

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
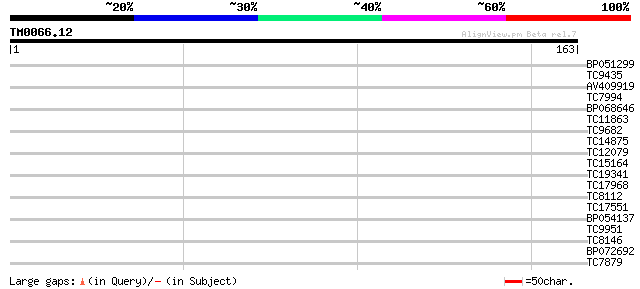
Query= TM0066.12
(163 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP051299 32 0.054
TC9435 similar to UP|Q9AXJ7 (Q9AXJ7) Phosphatase-like protein Mt... 30 0.21
AV409919 29 0.35
TC7994 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomeras... 29 0.35
BP068646 29 0.35
TC11863 weakly similar to UP|O22458 (O22458) Hydroxyproline-rich... 29 0.46
TC9682 similar to UP|Q8VZC0 (Q8VZC0) DTDP-glucose 4-6-dehydratas... 28 0.60
TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%) 28 0.60
TC12079 28 0.60
TC15164 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, p... 28 0.78
TC19341 similar to UP|Q9CAD1 (Q9CAD1) GMP synthase, partial (28%) 28 1.0
TC17968 similar to PIR|F86340|F86340 protein F2D10.34 [imported]... 28 1.0
TC8112 similar to UP|ODPA_PEA (P52902) Pyruvate dehydrogenase E1... 28 1.0
TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, part... 28 1.0
BP054137 27 1.3
TC9951 homologue to UP|TBB3_SOYBN (P28551) Tubulin beta chain (F... 27 1.7
TC8146 similar to UP|Q94FP3 (Q94FP3) Succinate dehydrogenase sub... 27 1.7
BP072692 27 1.7
TC7879 weakly similar to GB|AAM16239.1|20334756|AY093978 At2g321... 27 1.7
>BP051299
Length = 539
Score = 32.0 bits (71), Expect = 0.054
Identities = 33/151 (21%), Positives = 60/151 (38%), Gaps = 8/151 (5%)
Frame = +3
Query: 7 PPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPN---SWD 63
PPP +P+ + PSS ++S+ + F + S + +PN S
Sbjct: 105 PPPSSPSTATTHPPPSS-----------SSSATSVLFPISSTTPSAATTLFSPNPPPSAS 251
Query: 64 PLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHG-----THLLSPHEVRA 118
P +T+ P + T S+S P +S + + V + H T L +P +
Sbjct: 252 PPPSTVNSP--SSESVTASSASPPPKTAVISSSATPLFVSISHSPNPPTTLLFTPPLSPS 425
Query: 119 LMVPTSLTASLNSWVFIGKLISVVLFRFRLR 149
+ +P ++T L+ + V R RL+
Sbjct: 426 VSIPGTMTTRLSESLACMMTKGTVFLRLRLK 518
>TC9435 similar to UP|Q9AXJ7 (Q9AXJ7) Phosphatase-like protein Mtc923,
partial (17%)
Length = 486
Score = 30.0 bits (66), Expect = 0.21
Identities = 28/123 (22%), Positives = 48/123 (38%)
Frame = +3
Query: 8 PPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLSN 67
PP + + P S+P + P PTTT ++T PF P S +PL++
Sbjct: 135 PPSSTTTTTP*STPPQP--PPQPPPPTTTKTKTTPFH--------------PPSLNPLTS 266
Query: 68 TLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSPHEVRALMVPTSLTA 127
TL R P + P+ + ++ R+ + S V ++ P +
Sbjct: 267 TLPRLPHARLPLPTPPLAPPTSSDPAGHSARNLPPRLPEPAAVESARSVISIADPVPILN 446
Query: 128 SLN 130
S+N
Sbjct: 447 SMN 455
>AV409919
Length = 423
Score = 29.3 bits (64), Expect = 0.35
Identities = 18/49 (36%), Positives = 24/49 (48%), Gaps = 6/49 (12%)
Frame = +3
Query: 6 KPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETE------PFRLDQA 48
+PPP +P+ P +S +S+ P PTTTS T P LD A
Sbjct: 276 RPPPSSPSSPPPSTSSASS----SAPPPTTTSPTTRSWPRKIPVTLDHA 410
>TC7994 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomerase
precursor , partial (81%)
Length = 1355
Score = 29.3 bits (64), Expect = 0.35
Identities = 39/128 (30%), Positives = 49/128 (37%)
Frame = +2
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWD 63
L PP PN S P SP +T D P+TTSS A S P S
Sbjct: 347 LNYAPPPLPNPSVPSPSPKTTSRDSPPTRPSTTSS--------PAWSSASAPAPPPPSSS 502
Query: 64 PLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSPHEVRALMVPT 123
P S + + P ++ SS+SP S+R S A R T + P + PT
Sbjct: 503 PSSASFSPP-----ANSPTSSASPPPSARRSRRDPS-ASRSPSSTTI--PASISPSTAPT 658
Query: 124 SLTASLNS 131
T S S
Sbjct: 659 RSTLSSTS 682
>BP068646
Length = 491
Score = 29.3 bits (64), Expect = 0.35
Identities = 11/29 (37%), Positives = 15/29 (50%)
Frame = +2
Query: 9 PCNPNCSKPESSPSSTWFDMELPLPTTTS 37
PC+P+CS E + W LP PT +
Sbjct: 227 PCSPHCSPHEFLTNQYWVPPSLPYPTAVT 313
>TC11863 weakly similar to UP|O22458 (O22458) Hydroxyproline-rich
glycoprotein gas28p precursor, partial (7%)
Length = 341
Score = 28.9 bits (63), Expect = 0.46
Identities = 14/32 (43%), Positives = 19/32 (58%)
Frame = +1
Query: 7 PPPCNPNCSKPESSPSSTWFDMELPLPTTTSS 38
P P NP+ S P +PS+ LPLPT ++S
Sbjct: 166 PDPSNPSQSNPFHTPSNPKTPPLLPLPTPSAS 261
>TC9682 similar to UP|Q8VZC0 (Q8VZC0) DTDP-glucose 4-6-dehydratase-like
protein, partial (29%)
Length = 731
Score = 28.5 bits (62), Expect = 0.60
Identities = 19/57 (33%), Positives = 27/57 (47%), Gaps = 1/57 (1%)
Frame = +1
Query: 58 APNSWDP-LSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSP 113
A W+P L +T P D DPS++S V+ S SES + ++ LSP
Sbjct: 229 ATEKWEPKLHHTPPNPPNTPDPSPDPSTTSSVNSVSSSSSSESSSAPPSSSSNPLSP 399
>TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%)
Length = 1240
Score = 28.5 bits (62), Expect = 0.60
Identities = 28/90 (31%), Positives = 35/90 (38%)
Frame = +1
Query: 8 PPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLSN 67
PP NP S +P+S P +TS T P + S AP++ PL+
Sbjct: 136 PPSNPKTSTSTPTPASP--------PASTSPLTPPTTSPSSSTS----TAAPSASAPLTT 279
Query: 68 TLTRPLRLHDQDTDPSSSSPSFIVTVSQRS 97
T T SSSPS T S RS
Sbjct: 280 TPTTATSTTSPRRQTPSSSPS--TTASPRS 363
>TC12079
Length = 414
Score = 28.5 bits (62), Expect = 0.60
Identities = 18/61 (29%), Positives = 27/61 (43%)
Frame = +2
Query: 7 PPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLS 66
P P +P P SPS+T + P + + ++ P D + HG MAP P S
Sbjct: 89 PEPASP----PNPSPSTTKPPSTIGRPKSKTPKSPPSSXDPRIRPHGDLAMAPTGPIPRS 256
Query: 67 N 67
+
Sbjct: 257 S 259
>TC15164 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, partial
(57%)
Length = 1185
Score = 28.1 bits (61), Expect = 0.78
Identities = 34/147 (23%), Positives = 52/147 (35%), Gaps = 21/147 (14%)
Frame = +2
Query: 6 KPPPCNPN--CSKPESSPSSTWFDMELPLP----------TTTSSETEPFRLDQAVCSH- 52
KPPP +P P +SP +T P P +TT++ T P + +
Sbjct: 245 KPPPPSPARLTPLPSASPVTTTSTPPTPSPAATNASPSRRSTTTTTTPPTSPTPMLLTSP 424
Query: 53 --------GLFMMAPNSWDPLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRV 104
G F+ P P+S LT P S+PS ++ +SI
Sbjct: 425 PMKLKPLPGYFLPLPTPKAPISTPLTTP---------TPKSTPSSSISPKLTRQSITAPP 577
Query: 105 HHGTHLLSPHEVRALMVPTSLTASLNS 131
H R+L T+ T +L +
Sbjct: 578 PPTESFRCNHRARSLQSSTTTTTTLTT 658
>TC19341 similar to UP|Q9CAD1 (Q9CAD1) GMP synthase, partial (28%)
Length = 484
Score = 27.7 bits (60), Expect = 1.0
Identities = 15/38 (39%), Positives = 21/38 (54%), Gaps = 2/38 (5%)
Frame = +1
Query: 7 PPPCNPNCSKPESSPSS--TWFDMELPLPTTTSSETEP 42
P P +PN + P SS + T + PLP+ T+S T P
Sbjct: 70 PSPPSPNSTPPSSSSPAAPTPSTLPTPLPSLTASSTGP 183
>TC17968 similar to PIR|F86340|F86340 protein F2D10.34 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 607
Score = 27.7 bits (60), Expect = 1.0
Identities = 17/52 (32%), Positives = 23/52 (43%), Gaps = 3/52 (5%)
Frame = +1
Query: 66 SNTLTRPLRLHDQDT--DPSSSSPSFIVTVS-QRSESIAVRVHHGTHLLSPH 114
+NT PL HD PS F +R + + +R HHG+ L PH
Sbjct: 241 NNTKLTPL*FHDSHA*NPPSLKRRRFGTAAGYRRKQGVRLRCHHGSTPLRPH 396
>TC8112 similar to UP|ODPA_PEA (P52902) Pyruvate dehydrogenase E1 component
alpha subunit, mitochondrial precursor (PDHE1-A) ,
partial (90%)
Length = 1816
Score = 27.7 bits (60), Expect = 1.0
Identities = 34/129 (26%), Positives = 48/129 (36%), Gaps = 4/129 (3%)
Frame = +1
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSE-TEPFRLDQAVCSHGLFMMAPNSW 62
L P P +PN S P PS + P PT + S+ P A H P S+
Sbjct: 148 LHLPQPSDPNSSNP*PPPSPS-AARSPPTPTQSQSKPPSPSPRTTATLPHAPSKPPPQSF 324
Query: 63 DPLSNTLTRPLRLHDQ---DTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSPHEVRAL 119
P S R + T P+SS+ S T +R R L SP + +L
Sbjct: 325 IPSSARWPRCAAWRSRLIPSTRPNSSADSATSTTGRRPSPSVWR------LGSPRRMPSL 486
Query: 120 MVPTSLTAS 128
++ + S
Sbjct: 487 LLTVTTVPS 513
>TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, partial (22%)
Length = 505
Score = 27.7 bits (60), Expect = 1.0
Identities = 14/38 (36%), Positives = 20/38 (51%)
Frame = +3
Query: 7 PPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFR 44
PP PN + P SSP++T P P+ S+ + P R
Sbjct: 327 PPSSTPNGTPPSSSPATT------PSPSRPSTPSSPTR 422
>BP054137
Length = 474
Score = 27.3 bits (59), Expect = 1.3
Identities = 12/37 (32%), Positives = 21/37 (56%)
Frame = +3
Query: 7 PPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPF 43
PPP + +P S ++T D+ + + TTT + + PF
Sbjct: 258 PPPTPSSSQQPPRSTTATIVDIVIVIATTTIALSPPF 368
>TC9951 homologue to UP|TBB3_SOYBN (P28551) Tubulin beta chain (Fragment),
partial (44%)
Length = 600
Score = 26.9 bits (58), Expect = 1.7
Identities = 33/134 (24%), Positives = 51/134 (37%)
Frame = +2
Query: 17 PESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLSNTLTRPLRLH 76
PES+ + F + + TTT E G F AP+SW LS P
Sbjct: 161 PESTTAIPSFSLNASMSTTTKPAAE-----------GTFH-APSSWI-LSQEPWIP---- 289
Query: 77 DQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSPHEVRALMVPTSLTASLNSWVFIG 136
+DP+ S+ S +T S + P + P +L +S+ SW+ G
Sbjct: 290 ---SDPARSARSSALTTSSSGNPVP-------ETTGPKGI----TPKALNSSIPSWML*G 427
Query: 137 KLISVVLFRFRLRC 150
K + + + RC
Sbjct: 428 KRLRIAIACRDFRC 469
>TC8146 similar to UP|Q94FP3 (Q94FP3) Succinate dehydrogenase subunit 3
(Fragment), partial (92%)
Length = 1038
Score = 26.9 bits (58), Expect = 1.7
Identities = 11/34 (32%), Positives = 21/34 (61%)
Frame = +3
Query: 87 PSFIVTVSQRSESIAVRVHHGTHLLSPHEVRALM 120
P+ +VT+S S+++ +HH H+V+AL+
Sbjct: 9 PNSLVTLSSLRNSLSLTIHHYVVASQIHQVQALL 110
>BP072692
Length = 422
Score = 26.9 bits (58), Expect = 1.7
Identities = 29/116 (25%), Positives = 43/116 (37%), Gaps = 14/116 (12%)
Frame = +1
Query: 25 WFDMELPLPTTTSSE--TEPFRLDQAVCSHGLFMMAPNSWDPL--------SNTLTRPLR 74
W+ + T +S+ PF + C + P+ W P+ S T +
Sbjct: 34 WWRLSRHSKLTPNSDFGARPFSSSSSTCMPSMTHQTPSPWSPIPPSAAASSSATDCHDVL 213
Query: 75 LHDQ--DTDPSSSSPSFIVTV-SQRSESIAVRVHHGTHL-LSPHEVRALMVPTSLT 126
L D +PSFI + S + V GT L L PH ++PTS T
Sbjct: 214 LRDSLLLAHTLGGAPSFITPILDSLSLPFHLIVQGGTRLPLLPHPPPVSLIPTSHT 381
>TC7879 weakly similar to GB|AAM16239.1|20334756|AY093978
At2g32150/F22D22.10 {Arabidopsis thaliana;}, partial
(92%)
Length = 1344
Score = 26.9 bits (58), Expect = 1.7
Identities = 17/40 (42%), Positives = 20/40 (49%), Gaps = 2/40 (5%)
Frame = +3
Query: 6 KPPPCNPNCSKPESSPSSTWFD--MELPLPTTTSSETEPF 43
KPPP N SKP ++PS M L TT S TE +
Sbjct: 249 KPPPSALNSSKPTAAPSPVCEH*AMMSALKITTVSSTEGY 368
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.132 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,974,131
Number of Sequences: 28460
Number of extensions: 62752
Number of successful extensions: 613
Number of sequences better than 10.0: 90
Number of HSP's better than 10.0 without gapping: 603
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 611
length of query: 163
length of database: 4,897,600
effective HSP length: 83
effective length of query: 80
effective length of database: 2,535,420
effective search space: 202833600
effective search space used: 202833600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0066.12


