

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
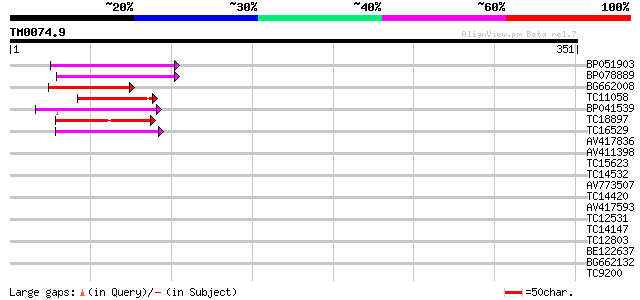
Query= TM0074.9
(351 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP051903 55 1e-08
BP078889 55 2e-08
BG662008 44 3e-05
TC11058 43 7e-05
BP041539 42 2e-04
TC18897 weakly similar to UP|Q9XEV3 (Q9XEV3) TIA-1 related prote... 42 2e-04
TC16529 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, p... 40 6e-04
AV417836 39 0.002
AV411398 38 0.003
TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding p... 38 0.003
TC14532 similar to UP|Q9FYA8 (Q9FYA8) SC35-like splicing factor ... 37 0.004
AV773507 37 0.004
TC14420 similar to UP|ROC3_NICSY (P19682) 28 kDa ribonucleoprote... 37 0.004
AV417593 36 0.008
TC12531 similar to UP|Q9ASQ8 (Q9ASQ8) At1g22910/F19G10_13, parti... 36 0.008
TC14147 similar to UP|ROC1_NICSY (Q08935) 29 kDa ribonucleoprote... 35 0.025
TC12803 similar to UP|Q9FYW5 (Q9FYW5) BAC19.11, partial (32%) 33 0.055
BE122637 33 0.055
BG662132 32 0.12
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 32 0.12
>BP051903
Length = 342
Score = 55.5 bits (132), Expect = 1e-08
Identities = 33/80 (41%), Positives = 46/80 (57%)
Frame = +2
Query: 26 NIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAM 85
++ VDG+ T + Q+R LF +FG VV+VFV + K R +FGFV F S A+
Sbjct: 53 HLHSLVDGIPDRTDYYQIRGLFAQFGRVVNVFVQRQRKIGRLFRFGFVRFASLEVASKAI 232
Query: 86 RSLHGTRLNSFFLSINPARF 105
L+G RL LS++ ARF
Sbjct: 233 DFLNGFRLGDAALSVSMARF 292
>BP078889
Length = 474
Score = 55.1 bits (131), Expect = 2e-08
Identities = 30/76 (39%), Positives = 43/76 (56%)
Frame = +1
Query: 30 FVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSLH 89
FVDG++ ++ +R LF FG + ++FV + K RR +FGFV F S+ A L
Sbjct: 247 FVDGIEEAIEYHHLRGLFALFGRISNLFVQRKHKFGRRFRFGFVRFFSEEQAAAVAFVLD 426
Query: 90 GTRLNSFFLSINPARF 105
G R+ LS+ PARF
Sbjct: 427 GLRVGGTALSVAPARF 474
>BG662008
Length = 457
Score = 44.3 bits (103), Expect = 3e-05
Identities = 21/53 (39%), Positives = 34/53 (63%)
Frame = +3
Query: 25 ENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQS 77
+++ LFVDG+ +SQ+R LF G++++VFV + + R +FGFV F S
Sbjct: 93 QSVTLFVDGISDFVDYSQIRGLFSHIGSLLNVFVQRQRRIGRAFRFGFVRFPS 251
>TC11058
Length = 539
Score = 43.1 bits (100), Expect = 7e-05
Identities = 24/49 (48%), Positives = 33/49 (66%)
Frame = +1
Query: 43 VRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSLHGT 91
VR LF RFG V SV + + +K+ RR + F+ + S+R+ AAM SLHGT
Sbjct: 22 VRYLFARFGNVASVRLLQDLKQGRRRAYAFISYLSERERDAAM-SLHGT 165
>BP041539
Length = 521
Score = 42.0 bits (97), Expect = 2e-04
Identities = 28/83 (33%), Positives = 43/83 (51%), Gaps = 5/83 (6%)
Frame = +2
Query: 17 HERSEVAEENIR-----LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFG 71
HE VA+E + L+V LD + SQ+ LF + G VVSV V + + + +G
Sbjct: 242 HETDAVAQEANQPMPTSLYVGDLDTEINDSQLYDLFNQIGQVVSVRVCRDLATQQSLGYG 421
Query: 72 FVLFQSKRDGIAAMRSLHGTRLN 94
+V F + +D A+ L+ T LN
Sbjct: 422 YVNFTNPKDAATALDVLNFTPLN 490
>TC18897 weakly similar to UP|Q9XEV3 (Q9XEV3) TIA-1 related protein, partial
(5%)
Length = 481
Score = 41.6 bits (96), Expect = 2e-04
Identities = 21/63 (33%), Positives = 39/63 (61%), Gaps = 1/63 (1%)
Frame = +1
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRRE-KFGFVLFQSKRDGIAAMRS 87
LF+DG+ T + +R LF + G++++VF+ K +R+RR +FGFV + ++ A+
Sbjct: 283 LFIDGISEPTNYYHIRGLFGQIGSLLNVFLQKQ-RRIRRNFRFGFVRYSTRDVASMAINL 459
Query: 88 LHG 90
+G
Sbjct: 460 FNG 468
>TC16529 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(22%)
Length = 761
Score = 40.0 bits (92), Expect = 6e-04
Identities = 24/67 (35%), Positives = 37/67 (54%)
Frame = +1
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L+V LD++ SQ+ LF + G VVSV V + + R +G+V F + D A+ L
Sbjct: 337 LYVGDLDQNVNDSQLYDLFNQVGQVVSVRVCRDLTTRRSLGYGYVNFSNPPDAARALDVL 516
Query: 89 HGTRLNS 95
+ T LN+
Sbjct: 517 NFTPLNN 537
>AV417836
Length = 429
Score = 38.5 bits (88), Expect = 0.002
Identities = 29/101 (28%), Positives = 40/101 (38%), Gaps = 12/101 (11%)
Frame = +1
Query: 3 PTGSARAPPPRWSCHERSEVAEENIR------------LFVDGLDRDTKHSQVRKLFERF 50
P +R PPPR R ++ R L V L D + +R FER+
Sbjct: 25 PRRRSRTPPPRG----RKRYDDDRTRSYRDRRSPLPSGLLVRNLPLDARPEDLRGPFERY 192
Query: 51 GTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSLHGT 91
G V V++PK FGFV ++ D A L+ T
Sbjct: 193 GPVKDVYLPKNYYTGEPRGFGFVKYRFGEDAAEAKHHLNHT 315
>AV411398
Length = 427
Score = 37.7 bits (86), Expect = 0.003
Identities = 27/93 (29%), Positives = 39/93 (41%)
Frame = +3
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
RL + + VR LFER+GTV+ V + + R FV S + + A+
Sbjct: 81 RLVAQNVPWTSTPEDVRSLFERYGTVLEVEL-SMYNKTRSRGLAFVEMSSPEEALEALNK 257
Query: 88 LHGTRLNSFFLSINPARFVDRKYKWNPPAKQHE 120
L L +N AR +K K PP Q +
Sbjct: 258 LESYEFEGRVLKLNYAR--PKKKKAPPPVVQRK 350
>TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (88%)
Length = 683
Score = 37.7 bits (86), Expect = 0.003
Identities = 22/67 (32%), Positives = 29/67 (42%)
Frame = +1
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+LF+ GL + F FGTV V R FGFV F S+ +A+ S
Sbjct: 187 KLFIGGLSYGVDDKSLEDAFSSFGTVAEARVIVDRDTGRSRGFGFVSFDSEESASSALSS 366
Query: 88 LHGTRLN 94
+ G LN
Sbjct: 367 MDGQDLN 387
>TC14532 similar to UP|Q9FYA8 (Q9FYA8) SC35-like splicing factor SCL30a, 30a
kD, partial (71%)
Length = 1073
Score = 37.4 bits (85), Expect = 0.004
Identities = 23/86 (26%), Positives = 35/86 (39%)
Frame = +3
Query: 5 GSARAPPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKR 64
G R+P PR + L V L D + +R+ F +FG + +++PK
Sbjct: 102 GGGRSPSPRGGGRYGPRPRDLPTSLLVRNLRHDCRPEDLRRPFGQFGPLKDIYLPKDYYT 281
Query: 65 VRREKFGFVLFQSKRDGIAAMRSLHG 90
+ FGFV F D A + G
Sbjct: 282 GQPRGFGFVQFVDPADAADAKYHMDG 359
>AV773507
Length = 496
Score = 37.4 bits (85), Expect = 0.004
Identities = 19/72 (26%), Positives = 33/72 (45%)
Frame = +2
Query: 33 GLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSLHGTR 92
GL D Q+ F+R G ++ + R FGF+ F +R A++ +HG
Sbjct: 29 GLSWDVTERQLEHAFDRDGKILECQIMMERDTGRPRGFGFITFADRRGMEDAIKEMHGRE 208
Query: 93 LNSFFLSINPAR 104
+ +S+N A+
Sbjct: 209 IGDRIISVNKAQ 244
>TC14420 similar to UP|ROC3_NICSY (P19682) 28 kDa ribonucleoprotein,
chloroplast precursor (28RNP), partial (34%)
Length = 637
Score = 37.4 bits (85), Expect = 0.004
Identities = 26/85 (30%), Positives = 40/85 (46%)
Frame = +1
Query: 19 RSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSK 78
RS A+ R++V L D +SQ+ ++F G VVS V + R FGFV +
Sbjct: 16 RSFDAQPAFRVYVGNLPWDFDNSQLEQVFSEHGKVVSARVVYDRETGRSRGFGFVTMSDE 195
Query: 79 RDGIAAMRSLHGTRLNSFFLSINPA 103
+ A+ +L G L + +N A
Sbjct: 196 AELNDAIAALDGQSLGGRTIRVNVA 270
>AV417593
Length = 423
Score = 36.2 bits (82), Expect = 0.008
Identities = 20/76 (26%), Positives = 35/76 (45%), Gaps = 2/76 (2%)
Frame = +2
Query: 9 APPPRWSCHER--SEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVR 66
AP W+ + S + + ++V L ++ QVRKLFE G + V +P
Sbjct: 194 APTVSWADPKNADSSASSQVKAVYVKNLPKNVTQEQVRKLFEHHGKITKVVLPPPKSGQE 373
Query: 67 REKFGFVLFQSKRDGI 82
+ + GFV F + + +
Sbjct: 374 KNRIGFVHFAERSNAM 421
>TC12531 similar to UP|Q9ASQ8 (Q9ASQ8) At1g22910/F19G10_13, partial (39%)
Length = 688
Score = 36.2 bits (82), Expect = 0.008
Identities = 26/84 (30%), Positives = 41/84 (47%), Gaps = 7/84 (8%)
Frame = -1
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
++FV GL +T+ ++K FE+FG ++ V R + +GFV F R+ AAMR+
Sbjct: 358 KVFVGGLAWETQKETMKKYFEQFGDILEAVVITDKATGRSKGYGFVTF---REPEAAMRA 188
Query: 88 -------LHGTRLNSFFLSINPAR 104
+ G R N S+ R
Sbjct: 187 CVDAAPVIDGRRANCNLASLGVQR 116
>TC14147 similar to UP|ROC1_NICSY (Q08935) 29 kDa ribonucleoprotein A,
chloroplast precursor (CP29A), partial (63%)
Length = 1602
Score = 34.7 bits (78), Expect = 0.025
Identities = 24/76 (31%), Positives = 35/76 (45%)
Frame = +3
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
R+ V L ++ + LF G VV V + R FGFV F S + +A+RS
Sbjct: 741 RVHVGNLAWGVDNAALESLFREQGRVVDAKVIYDRESGRSRGFGFVTFSSPDEVNSAIRS 920
Query: 88 LHGTRLNSFFLSINPA 103
L G LN + ++ A
Sbjct: 921 LDGADLNGRAIKVSQA 968
>TC12803 similar to UP|Q9FYW5 (Q9FYW5) BAC19.11, partial (32%)
Length = 892
Score = 33.5 bits (75), Expect = 0.055
Identities = 20/76 (26%), Positives = 37/76 (48%)
Frame = +3
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
R+FV GL Q+R+L + FG + S + + + + +GF ++Q A S
Sbjct: 93 RIFVGGLPYYFAEEQIRELLQAFGPLRSFDLVRDKETGNSKGYGFCIYQDPAVTDMACAS 272
Query: 88 LHGTRLNSFFLSINPA 103
L+G ++ L++ A
Sbjct: 273 LNGLKMGDKTLTVRRA 320
>BE122637
Length = 332
Score = 33.5 bits (75), Expect = 0.055
Identities = 19/56 (33%), Positives = 25/56 (43%)
Frame = +2
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAA 84
L V + D + ++R FERFG V V++PK F FV F D A
Sbjct: 29 LLVRNIPLDCRADEIRVPFERFGPVRDVYIPKDYYSGEPRGFAFVQFVDPYDASEA 196
>BG662132
Length = 409
Score = 32.3 bits (72), Expect = 0.12
Identities = 27/93 (29%), Positives = 43/93 (46%), Gaps = 16/93 (17%)
Frame = +3
Query: 17 HERSEVAE--------------ENIRLFVDGLDRDTKHSQVRKLFERFGTV--VSVFVPK 60
HE SE+ + + +LFV G+ RDT ++ F ++G V ++ + +
Sbjct: 9 HEESEIGDLGFCTVTMNWKMDSDRAKLFVGGISRDTTEEILKHHFTKYGNVSDSTISIDR 188
Query: 61 TIKRVRREKFGFVLFQSKRDGIAAMRSLHGTRL 93
T + R FGFV F D AA R+L T +
Sbjct: 189TTRSPR--GFGFVTF---FDLSAAHRALQDTHV 272
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 32.3 bits (72), Expect = 0.12
Identities = 18/69 (26%), Positives = 38/69 (54%), Gaps = 3/69 (4%)
Frame = +1
Query: 21 EVAEENI---RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQS 77
+VA+E++ ++FV GL DT + + F+++G + + + +GF+LF+S
Sbjct: 439 KVADEDVSHRKIFVHGLGWDTTATTLVYAFQQYGAIEDCKAVTDKVTGKSKGYGFILFKS 618
Query: 78 KRDGIAAMR 86
+R A++
Sbjct: 619 RRGARNALK 645
Score = 27.7 bits (60), Expect = 3.0
Identities = 28/127 (22%), Positives = 50/127 (39%), Gaps = 18/127 (14%)
Frame = +2
Query: 91 TRLNSFFLSINPARFVDRKYKWNPPAKQHEELPG-------RKKIRQEWRIKHRVGAPFS 143
T +++ S++ + + NP + P R R WR++ F
Sbjct: 32 TLVSTLTHSLHSKNHQTKPWLTNPSPRSESSSPNPLNRRSYRSSCRTSWRLR------FK 193
Query: 144 GTGGFSQESQKVWR-----------VKKSTRKSPEKKVEEQERSGMTMNQVRVGLNMDLS 192
T ++ +K WR VKK +K +K+ +++R TMN+ R N
Sbjct: 194 STKKSKKK*RKKWRRKKKRRKRKRRVKKKKKKKKKKRKWKRKRRRKTMNRSRSFSNPWGR 373
Query: 193 RMAIASL 199
++ASL
Sbjct: 374 SRSLASL 394
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,294,099
Number of Sequences: 28460
Number of extensions: 81573
Number of successful extensions: 480
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 478
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 480
length of query: 351
length of database: 4,897,600
effective HSP length: 91
effective length of query: 260
effective length of database: 2,307,740
effective search space: 600012400
effective search space used: 600012400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0074.9


