

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
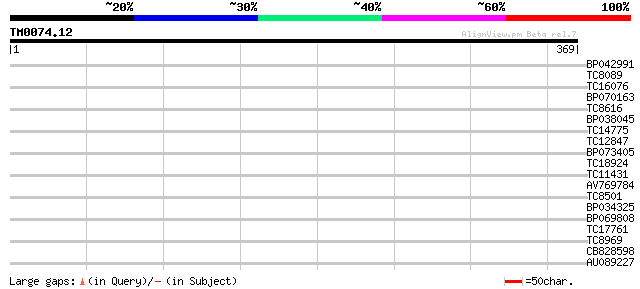
Query= TM0074.12
(369 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP042991 33 0.099
TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinog... 30 0.49
TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG... 30 0.64
BP070163 29 1.1
TC8616 UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like pro... 29 1.4
BP038045 28 1.9
TC14775 PIR|S14181|S14181 DNA-directed RNA polymerase largest c... 28 1.9
TC12847 similar to GB|AAM66074.1|21595125|AY088542 endosperm-spe... 28 2.4
BP073405 28 2.4
TC18924 homologue to UP|Q94FN3 (Q94FN3) Phosphatidylinositol tra... 28 2.4
TC11431 28 2.4
AV769784 28 3.2
TC8501 similar to UP|Q9FFY9 (Q9FFY9) H-protein promoter binding ... 23 3.5
BP034325 27 5.4
BP069808 27 5.4
TC17761 weakly similar to UP|Q38889 (Q38889) OBP33pep (Fragment)... 27 5.4
TC8969 27 7.1
CB828598 27 7.1
AU089227 27 7.1
>BP042991
Length = 534
Score = 32.7 bits (73), Expect = 0.099
Identities = 25/81 (30%), Positives = 35/81 (42%), Gaps = 19/81 (23%)
Frame = +1
Query: 10 SILSAQRDPRSIFYSPPS----QPNTS-----PY-YCHNVPSYYPQTHSS---------L 50
S S+ P +YSP QPN S PY Y H+ P + Q S L
Sbjct: 292 SFTSSYPQPAPDYYSPSQPYIPQPNVSSQHYYPYIYPHSSPPHDTQLSFSSPRAKSQPKL 471
Query: 51 QLSEFNGENPEYWLFMADCYF 71
+++ FNG +P WL + +F
Sbjct: 472 KITVFNGSDPTMWLLNTETFF 534
>TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (43%)
Length = 890
Score = 30.4 bits (67), Expect = 0.49
Identities = 16/38 (42%), Positives = 20/38 (52%)
Frame = +2
Query: 12 LSAQRDPRSIFYSPPSQPNTSPYYCHNVPSYYPQTHSS 49
LS QR S + PP P TSP + + PS P T +S
Sbjct: 176 LSRQRCSSSPWLLPPQTPTTSPAFSPSTPSSPPSTTTS 289
>TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG
{Arabidopsis thaliana;} , partial (24%)
Length = 1048
Score = 30.0 bits (66), Expect = 0.64
Identities = 12/42 (28%), Positives = 23/42 (54%)
Frame = -3
Query: 78 WETRFQLLAIHLGGKALTWFRLSHQQIHSWEEFKVAFRLYFV 119
W R Q ++IH+ + WFR++ + I+ F + +R Y +
Sbjct: 557 WTHRCQEISIHIPANRVYWFRMTLECINLSVTFDIKYRHYLI 432
>BP070163
Length = 257
Score = 29.3 bits (64), Expect = 1.1
Identities = 15/50 (30%), Positives = 24/50 (48%)
Frame = +3
Query: 40 PSYYPQTHSSLQLSEFNGENPEYWLFMADCYFLQNPYPWETRFQLLAIHL 89
P+YY +SS L + G + L + Y L +PW F+L+ H+
Sbjct: 102 PNYYIIQYSSNYLVQ*CGGHSPEKLILDPIYILPKMFPWTINFKLIIPHI 251
>TC8616 UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like protein III,
complete
Length = 2302
Score = 28.9 bits (63), Expect = 1.4
Identities = 27/95 (28%), Positives = 42/95 (43%)
Frame = +3
Query: 228 DVNKTFVLKTTDLVLRNPRQVFGNLLEGKFVQKNNEAQFEKHFDKLLMQFHNSSTTVDVQ 287
D NK + T D +R Q F KF A+ +H D +S+T +DVQ
Sbjct: 624 DPNKLMQVTTMDRYVRYHVQEFEKSFAIKFPACTMAAK--RHID-------SSTTILDVQ 776
Query: 288 KIGVPCPKVFEKMLVTRNTRICVDNTFEKLLETYI 322
+G+ + L+TR ++ DN E L + +I
Sbjct: 777 GVGLKNFTKSARELITRLQKVDGDNYPETLCQMFI 881
>BP038045
Length = 556
Score = 28.5 bits (62), Expect = 1.9
Identities = 19/57 (33%), Positives = 25/57 (43%), Gaps = 2/57 (3%)
Frame = -3
Query: 30 NTSPYYCHNVPSYYPQTHSSLQLSEFNGENPEYWLFMADCYFLQNP--YPWETRFQL 84
+T PYYC +P ++S L E +NP +L F YP TRF L
Sbjct: 515 STVPYYCKQIP*PNSFSYSQLLADEAAPKNPSRFLTKFTILFPSYVILYPMTTRFVL 345
>TC14775 PIR|S14181|S14181 DNA-directed RNA polymerase largest chain
(isoform B1) - soybean (fragment) {Glycine
max;} , partial (22%)
Length = 848
Score = 28.5 bits (62), Expect = 1.9
Identities = 17/43 (39%), Positives = 23/43 (52%), Gaps = 3/43 (6%)
Frame = +1
Query: 23 YSP--PSQPNTSPYYCHNVPSYYPQTHSSLQLSEFN-GENPEY 62
YSP P+ TSP Y PSY P + + S +N G +P+Y
Sbjct: 190 YSPTSPNYSPTSPSYSPTSPSYSPSSPTYSPSSPYNSGVSPDY 318
Score = 28.1 bits (61), Expect = 2.4
Identities = 16/31 (51%), Positives = 17/31 (54%), Gaps = 2/31 (6%)
Frame = +1
Query: 18 PRSIFYSP--PSQPNTSPYYCHNVPSYYPQT 46
P S YSP PS TSP Y PSY PQ+
Sbjct: 34 PTSPGYSPTSPSYSPTSPSYSPTSPSYNPQS 126
>TC12847 similar to GB|AAM66074.1|21595125|AY088542 endosperm-specific
protein-like protein {Arabidopsis thaliana;} , partial
(11%)
Length = 245
Score = 28.1 bits (61), Expect = 2.4
Identities = 17/41 (41%), Positives = 20/41 (48%)
Frame = +1
Query: 9 SSILSAQRDPRSIFYSPPSQPNTSPYYCHNVPSYYPQTHSS 49
SS+LS P PPS P TSP + P+ P T SS
Sbjct: 76 SSLLSPSPPP------PPSPPTTSPRFSPPTPTTAPTTPSS 180
>BP073405
Length = 400
Score = 28.1 bits (61), Expect = 2.4
Identities = 18/63 (28%), Positives = 29/63 (45%)
Frame = -3
Query: 69 CYFLQNPYPWETRFQLLAIHLGGKALTWFRLSHQQIHSWEEFKVAFRLYFVYNRPNGFST 128
CYFL+ PW+ +FQLL + ++ + + S + + R Y + R FST
Sbjct: 383 CYFLEICLPWKYKFQLLILQSHNQSCEIQNIGLWMLQSSTQVQPLCRHYSIQER--FFST 210
Query: 129 TTV 131
V
Sbjct: 209 IQV 201
>TC18924 homologue to UP|Q94FN3 (Q94FN3) Phosphatidylinositol transfer-like
protein II, complete
Length = 1976
Score = 28.1 bits (61), Expect = 2.4
Identities = 26/95 (27%), Positives = 42/95 (43%)
Frame = +3
Query: 228 DVNKTFVLKTTDLVLRNPRQVFGNLLEGKFVQKNNEAQFEKHFDKLLMQFHNSSTTVDVQ 287
D K + T D ++ + F + KF + A+ KH D+ S+T +DVQ
Sbjct: 501 DATKLMQVTTMDRYIKYHVKEFERTFDLKFAACSIAAK--KHIDQ-------STTILDVQ 653
Query: 288 KIGVPCPKVFEKMLVTRNTRICVDNTFEKLLETYI 322
+G+ + L+TR +I DN E L +I
Sbjct: 654 GVGLKNFNKHARELITRLQKIDGDNYPETLNRMFI 758
>TC11431
Length = 551
Score = 28.1 bits (61), Expect = 2.4
Identities = 16/46 (34%), Positives = 22/46 (47%)
Frame = -2
Query: 17 DPRSIFYSPPSQPNTSPYYCHNVPSYYPQTHSSLQLSEFNGENPEY 62
DP+ + P SQ N + HN P + LQL F G+N +Y
Sbjct: 364 DPKPSYNRPRSQTNN*E*HTHN-----PYSLRLLQLHSFRGQNHQY 242
>AV769784
Length = 535
Score = 27.7 bits (60), Expect = 3.2
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = +3
Query: 22 FYSPPSQPNTSPYYCHNVPSYYPQTHSSLQL 52
F+ P P+TS H+ P Y PQ+HSS+ +
Sbjct: 306 FHHLPPSPSTSSQ--HSPPPYPPQSHSSVPI 392
>TC8501 similar to UP|Q9FFY9 (Q9FFY9) H-protein promoter binding factor-1
(Telomeric DNA-binding protein 1), partial (5%)
Length = 616
Score = 23.5 bits (49), Expect(2) = 3.5
Identities = 10/34 (29%), Positives = 16/34 (46%)
Frame = -3
Query: 45 QTHSSLQLSEFNGENPEYWLFMADCYFLQNPYPW 78
Q L + F+ E+WLF+ + P+PW
Sbjct: 407 QAEFLLDIF*FSPSEAEFWLFIGAD*SIGPPWPW 306
Score = 22.3 bits (46), Expect(2) = 3.5
Identities = 11/30 (36%), Positives = 13/30 (42%)
Frame = -2
Query: 18 PRSIFYSPPSQPNTSPYYCHNVPSYYPQTH 47
PR+ + P Q S CHN S P H
Sbjct: 474 PRTSNQNSPLQKLNSMSICHNFSSRVPLRH 385
>BP034325
Length = 417
Score = 26.9 bits (58), Expect = 5.4
Identities = 14/40 (35%), Positives = 19/40 (47%)
Frame = +3
Query: 2 VLNYDVLSSILSAQRDPRSIFYSPPSQPNTSPYYCHNVPS 41
+L D S + +AQ P S P QP++ P HN S
Sbjct: 225 LLCLDFRSGMCAAQNHPHRTHTSKPHQPHSIPTATHNCKS 344
>BP069808
Length = 291
Score = 26.9 bits (58), Expect = 5.4
Identities = 17/50 (34%), Positives = 23/50 (46%), Gaps = 3/50 (6%)
Frame = +3
Query: 43 YPQTHSSLQLSEFNGENPEYWLFMADCYFLQNPYPWETRFQLL---AIHL 89
Y Q SLQ + + + EY L F+ PYPW + L +IHL
Sbjct: 114 YSQKGLSLQRAGWLRHSYEYLLHSLPSSFILYPYPWSLSYNNLL*SSIHL 263
>TC17761 weakly similar to UP|Q38889 (Q38889) OBP33pep (Fragment), partial
(27%)
Length = 549
Score = 26.9 bits (58), Expect = 5.4
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = -2
Query: 87 IHLGGKALTWFRLSHQQIHSWEEFKV 112
+H GG + + SHQ+ H W+ F V
Sbjct: 176 MHQGGLKMNFHSQSHQRNHHWQHFLV 99
>TC8969
Length = 892
Score = 26.6 bits (57), Expect = 7.1
Identities = 9/25 (36%), Positives = 19/25 (76%)
Frame = -1
Query: 325 LLDWQRMRPMNLTSEIALSSLSGPG 349
+L+W+ M+P N++ +I L++L+ G
Sbjct: 865 ILNWRLMQPPNVSQKITLNNLAAKG 791
>CB828598
Length = 559
Score = 26.6 bits (57), Expect = 7.1
Identities = 15/47 (31%), Positives = 22/47 (45%)
Frame = -1
Query: 107 WEEFKVAFRLYFVYNRPNGFSTTTVKYGFEKSSLVEIQRSSEQVSPH 153
W+ K ++ V + NG +TTT + G +SSL R S H
Sbjct: 352 WQASKTLLKVN*VITKWNGTTTTTERCGCTRSSLRARYRRRRHSSAH 212
>AU089227
Length = 447
Score = 26.6 bits (57), Expect = 7.1
Identities = 13/48 (27%), Positives = 21/48 (43%)
Frame = -1
Query: 60 PEYWLFMADCYFLQNPYPWETRFQLLAIHLGGKALTWFRLSHQQIHSW 107
PE F + +F +P R + + LGG ++ H +HSW
Sbjct: 246 PELVRFSSFIFFXSSPSVTXLR*SEVFLVLGGSLVSLILCKHASVHSW 103
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,824,356
Number of Sequences: 28460
Number of extensions: 105070
Number of successful extensions: 636
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 626
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 633
length of query: 369
length of database: 4,897,600
effective HSP length: 92
effective length of query: 277
effective length of database: 2,279,280
effective search space: 631360560
effective search space used: 631360560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0074.12


