656 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2,... 227 4e-60
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 216 7e-57
TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3,... 211 4e-55
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 205 2e-53
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 205 2e-53
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 196 1e-50
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 196 1e-50
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 191 3e-49
AI967315 179 1e-45
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 178 3e-45
TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine prote... 177 5e-45
BP032158 177 6e-45
CN825263 175 2e-44
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 166 8e-42
TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P... 159 1e-39
TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1,... 157 7e-39
TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein k... 155 3e-38
TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase... 154 3e-38
AW719817 154 6e-38
CB829158 150 8e-37
>TC17497 UP|CAE45595 (CAE45595) S-receptor kinase-like protein 2, complete
Length = 2946
Score = 227 bits (579), Expect = 4e-60
Identities = 128/296 (43%), Positives = 177/296 (59%), Gaps = 3/296 (1%)
Frame = +1
Query: 322 YEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIK 381
+ + ATN+F SNKLGEGG G VYKG L +G +A+KRLS + Q + F NE+ LI
Sbjct: 1642 FSTISSATNHFSLSNKLGEGGFGPVYKGLLANGQEIAVKRLSNTSGQGMEEFKNEIKLIA 1821
Query: 382 DLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEG 441
LQH+NLVKL GCS+ E+ N + L R+ + + W R +II G A G
Sbjct: 1822 RLQHRNLVKLFGCSVHQDENSHA-----NKKMKILLDSTRS-KLVDWNKRLQIIDGIARG 1983
Query: 442 LAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIST-AICGTLGYM 500
L YLH++S+L+IIHRD+K SNILLDD PKI+DFGLAR+F DQ + T + GT GYM
Sbjct: 1984 LLYLHQDSRLRIIHRDLKTSNILLDDEMNPKISDFGLARIFIGDQVEARTKRVMGTYGYM 2163
Query: 501 APEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFV--QNSCSILHMVWSLYGSNRLCDI 558
PEY V G + K+D++SFGV+V+E++SGK F + ++L W L+ R ++
Sbjct: 2164 PPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIGRFYDPHHHLNLLSHAWRLWIEERPLEL 2343
Query: 559 VDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIADPTQPPF 614
VD +L+ E + + + LLC Q E RP M +V M+N ++ P P F
Sbjct: 2344 VDELLDDPVIPTEILRYIHVALLCVQRRPENRPDMLSIVLMLNGEKELPKPRLPAF 2511
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 216 bits (551), Expect = 7e-57
Identities = 128/324 (39%), Positives = 185/324 (56%), Gaps = 13/324 (4%)
Frame = +1
Query: 299 QRRERRQFGAFLFSENNSK------LNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALP 352
++RE + G + S N S + + L AT F ++N +GEGG G VYKG L
Sbjct: 334 RKRETKGKGKGVSSSNGSSNGKTAAASFGFRELADATRNFKEANLIGEGGFGKVYKGRLT 513
Query: 353 DGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLS 412
G VA+K+LS + Q F EV ++ L H NLV+L+G G + LLVYEY+P S
Sbjct: 514 TGEAVAVKQLSHDGRQGFQEFVMEVLMLSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGS 693
Query: 413 LHDHL-SVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTP 471
L DHL + + + L W R K+ +G A GL YLH + +I+RD+K +NILLD+ F P
Sbjct: 694 LEDHLFELSHDKEPLNWSTRMKVAVGAARGLEYLHCTADPPVIYRDLKSANILLDNEFNP 873
Query: 472 KIADFGLARLFP-EDQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGK 530
K++DFGLA+L P D + +ST + GT GY APEY + GKLT K+DIYSFGV+++ELL+G+
Sbjct: 874 KLSDFGLAKLGPVGDNTHVSTRVMGTYGYCAPEYAMSGKLTLKSDIYSFGVVLLELLTGR 1053
Query: 531 -----SRTSFVQNSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQA 585
SR QN S +S R +VDP+L+G +P+ + + I +C Q
Sbjct: 1054RAIDTSRRPGEQNLVSWARPYFS--DRRRFGHMVDPLLQGRFPSRCLHQAIAITAMCLQE 1227
Query: 586 SAELRPPMSVVVKMINNIHKIADP 609
+ RP ++ +V + + P
Sbjct: 1228QPKFRPLITDIVVALEYLASQGSP 1299
>TC12704 UP|CAE45596 (CAE45596) S-receptor kinase-like protein 3, complete
Length = 2481
Score = 211 bits (536), Expect = 4e-55
Identities = 136/374 (36%), Positives = 191/374 (50%), Gaps = 34/374 (9%)
Frame = +1
Query: 275 SFAAVATLLIVATIIFFVRKNVLRQRR---ERRQFGAFLFSENNSKLNMP----YEVLEK 327
+F T+L +AT+ RK R+ E R + + +++ + +
Sbjct: 1261 AFIICITILGLATVTCIRRKKNEREDEGGIETRIINHWKDKRGDEDIDLATIFDFSTISS 1440
Query: 328 ATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKN 387
TN+F +SNKLGEGG G VYKG L +G +A+KRLS + Q + F NEV LI LQH+N
Sbjct: 1441 TTNHFSESNKLGEGGFGPVYKGVLANGQEIAVKRLSNTSGQGMEEFKNEVKLIARLQHRN 1620
Query: 388 LVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHE 447
LVKLLGCSI E LL+YE++ N SL Y
Sbjct: 1621 LVKLLGCSIHHDEMLLIYEFMHNRSLD-----------------------------YFIF 1713
Query: 448 ESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIST-AICGTLGYMAPEYIV 506
+S+L+IIHRD+K SNILLD PKI+DFGLAR+F DQ + T + GT GYM+PEY V
Sbjct: 1714 DSRLRIIHRDLKTSNILLDSEMNPKISDFGLARIFTGDQVEAKTKRVMGTYGYMSPEYAV 1893
Query: 507 LGKLTEKADIYSFGVLVIELLSGKS----------RTSFVQNSCSILHMV---------- 546
G + K+D++SFGV+V+E++SGK R +S + ++
Sbjct: 1894 HGSFSVKSDVFSFGVIVLEIISGKKIGRFCDPHHHRNLLSHSSNFAVFLIKALRICMFEN 2073
Query: 547 ------WSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMI 600
W L+ R ++VD +L+G E + + I LLC Q E RP M VV M+
Sbjct: 2074 VKNRKAWRLWIEERPLELVDELLDGLAIPTEILRYIHIALLCVQQRPEYRPDMLSVVLML 2253
Query: 601 NNIHKIADPTQPPF 614
N ++ P+ P F
Sbjct: 2254 NGEKELPKPSLPAF 2295
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 205 bits (522), Expect = 2e-53
Identities = 109/246 (44%), Positives = 154/246 (62%), Gaps = 3/246 (1%)
Frame = +1
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQ 384
L ATN F+ NKLGEGG GSVY G L DG+ +A+KRL + + F EV ++ ++
Sbjct: 43 LHSATNNFNYDNKLGEGGFGSVYWGQLWDGSQIAVKRLKVWSNKADMEFAVEVEILARVR 222
Query: 385 HKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQ-QLTWEVRHKIILGTAEGLA 443
HKNL+ L G G E L+VY+Y+PNLSL HL + + + L W R I +G+AEG+
Sbjct: 223 HKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIAIGSAEGIV 402
Query: 444 YLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAPE 503
YLH ++ IIHRDIK SN+LLD +F ++ADFG A+L P+ + ++T + GTLGY+APE
Sbjct: 403 YLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGATHVTTRVKGTLGYLAPE 582
Query: 504 YIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSC--SILHMVWSLYGSNRLCDIVDP 561
Y +LGK E D++SFG+L++EL SGK + ++ SI L + + + DP
Sbjct: 583 YAMLGKANECCDVFSFGILLLELASGKKPLEKLSSTVKRSINDWALPLACAKKFTEFADP 762
Query: 562 ILEGNY 567
L G Y
Sbjct: 763 RLNGEY 780
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 205 bits (521), Expect = 2e-53
Identities = 105/220 (47%), Positives = 147/220 (66%), Gaps = 1/220 (0%)
Frame = +2
Query: 312 SENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWAD 371
S NNS Y+ L AT F D NKLGEGG GSVY G DG +A+K+L ++
Sbjct: 242 SVNNSWRIFTYKELHAATGGFSDDNKLGEGGFGSVYWGRTSDGLQIAVKKLKAMNSKAEM 421
Query: 372 HFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQ-QLTWEV 430
F EV ++ ++HKNL+ L G + + L+VY+Y+PNLSL HL + ++ QL W+
Sbjct: 422 EFAVEVEVLGRVRHKNLLGLRGYCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQK 601
Query: 431 RHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIS 490
R KI +G+AEG+ YLH E IIHRDIK SN+LL+ +F P +ADFG A+L PE S ++
Sbjct: 602 RMKIAIGSAEGILYLHHEVTPHIIHRDIKASNVLLNSDFEPLVADFGFAKLIPEGVSHMT 781
Query: 491 TAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGK 530
T + GTLGY+APEY + GK++E D+YSFG+L++EL++G+
Sbjct: 782 TRVKGTLGYLAPEYAMWGKVSESCDVYSFGILLLELVTGR 901
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 196 bits (498), Expect = 1e-50
Identities = 151/489 (30%), Positives = 246/489 (49%), Gaps = 36/489 (7%)
Frame = +2
Query: 150 DLTLCGNIDFSGNRSV--------HKANV--VELVRN-LSVEAPKNDGFFVGVLSKRNVS 198
++ + ++ SGN H+A++ V+L RN L+ E PK G+ + ++S
Sbjct: 1622 EIPMLTKVNISGNNLTGPIPTTITHRASLTAVDLSRNNLAGEVPK------GMKNLMDLS 1783
Query: 199 VYGLAQCWNFVNESVCQNC-LVEAVTRIDSCALKEEGRVLNAGCYLRFSTDKFYDNSSNI 257
+ L++ N ++ V + ++T +D + G V G +L F+ DK + + N+
Sbjct: 1784 ILNLSR--NEISGPVPDEIRFMTSLTTLDLSSNNFTGTVPTGGQFLVFNYDKTFAGNPNL 1957
Query: 258 APPG--------------TKGST-KVGIIVAESFAAVATLLIVATIIFFVRKNVLRQRRE 302
P T+ T +V IV A A LL+ T+ +V+R+RR
Sbjct: 1958 CFPHRASCPSVLYDSLRKTRAKTARVRAIVIGIALATAVLLVAVTV------HVVRKRRL 2119
Query: 303 RRQFGAFLFSENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRL 362
R L + ++ E + N +G+GG+G VY+G++P+GT VAIKRL
Sbjct: 2120 HRAQAWKLTAFQRLEIKA-----EDVVECLKEENIIGKGGAGIVYRGSMPNGTDVAIKRL 2284
Query: 363 SFNTTQWADHFFN-EVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRR 421
+ D+ F E+ + ++H+N+++LLG +LL+YEY+PN SL + L +
Sbjct: 2285 VGQGSGRNDYGFRAEIETLGKIRHRNIMRLLGYVSNKDTNLLLYEYMPNGSLGEWLHGAK 2464
Query: 422 NLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLAR- 480
L WE+R+KI + A GL Y+H + IIHRD+K +NILLD +F +ADFGLA+
Sbjct: 2465 G-GHLRWEMRYKIAVEAARGLCYMHHDCSPLIIHRDVKSNNILLDADFEAHVADFGLAKF 2641
Query: 481 LFPEDQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSC 540
L+ SQ ++I G+ GY+APEY K+ EK+D+YSFGV+++EL+ G+ +
Sbjct: 2642 LYDPGASQSMSSIAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELIIGRKPVGEFGDGV 2821
Query: 541 SILHMVWSLYG-------SNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPM 593
I+ V + + +VDP L G YP + I ++C + RP M
Sbjct: 2822 DIVGWVNKTMSELSQPSDTALVLAVVDPRLSG-YPLTSVIHMFNIAMMCVKEMGPARPTM 2998
Query: 594 SVVVKMINN 602
VV M+ N
Sbjct: 2999 REVVHMLTN 3025
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 196 bits (497), Expect = 1e-50
Identities = 120/337 (35%), Positives = 185/337 (54%), Gaps = 8/337 (2%)
Frame = +3
Query: 323 EVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKD 382
E +E AT + +GEGG GSVY+G L DG VA+K S +TQ F NE+NL+
Sbjct: 2142 EYIEVATERY--KTLIGEGGFGSVYRGTLNDGQEVAVKVRSSTSTQGTREFDNELNLLSA 2315
Query: 383 LQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHL---SVRRNLQQLTWEVRHKIILGTA 439
+QH+NLV LLG + +LVY ++ N SL D L +R + L W R I LG A
Sbjct: 2316 IQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAKRKI--LDWPTRLSIALGAA 2489
Query: 440 EGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFP-EDQSQISTAICGTLG 498
GLAYLH +IHRDIK SNILLD + K+ADFG ++ P E S +S + GT G
Sbjct: 2490 RGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAPQEGDSYVSLEVRGTAG 2669
Query: 499 YMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSF--VQNSCSILHMVWSLYGSNRLC 556
Y+ PEY +L+EK+D++SFGV+++E++SG+ + + S++ +++
Sbjct: 2670 YLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRTEWSLVEWATPYIRGSKVD 2849
Query: 557 DIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIADPTQPPFLN 616
+IVDP ++G Y AE +++++ L C + + RP M +V+ + + I + +
Sbjct: 2850 EIVDPGIKGGYHAEAMWRVVEVALQCLEPFSTYRPSMVAIVRELEDALIIENNASEYMKS 3029
Query: 617 FGS--GEFSRSSLPEESLQPGSNTQSSGDSMTESSEH 651
S G S + E+ + P + + + T+S H
Sbjct: 3030 IDSLGGSNRYSIVIEKRVLPSTTSTAESTITTQSLSH 3140
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 191 bits (485), Expect = 3e-49
Identities = 109/291 (37%), Positives = 167/291 (56%), Gaps = 10/291 (3%)
Frame = +2
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTT-QWADHFFNEVNLIKDL 383
L++ T+ F +GEG G VY L DG VA+K+L ++ + + F +V+++ L
Sbjct: 338 LKEKTDNFGSKALIGEGSYGRVYYATLNDGNAVAVKKLDVSSEPETNNEFLTQVSMVSRL 517
Query: 384 QHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQ------QLTWEVRHKIILG 437
++ N V+L G + G +L YE+ SLHD L R+ +Q L W R +I +
Sbjct: 518 KNDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVRIAVD 697
Query: 438 TAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQI-STAICGT 496
A GL YLHE+ Q IIHRDI+ SN+L+ +++ KIADF L+ P+ +++ ST + GT
Sbjct: 698 AARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLHSTRVLGT 877
Query: 497 LGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSL--YGSNR 554
GY APEY + G+LT+K+D+YSFGV+++ELL+G+ + W+ ++
Sbjct: 878 FGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSEDK 1057
Query: 555 LCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHK 605
+ VDP L+G YP + KL + LC Q AE RP MS+VVK + + K
Sbjct: 1058VKQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVKALQPLLK 1210
>AI967315
Length = 1308
Score = 179 bits (454), Expect = 1e-45
Identities = 115/289 (39%), Positives = 165/289 (56%), Gaps = 10/289 (3%)
Frame = +1
Query: 322 YEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLS--FNTTQWADHFFNEVNL 379
YE L ATN F N +G+GG VYKG L G +A+KRL+ + F E+
Sbjct: 118 YEELFHATNGFSSENMVGKGGYAEVYKGRLESGDEIAVKRLTRTCRDERKEKEFLTEIGT 297
Query: 380 IKDLQHKNLVKLLGCSITGPESLLVYEY-----VPNLSLHDHLSVRRNLQQLTWEVRHKI 434
I + H N++ LLGC I LV+E V +L +HD + L W+ R+KI
Sbjct: 298 IGHVCHSNVMPLLGCCIDNG-LYLVFELSTVGSVASL-IHDE-----KMAPLDWKTRYKI 456
Query: 435 ILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTA-I 493
+LGTA GL YLH+ Q +IIHRDIK SNILL ++F P+I+DFGLA+ P + S A I
Sbjct: 457 VLGTARGLHYLHKGCQRRIIHRDIKASNILLTEDFEPQISDFGLAKWLPSQWTHHSIAPI 636
Query: 494 CGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWS--LYG 551
GT G++APEY + G + EK D+++FGV ++E++SG+ V S LH W+ +
Sbjct: 637 EGTFGHLAPEYYMHGVVDEKTDVFAFGVFLLEVISGRKP---VDGSHQSLH-TWAKPILS 804
Query: 552 SNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMI 600
+ +VDP LEG Y + ++ LC +AS+ RP MS V++++
Sbjct: 805 KWEIEKLVDPRLEGCYDVTQFNRVAFAASLCIRASSTWRPTMSEVLEVM 951
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 178 bits (451), Expect = 3e-45
Identities = 106/299 (35%), Positives = 160/299 (53%), Gaps = 8/299 (2%)
Frame = +1
Query: 318 LNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEV 377
+ Y+ L KATN F NK+G+GG G+VY L G AIK++ Q + F E+
Sbjct: 1045 MEFSYQELAKATNNFSLDNKIGQGGFGAVYYAELR-GKKTAIKKMD---VQASTEFLCEL 1212
Query: 378 NLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILG 437
++ + H NLV+L+G + G LVYE++ N +L +L + L W R +I L
Sbjct: 1213 KVLTHVHHLNLVRLIGYCVEG-SLFLVYEHIDNGNLGQYLH-GSGKEPLPWSSRVQIALD 1386
Query: 438 TAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTL 497
A GL Y+HE + IHRD+K +NIL+D N K+ADFGL +L S + T + GT
Sbjct: 1387 AARGLEYIHEHTVPVYIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQTRLVGTF 1566
Query: 498 GYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRT----SFVQNSCSILHMVWSLYGSN 553
GYM PEY G ++ K D+Y+FGV++ EL+S K+ V S ++ + +
Sbjct: 1567 GYMPPEYAQYGDISPKIDVYAFGVVLFELISAKNAVLKTGELVAESKGLVALFEEALNKS 1746
Query: 554 RLCD----IVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIAD 608
CD +VDP L NYP + K+ ++G CT+ + LRP M +V + + + +
Sbjct: 1747 DPCDALRKLVDPRLGENYPIDSVLKIAQLGRACTRDNPLLRPSMRSLVVALMTLSSLTE 1923
>TC12343 similar to UP|CDK9_SCHPO (Q96WV9) Serine/threonine protein kinase
Cdk9 (Cyclin-dependent kinase Cdk9) , partial (7%)
Length = 723
Score = 177 bits (449), Expect = 5e-45
Identities = 92/170 (54%), Positives = 112/170 (65%)
Frame = +3
Query: 322 YEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIK 381
YE L AT FH NKLGEGG G V+KG L DG +A+K+LS + Q F NE L+
Sbjct: 213 YETLVAATKNFHAVNKLGEGGFGPVFKGKLNDGREIAVKKLSRRSNQGRTQFINEAKLLT 392
Query: 382 DLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEG 441
+QH+N+V L G G E LLVYEYVP SL L + +QL W+ R II G A G
Sbjct: 393 RVQHRNVVSLFGYCAHGSEKLLVYEYVPRESLDKLLFRSQKKEQLDWKRRFDIISGVARG 572
Query: 442 LAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIST 491
L YLHE+S IIHRDIK +NILLD+ + PKIADFGLAR+FPEDQ+ ++T
Sbjct: 573 LLYLHEDSHDCIIHRDIKAANILLDEKWVPKIADFGLARIFPEDQTHVNT 722
>BP032158
Length = 555
Score = 177 bits (448), Expect = 6e-45
Identities = 92/175 (52%), Positives = 118/175 (66%), Gaps = 1/175 (0%)
Frame = +2
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQ 384
++ ATN F +NK+GEGG G VYKG L +G +A+K+LS + Q F NE+ +I LQ
Sbjct: 29 IKAATNNFDPANKIGEGGFGPVYKGVLSEGDVIAVKQLSSKSKQGNREFINEIGMISALQ 208
Query: 385 HKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQ-QLTWEVRHKIILGTAEGLA 443
H NLVKL GC I G + LLVYEY+ N SL L + L W R KI +G A+GLA
Sbjct: 209 HPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNWRTRMKICVGIAKGLA 388
Query: 444 YLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLG 498
YLHEES+LKI+HRDIK +N+LLD + KI+DFGLA+L E+ + IST I GT+G
Sbjct: 389 YLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEEENTHISTRIAGTIG 553
>CN825263
Length = 663
Score = 175 bits (444), Expect = 2e-44
Identities = 94/190 (49%), Positives = 120/190 (62%), Gaps = 2/190 (1%)
Frame = +1
Query: 342 GSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPES 401
G G VYKG L DG VA+K L + + F EV ++ L H+NLVKL+G I
Sbjct: 1 GFGLVYKGILNDGRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQTR 180
Query: 402 LLVYEYVPNLSLHDHL-SVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKL 460
L+YE VPN S+ HL + L W R KI LG A GLAYLHE+S +IHRD K
Sbjct: 181 CLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFKS 360
Query: 461 SNILLDDNFTPKIADFGLAR-LFPEDQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSF 519
SNILL+ +FTPK++DFGLAR E IST + GT GY+APEY + G L K+D+YS+
Sbjct: 361 SNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYSY 540
Query: 520 GVLVIELLSG 529
GV+++ELL+G
Sbjct: 541 GVVLLELLTG 570
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 166 bits (421), Expect = 8e-42
Identities = 95/263 (36%), Positives = 145/263 (55%), Gaps = 3/263 (1%)
Frame = +3
Query: 338 LGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSIT 397
LG GG G VY G + GT VAIKR + + Q F E+ ++ L+H++LV L+G
Sbjct: 15 LGVGGFGKVYYGEVDGGTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIGYCEE 194
Query: 398 GPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRD 457
E +LVY+++ +L +HL + L W+ R +I +G A GL YLH ++ IIHRD
Sbjct: 195 NTEMILVYDHMAYGTLREHL-YKTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTIIHRD 371
Query: 458 IKLSNILLDDNFTPKIADFGLARLFPE-DQSQISTAICGTLGYMAPEYIVLGKLTEKADI 516
+K +NILLD+ + K++DFGL++ P D + +ST + G+ GY+ PEY +LT+K+D+
Sbjct: 372 VKTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTDKSDV 551
Query: 517 YSFGVLVIELLSGKS--RTSFVQNSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACK 574
YSFGV++ E+L + S + S+ Y L I+DP L+G E K
Sbjct: 552 YSFGVVLFEILCARPALNPSLAKEQVSLAEWASHCYNKGILDQILDPYLKGKIAPECFKK 731
Query: 575 LLKIGLLCTQASAELRPPMSVVV 597
+ + C RP M V+
Sbjct: 732 FAETAMKCVSDQGIERPSMGDVL 800
>TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P14.15
{Arabidopsis thaliana;}, partial (37%)
Length = 1079
Score = 159 bits (402), Expect = 1e-39
Identities = 89/225 (39%), Positives = 135/225 (59%), Gaps = 4/225 (1%)
Frame = +3
Query: 376 EVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKII 435
EV L+ + H+NLVKLLG G E +L+ EYVPN +L +HL R + L + R +I
Sbjct: 6 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRG-KILDFNQRLEIA 182
Query: 436 LGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFP--EDQSQISTAI 493
+ A GL YLH ++ +IIHRD+K SNILL ++ K+ADFG ARL P DQ+ IST +
Sbjct: 183 IDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKV 362
Query: 494 CGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSL--YG 551
GT+GY+ PEY+ +LT K+D+YSFG+L++E+L+G+ + + + + W+ Y
Sbjct: 363 KGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYN 542
Query: 552 SNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVV 596
+ +++DP++E A+ K+L + C RP M V
Sbjct: 543 EGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSV 677
>TC18923 UP|CAE45594 (CAE45594) S-receptor kinase-like protein 1, complete
Length = 2061
Score = 157 bits (396), Expect = 7e-39
Identities = 89/239 (37%), Positives = 135/239 (56%), Gaps = 12/239 (5%)
Frame = +1
Query: 388 LVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRN---------LQQLTWEVRHKIILGT 438
+ K L S+ G +L++ + L+ + ++N + L W R +II G
Sbjct: 1258 MTKKLAGSLAGIVALVICIIILGLATSTCIQRKKNERGDGDSTRSKLLDWNKRLQIIDGI 1437
Query: 439 AEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIST-AICGTL 497
A GL YLH++S+L+IIHRD+K SNILLD+ PKI+DFGLAR+F DQ + T + GT
Sbjct: 1438 ARGLLYLHQDSRLRIIHRDLKTSNILLDNEMNPKISDFGLARIFIGDQVEARTKRVMGTY 1617
Query: 498 GYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFV--QNSCSILHMVWSLYGSNRL 555
GYM PEY V G + K+D++SFGV+V+E++SGK F + ++L W L+
Sbjct: 1618 GYMPPEYAVHGSFSIKSDVFSFGVIVLEIISGKKIRKFYDPHHHLNLLSHAWRLWIEGSP 1797
Query: 556 CDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIADPTQPPF 614
++VD + E + E + + + LLC Q E RP M +V M+N ++ P+ P F
Sbjct: 1798 LELVDKLFEDSIIPTEILRYIHVALLCVQRRPETRPDMLSIVLMLNGEKELPKPSLPAF 1974
>TC9538 similar to UP|RLK5_ARATH (P47735) Receptor-like protein kinase 5
precursor , partial (10%)
Length = 1067
Score = 155 bits (391), Expect = 3e-38
Identities = 92/227 (40%), Positives = 131/227 (57%), Gaps = 13/227 (5%)
Frame = +2
Query: 387 NLVKLLGCSITGPESLLVYEYVPNLSLHDHL-------SVRRNLQQLT---WEVRHKIIL 436
N+V+LL C LLVYEY+ N SL L SV +QQ T W R KI +
Sbjct: 5 NIVRLLCCISNEASMLLVYEYLENHSLDKWLHLKPKSSSVSGVVQQYTVLDWPKRLKIAI 184
Query: 437 GTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLAR-LFPEDQSQISTAICG 495
G A+GL+Y+H + I+HRD+K SNILLD F K+ADFGLAR L + I + + G
Sbjct: 185 GAAQGLSYMHHDCSPPIVHRDVKTSNILLDKQFNAKVADFGLARMLIKPGELNIMSTVIG 364
Query: 496 TLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWS--LYGSN 553
T GY+APEY+ +++EK D+YSFGV+++EL +GK Q+S S+ W L GSN
Sbjct: 365 TFGYIAPEYVQTTRISEKVDVYSFGVVLLELTTGKEANYGDQHS-SLAEWAWRHILIGSN 541
Query: 554 RLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMI 600
+ ++E +Y +E C + K+G++CT RP M V+ ++
Sbjct: 542 VXDLLXKDVMEASY-IDEMCSVFKLGVMCTATLPATRPSMKEVLPIL 679
>TC19726 similar to UP|Q9XIS3 (Q9XIS3) Lectin-like protein kinase, partial
(30%)
Length = 728
Score = 154 bits (390), Expect = 3e-38
Identities = 89/234 (38%), Positives = 130/234 (55%), Gaps = 6/234 (2%)
Frame = +3
Query: 284 IVATIIFFVRKNVLRQRRERRQFGAFLFSENNSKLNMPYEVLEKATNYFHDSNKLGEGGS 343
++ +++++R R+ Q L S + Y L+KATN F + +KLG+GG
Sbjct: 24 VIGGLLWWIRLKKKREEGSNSQILGTLKSLPGTPREFNYVELKKATNNFDEKHKLGQGGY 203
Query: 344 GSVYKGALP-DGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESL 402
G VY+G LP + VA+K S + + D F +E+ +I L+HK+LV+L G L
Sbjct: 204 GVVYRGMLPKEKLEVAVKMFSRDKMKSTDDFLSELIIINRLRHKHLVRLQGWCHKNGVLL 383
Query: 403 LVYEYVPNLSLHDHLSVRRN---LQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIK 459
LVY+Y+PN SL H+ L+W +R+KII G A L YLH E K++HRD+K
Sbjct: 384 LVYDYMPNGSLDSHIFCEEGGTITTPLSWPLRYKIISGVASALNYLHNEYDQKVVHRDLK 563
Query: 460 LSNILLDDNFTPKIADFGLARLFPEDQSQIS--TAICGTLGYMAPEYIVLGKLT 511
SNI+LD F K+ DFGLAR ++ + + GT+GY+APE GK T
Sbjct: 564 ASNIMLDSEFNAKLGDFGLARALENEKISYTELEGVHGTMGYIAPECFHTGKAT 725
>AW719817
Length = 499
Score = 154 bits (388), Expect = 6e-38
Identities = 77/150 (51%), Positives = 109/150 (72%), Gaps = 2/150 (1%)
Frame = +1
Query: 453 IIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAPEYIVLGKLTE 512
I+HRDIK SNILLD +F PKI DFGLA+LFP+D + IST I GT GY+APEY + G+LT
Sbjct: 16 IVHRDIKASNILLDRDFNPKIGDFGLAKLFPDDITHISTRIAGTTGYLAPEYAMGGQLTM 195
Query: 513 KADIYSFGVLVIELLSGK--SRTSFVQNSCSILHMVWSLYGSNRLCDIVDPILEGNYPAE 570
KAD+YSFGVL++E++SGK +RT++ ++ +L W L+ +L ++VDP ++ ++P E
Sbjct: 196 KADVYSFGVLILEVVSGKGSARTNWGGSNKYLLEWAWQLHEEGKLLELVDPDMD-DFPEE 372
Query: 571 EACKLLKIGLLCTQASAELRPPMSVVVKMI 600
+ +K+ CTQA+A RP MS VV M+
Sbjct: 373 *VIRYMKVAFFCTQAAASRRPLMSQVVDML 462
>CB829158
Length = 553
Score = 150 bits (378), Expect = 8e-37
Identities = 83/177 (46%), Positives = 118/177 (65%), Gaps = 9/177 (5%)
Frame = +1
Query: 449 SQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAPEYIVLG 508
S+++IIHRDIK+SNILLD KIADFGLAR F ED+S ISTAI GTLGYMAPEY+ G
Sbjct: 1 SKVRIIHRDIKVSNILLDAKLRAKIADFGLARSFQEDKSHISTAIAGTLGYMAPEYLAHG 180
Query: 509 KLTEKADIYSFGVLVIELLSGK--SRTSFVQNSCSILHMVWSLYGSNRLCDIVDPILE-- 564
+LTEK D+YSFGVL++E+++G+ +R+ + + S++ + W + + + DP +E
Sbjct: 181 QLTEKVDVYSFGVLLLEIVTGRQNNRSKESEYTDSLILVAWEHFQTGTAEQLFDPNIELH 360
Query: 565 ----GNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHK-IADPTQPPFLN 616
GN E +++ IGLLC Q LRP MS V++M+ + + P+ PPFL+
Sbjct: 361 EAHNGNV-KNEILRVIHIGLLCIQEIPSLRPTMSKVLQMLTKKEELLVAPSNPPFLD 528
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,137,344
Number of Sequences: 28460
Number of extensions: 183427
Number of successful extensions: 1808
Number of sequences better than 10.0: 342
Number of HSP's better than 10.0 without gapping: 1583
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1608
length of query: 656
length of database: 4,897,600
effective HSP length: 96
effective length of query: 560
effective length of database: 2,165,440
effective search space: 1212646400
effective search space used: 1212646400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0076c.7


